getwd()
training = read.csv("pml-training.csv")
testing = read.csv("pml-testing.csv")
summary(training)
str(training)
Cols not to be considered(row number, timestamp, value as NA/ DIV/0!):
cleanTraining = training[, colSums(is.na(training)) == 0]
cleanTraining = cleanTraining[, colSums(cleanTraining == '#DIV/0!') == 0]
cleanTraining = subset(cleanTraining, select=-c(X,user_name,raw_timestamp_part_1,raw_timestamp_part_2,cvtd_timestamp))
trainingCols = colnames(cleanTraining)
cleanTesting = subset(testing, select=trainingCols[0: (length(trainingCols)-1)])
set.seed(12345)
library(caret)
trainIndex = createDataPartition(cleanTraining$classe, p = 0.70,list=FALSE)
cleanTrainingData = cleanTraining[trainIndex,]
cleanValidationData = cleanTraining[-trainIndex,]
sample = cleanTrainingData[sample(nrow(cleanTrainingData), 1000), ]
library(caret)
ctrl = trainControl(preProcOptions = list(thresh = 0.8))
model1 = train(classe~., preProcess="pca", trControl = ctrl, data=sample, method="rpart")
predictions1 = predict(model1, cleanValidationData)
confusionMatrix(predictions1, cleanValidationData$classe)
Confusion Matrix and Statistics
Reference
Prediction A B C D E
A 1235 531 722 324 707
B 439 608 304 640 375
C 0 0 0 0 0
D 0 0 0 0 0
E 0 0 0 0 0
Overall Statistics
Accuracy : 0.3132
95% CI : (0.3013, 0.3252)
No Information Rate : 0.2845
P-Value [Acc > NIR] : 7.032e-07
Kappa : 0.0868
Mcnemar's Test P-Value : NA
library(randomForest)
ctrl = trainControl(preProcOptions = list(thresh = 0.8))
