Python lib for common ontology operations over a variety of backends.
This library provides a collection of different interfaces for different kinds of ontology operations, including:
- lookup of basic features of an ontology element, such as it's label, definition, relationships, or aliases
- search an ontology for a term
- validating an ontology
- updating, deleting, or modifying terms
- ontology term matching
- generating and visualizing subgraphs
- provide specialized object models for more advanced operations, such as graph traversal, or OWL axiom processing, or text annotation
These interfaces are separated from any particular backend. This means the same API can be used regardless of whether the ontology:
- is served by a remote API such as OLS or BioPortal
- is present locally on the filesystem in owl, obo, obojson, or sqlite formats
- is to be downloaded from a remote repository such as the OBO library
- is queried from a remote database, including SPARQL endpoints (Ontobee/Ubergraph), A SQL database, a Solr/ES endpoint
from src.oaklib.resource import OntologyResource
from src.oaklib.implementations.sqldb.sql_implementation import SqlImplementation
resource = OntologyResource(slug='tests/input/go-nucleus.db', local=True)
oi = SqlImplementation(resource)
for curie in oi.basic_search("cell"):
print(f'{curie} ! {oi.get_label_by_curie(curie)}')
for rel, fillers in oi.get_outgoing_relationship_map_by_curie(curie).items():
print(f' RELATION: {rel} ! {oi.get_label_by_curie(rel)}')
for filler in fillers:
print(f' * {filler} ! {oi.get_label_by_curie(filler)}')For more examples, see
Documentation here is incomplete.
See CLI docs
Use the pronto backend to fetch and parse an ontology from the OBO library, then use the search command
runoak -i obolibrary:pato.obo search osmol Returns:
PATO:0001655 ! osmolarity
PATO:0001656 ! decreased osmolarity
PATO:0001657 ! increased osmolarity
PATO:0002027 ! osmolality
PATO:0002028 ! decreased osmolality
PATO:0002029 ! increased osmolality
PATO:0045034 ! normal osmolality
PATO:0045035 ! normal osmolarity
Perform validation on PR using sqlite/rdftab instance:
runoak -i sqlite:../semantic-sql/db/pr.db validateList all terms obolibrary has for mondo
runoak -i obolibrary:mondo.obo terms Make a lexical index of all terms in Mondo:
runoak -i obolibrary:mondo.obo lexmatch -L mondo.index.yamlSearching over OBO using ontobee:
runoak -i ontobee: search tentacleyields:
http://purl.obolibrary.org/obo/CEPH_0000256 ! tentacle
http://purl.obolibrary.org/obo/CEPH_0000257 ! tentacle absence
http://purl.obolibrary.org/obo/CEPH_0000258 ! tentacle pad
...
Searching over a broader set of ontologies in bioportal (requires API KEY) (https://www.bioontology.org/wiki/BioPortal_Help#Getting_an_API_key)
runoak set-apikey bioportal YOUR-KEY-HERE
runoak -i bioportal: search tentacleyields:
BTO:0001357 ! tentacle
http://purl.jp/bio/4/id/200906071014668510 ! tentacle
CEPH:0000256 ! tentacle
http://www.projecthalo.com/aura#Tentacle ! Tentacle
CEPH:0000256 ! tentacle
...
Searching over more limited set of ontologies in Ubergraph:
runoak -v -i ubergraph: search tentacleyields
UBERON:0013206 ! nasal tentacle
runoak -i bioportal: annotate neuron from CA4 region of hippocampus of mouseyields:
object_id: CL:0000540
object_label: neuron
object_source: https://data.bioontology.org/ontologies/NIFDYS
match_type: PREF
subject_start: 1
subject_end: 6
subject_label: NEURON
object_id: http://www.co-ode.org/ontologies/galen#Neuron
object_label: Neuron
object_source: https://data.bioontology.org/ontologies/GALEN
match_type: PREF
subject_start: 1
subject_end: 6
subject_label: NEURON
...Create a SSSOM mapping file for a set of ontologies:
robot merge -I http://purl.obolibrary.org/obo/hp.owl -I http://purl.obolibrary.org/obo/mp.owl convert --check false -o hp-mp.obo
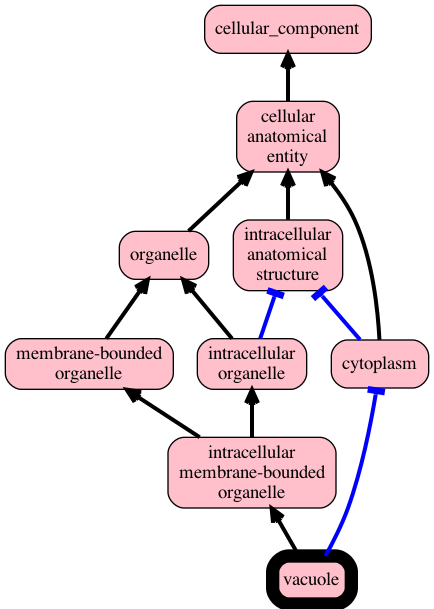
runoak lexmatch -i hp-mp.obo -o hp-mp.sssom.tsvUse the sqlite backend to visualize graph up from 'vacuole' using test ontology sqlite:
runoak -i sqlite:tests/input/go-nucleus.db viz GO:0005773Same using ubergraph, restricting to is-a and part-of
runoak -i ubergraph: viz GO:0005773 -p i,BFO:0000050Same using pronto, fetching ontology from obolibrary
runoak -i obolibrary:go.obo viz GO:0005773Currently all implementations exist in this repo/module, this results in a lot of dependencies
One possibility is to split out each implementation into its own repo and use a plugin architecture