This is the coder for the paper:
Multimodal Clinical Trial Outcome Prediction with Large Language Models
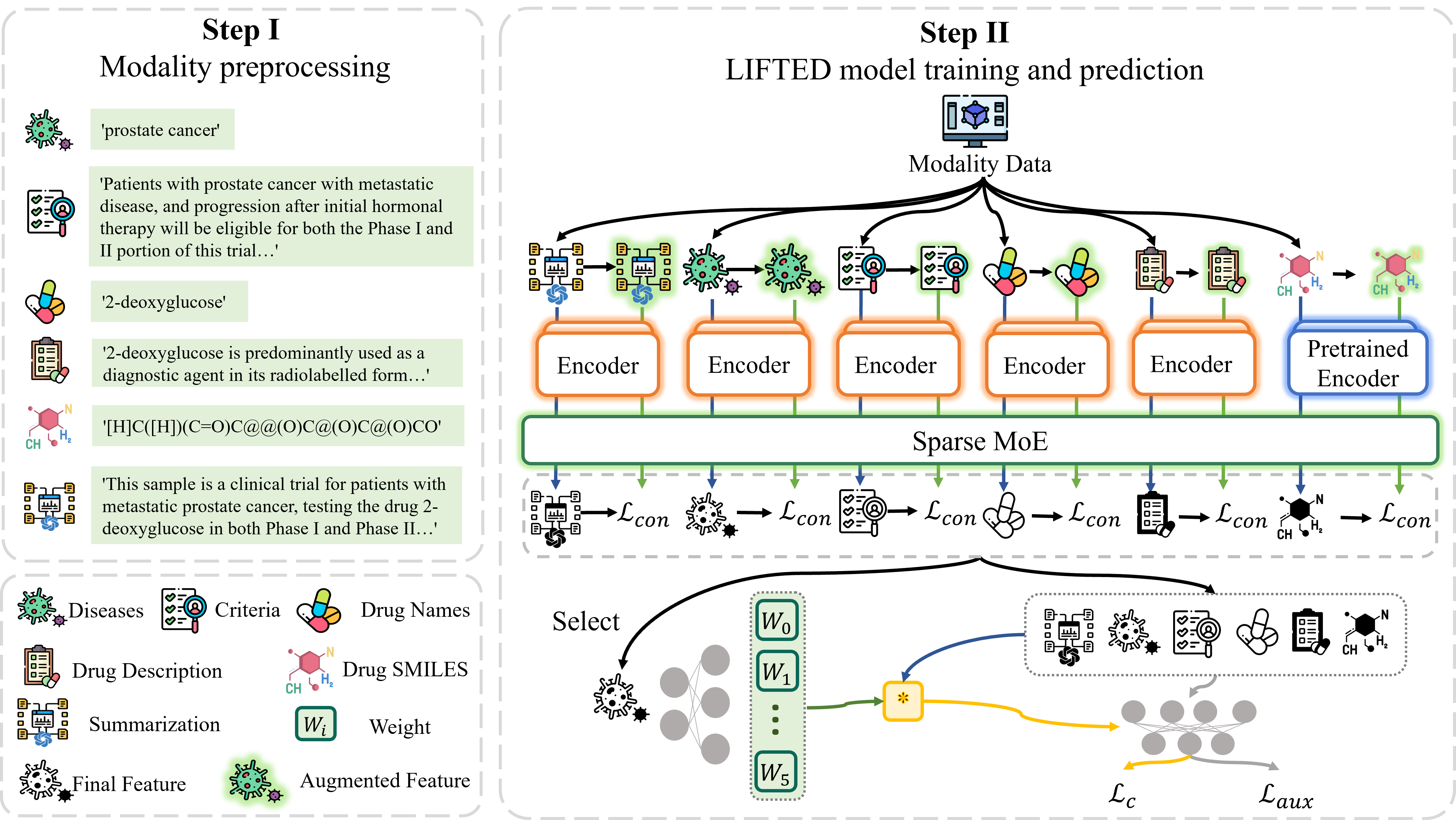
We propose an approach called muLti-modal mIx-of-experts For ouTcome prEDiction (LIFTED), which extracts information from different modalities with a transformer based unified encoder, enhances the extracted features by a Sparse Mixture-of-Experts framework and integrates multimodal information with Mixture-of-Experts, releasing the limitations of the extensibility to adapt new modalities and improving the ability to discern similar information patterns across different modalities.
