Left Atrium segmentation from LGE-MRI is the initial step for diagnosing left atrial fibrosis. However, because of the varying image quality and the different shapes of the left atrium, deep learning methods are gaining more attention on left atrium segmentation. So, we analyzed the deep learning models that perform well on medical imaging on MICCAI 2018 left atrium segmentation challenge dataset. After evaluating different metrics, the analysis shows that deep learning models with skip connection or self-attention-based models perform better on average in all metrics than other deep learning architectures. Apart from that LA wall segmentation was also implemented on University of Utah's LGE-MRI dataset. This research can be reference for other medical imaging analysis.
Project requirements/dependencies:
- PyTorch
- Albumentation
- Segmentation PyTorch Library
PyPI version:
$ pip install segmentation-models-pytorch - Monai
First Download the dataset of MICCAI Atrial Segmentation Challenge 2018.
Slice 3D MR images using:
$ python3 slice_image_mask.py
Then install Jupyter Notebook to run LA Cavity Segmentation
and LA Wall Segmentation
As it is a benchmark research, for Left Atrium segmentation and wall segmentation the following methods were used:
- Unet.
- Unet++
- MAnet
- DeepLabV3
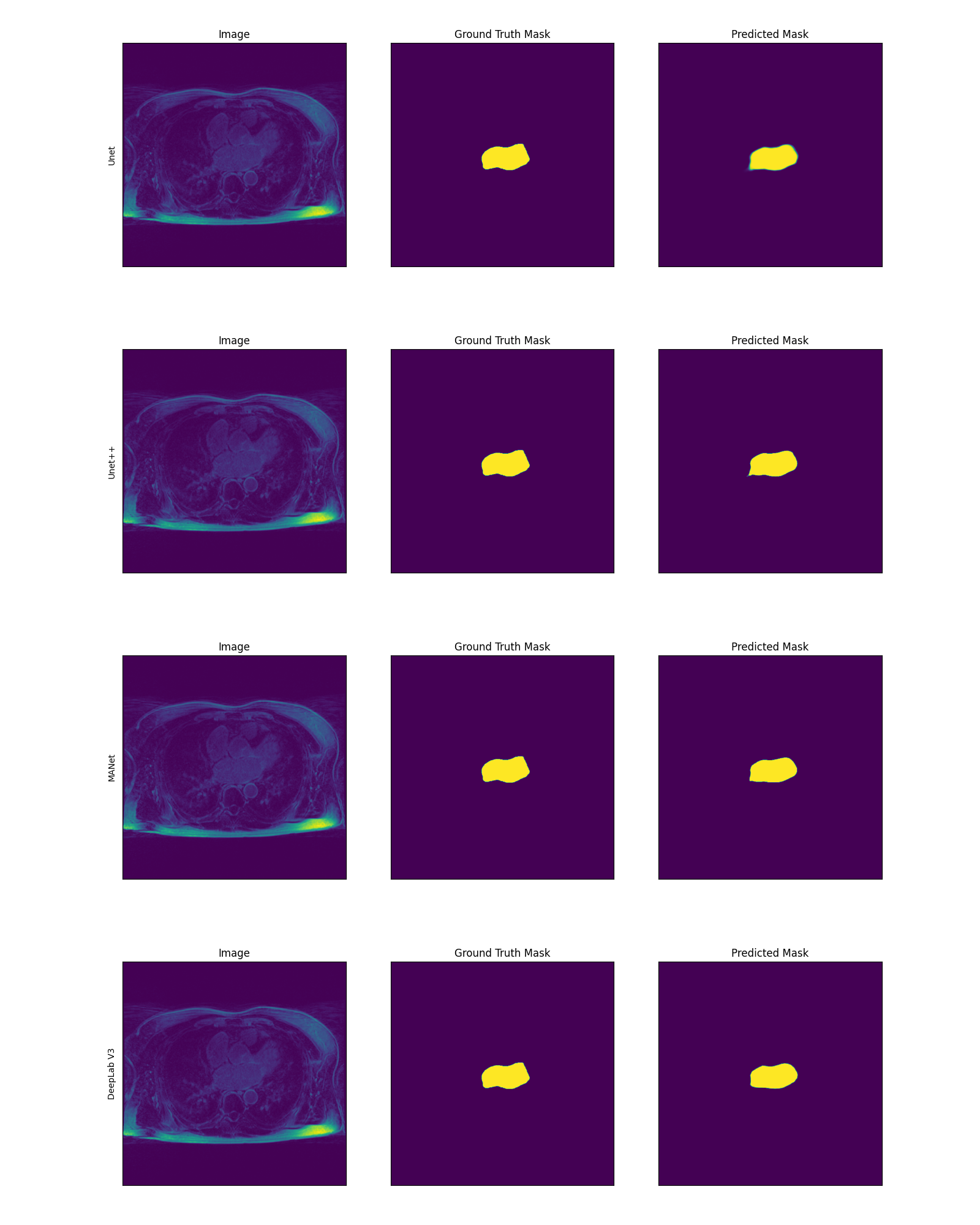
Prediction for testing dataset's Patient-3's 43rd slice with associated prediction mask and ground truth. Here every 3 image is a set and the
set of images shown are for Unet, Unet++, MANet and DeepLab V3 respectively:
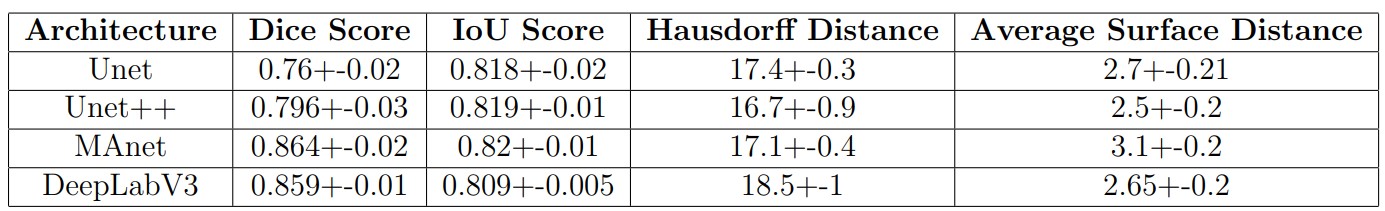
 Performance evaluation on different metrics (Dice Score, IOU Score, Hausdorff Distance, Average Surface Distance:
Performance evaluation on different metrics (Dice Score, IOU Score, Hausdorff Distance, Average Surface Distance:
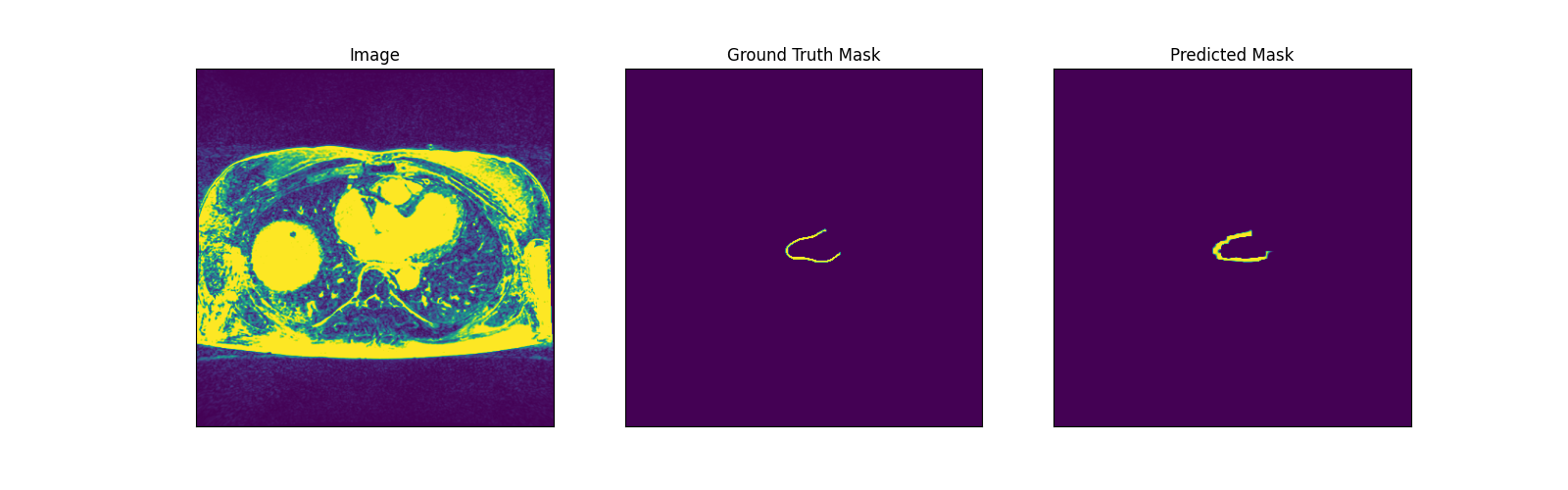
 LA Wall Prediction for Unet (Testing Data):
LA Wall Prediction for Unet (Testing Data):

- This project was conducted under the supervision of Dr. Shireen Elhabian
- Pre-trained models was based on Segmentation PyTorch Library
Created by @syedfahimahmed - feel free to contact me!