Ellis Hughes
Sandbox is intended to provide a solution for reproducible research by reducing the potential for interference of a dirty environment. This is done by executing either your scripts or your provided code in a completely separate r-session.
Currently this package is only available on GitHub:
devtools::install_github("thebioengineer/sandbox")The basic use of sandbox would be to wrap your current code in the sandbox function, as below. That's it!
library(sandbox)
sandbox({
suppressPackageStartupMessages({
library(tidyverse)
})
attach(diamonds)
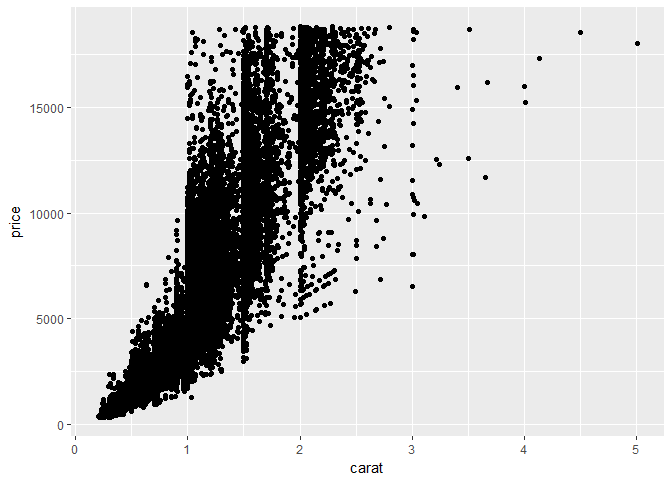
ggplot(diamonds, aes(x=carat, y=price)) + geom_point()
diamonds%>%
mutate(carat_binned=cut(carat, c(0,.5,.9,1,1.5,1.9,2,2.5,3,4,5,6)))%>%
group_by(carat_binned,cut)%>%
summarize(avgPrice=mean(price))%>%
spread(cut,avgPrice)
})By executing the code, it will return the plot generated by ggplot, and the output of the summarization of the diamonds dataset into your console. Your environment will remain clean, and none of the tidyverse packages will be loaded into your namespace on your current machine:
## # A tibble: 11 x 6
## carat_binned Fair Good `Very Good` Premium Ideal
## <fct> <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 (0,0.5] 1028. 786. 766. 863. 864.
## 2 (0.5,0.9] 2297. 2520. 2548. 2387. 2425.
## 3 (0.9,1] 3715. 4586. 4922. 4698. 4861.
## 4 (1,1.5] 5003. 5897. 6502. 6260. 7055.
## 5 (1.5,1.9] 7899. 9810. 11066. 11039. 11741.
## 6 (1.9,2] 11551. 14393. 14903. 14062. 14667.
## 7 (2,2.5] 11524. 14483. 15135. 14946. 15568.
## 8 (2.5,3] 13212. 15402. 15036. 15987. 16333
## 9 (3,4] 13611 18359 15669 14914. 14427.
## 10 (4,5] 17930 NA NA 15223 NA
## 11 (5,6] 18018 NA NA NA NA
ls()## character(0)
"tidyverse"%in%loadedNamespaces()## [1] FALSE
In situations where you may want return the outputs of the processing, you can 'leak' objects out of the sandbox environment into your global environment. This time, we are not plotting so there will be no output. However, when we ls() the current R session, summarizedDiamonds now exists.
library(sandbox)
sandbox({
suppressPackageStartupMessages({
library(tidyverse)
})
attach(diamonds)
summarizedDiamonds<-diamonds%>%
mutate(carat_binned=cut(carat, c(0,.5,.9,1,1.5,1.9,2,2.5,3,4,5,6)))%>%
group_by(carat_binned,cut)%>%
summarize(avgPrice=mean(price))%>%
spread(cut,avgPrice)
leak(summarizedDiamonds)
})
ls()## [1] "summarizedDiamonds"
While other packages, such as callr, also provide similar abilities, sandbox as the unique benefit of returning all the outputs of your code - print outs, plots, even objects - not just one output. Also, sandbox does not require using anonymous functions to pass complex workflows.