An overview of our framework, LIFTED.We recommend you use conda to install this project and all required packages with the specific version to recurrent our experiments and results of them. The following commands can be used to install this project and all required packages.
conda env create -f requirements/conda.yml -n <env_name>
conda activate <env_name>
pip install -e .See the installation docs for details.
You have to set the following API keys before running experiments.
We use Wandb as our logger, so you have to create an account and login with your API key before running experiments.
wandb login
### paste your api keyWe use OpenAI API to generate summaries. You have to set your API key to call the ChatGPT 3.5 API.
echo <your-openai-key> > openai_api_key.txtWe put all the datasets under the data directory. You can create it under the root directory of this project with the following command.
mkdir dataDownload the Hint dataset to the data directory.
git clone https://github.com/futianfan/clinical-trial-outcome-prediction.git data/clinical-trial-outcome-predictionPrepare the criteria data by running the following command.
python tools/datasets/encode_criteria.pyRun the following command to generate the summarization output from the LLM model.
python tools/datasets/convert_datasets.py --tasks hint_summaryYou can train the model with the following command.
CUDA_VISIBLE_DEVICES=<gpu_ids> cli fit --config-file configs/path/to/config.yamlwhere the configs/path/to/config.yaml is the path to the config file you want to use. For instance, run the following command to train the mmcto model with data augment and auxiliary loss features on the phase I dataset.
CUDA_VISIBLE_DEVICES=<gpu_ids> cli fit --config-file configs/runs/mmcto/base/mmcto_hint_phase_I_augment_aux-loss_1x.yamlYou can evaluate your model trained in the previous step on validation or test dataset with the following command.
CUDA_VISIBLE_DEVICES=<gpu_ids> cli {validate, test} --config-file configs/runs/mmcto/base/mmcto_hint_phase_I_augment_aux-loss_1x.yaml --ckpt_path work_dirs/mmcto_hint_phase_I_augment_aux-loss_1x/<run_id>/checkpoints/<ckpt_name>.ckptwhere the {validate, test} determines the dataset you want to evaluate on, and the configs/runs/mmcto/base/mmcto_hint_phase_I_augment_aux-loss_1x.yaml is the path to the config file you want to use, and the work_dirs/mmcto_hint_phase_I_augment_aux-loss_1x/<run_id>/checkpoints/<ckpt_name>.ckpt is the path to the checkpoint you want to evaluate.
You can predict the outcome of the clinical trials and plot the weights of the Sparse Mixture-of-Experts and Mixture-of-Experts modules with the following command.
CUDA_VISIBLE_DEVICES=<gpu_ids> cli predict --config-file configs/runs/mmcto/base/mmcto_hint_phase_I_augment_aux-loss_1x.yaml --ckpt_path work_dirs/mmcto_hint_phase_I_augment_aux-loss_1x/<run_id>/checkpoints/<ckpt_name>.ckptSimilarly, the configs/runs/mmcto/base/mmcto_hint_phase_I_augment_aux-loss_1x.yaml is the path to the config file you want to use, and the work_dirs/mmcto_hint_phase_I_augment_aux-loss_1x/<run_id>/checkpoints/<ckpt_name>.ckpt is the path to the checkpoint you want to predict.
See contribution docs for details.
If you find this repo is helpful to your research, please consider citing our paper:
@article{zheng2024multimodal,
title={Multimodal Clinical Trial Outcome Prediction with Large Language Models},
author={Zheng, Wenhao and Peng, Dongsheng and Xu, Hongxia and Zhu, Hongtu and Fu, Tianfan and Yao, Huaxiu},
journal={arXiv preprint arXiv:2402.06512},
year={2024}
}