model2 = train(classe~., preProcess="pca", trControl = ctrl, data=sample, method="rf")
predictions2 = predict(model2, cleanValidationData)
confusionMatrix(predictions2, cleanValidationData$classe)
Confusion Matrix and Statistics
Reference
Prediction A B C D E
A 1639 76 2 19 2
B 7 1015 64 26 15
C 9 44 952 118 32
D 10 4 8 787 21
E 9 0 0 14 1012
Overall Statistics
Accuracy : 0.9184
95% CI : (0.9111, 0.9253)
No Information Rate : 0.2845
P-Value [Acc > NIR] : < 2.2e-16
Kappa : 0.8967
Mcnemar's Test P-Value : < 2.2e-16
library(e1071)
ctrl = trainControl(preProcOptions = list(thresh = 0.8))
model3 = svm(classe~., preProcess="pca", trControl = ctrl, data=sample)
predictions3 = predict(model3, cleanValidationData)
confusionMatrix(predictions3, cleanValidationData$classe)
Confusion Matrix and Statistics
Reference
Prediction A B C D E
A 1445 108 79 66 12
B 39 781 58 16 85
C 80 165 802 132 84
D 95 49 79 719 55
E 15 36 8 31 846
Overall Statistics
Accuracy : 0.7805
95% CI : (0.7697, 0.791)
No Information Rate : 0.2845
P-Value [Acc > NIR] : < 2.2e-16
Kappa : 0.7224
Mcnemar's Test P-Value : < 2.2e-16
###ACCURACY based on sample size
sample rpart randomForest svm
1000 0.3132 0.9184 0.7805
10000 0.3579 0.9526 0.9269
ALL NA NA 0.9405
Based on the accuracy the Random Forest model is selected.
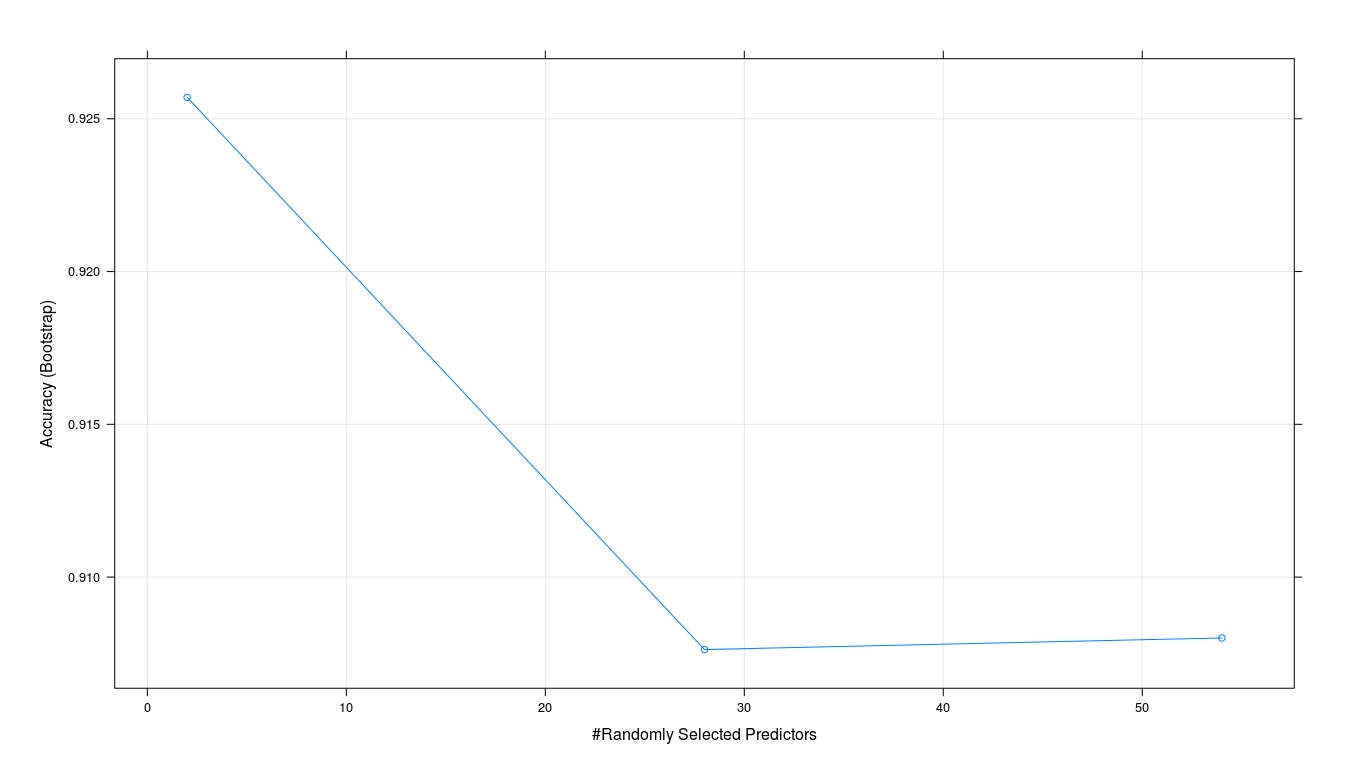
plot(model2)
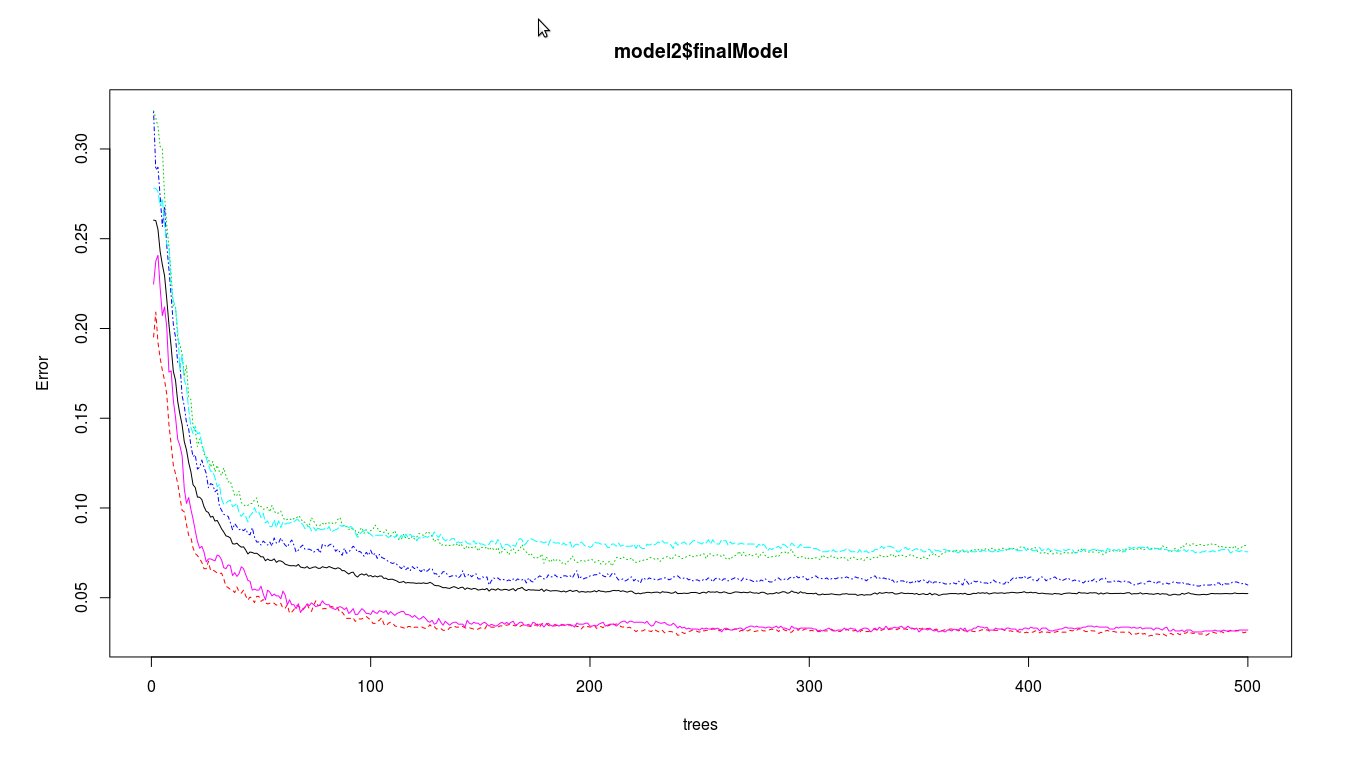
plot(model2$finalModel)
Using randomForest using 10000 training samples
predict(model2,cleanTesting)
Predictions obtained for the the 20 test cases
B A B A A E D B A A B C B A E E A B B B