A genotype-phenotype mapping associates genotypes with phenotypes, e.g., HIV genotypes with their resistance to a certain drug. In this project we pretrain a neural network on the genotypes and thereby achieve better genotype-phenotype regression performance in the low data regime.
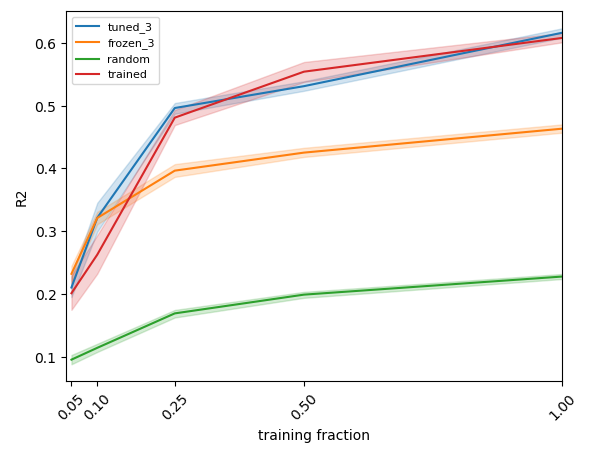
The following plot summarizes our results.
- random: we use a randomly initiallized fully connected neural network to extract features for a linear regression model,
- frozen_3: we use a pretrained neural network to extract features for a linear regression model,
- trained: we train a neural network to solve the regression task,
- tuned_3: we finetune the pretrained neural network on the regression task.
In order to compute confidence intervals, we repeated experiment 10 times.
First, install dependencies
# clone project
git clone git@github.com:wendlerc/ssl_genopheno.git
# install project
cd csss
conda create -n ssl_genopheno python=3.9
conda activate ssl_genopheno
pip install -r requirements.txt
conda install jupyterNext, pretrain a neural network with either
python genotype/pretrain.py --wandb_name pretrained or
python genotype/pretrain_vicreg.py --wandb_name pretrained_vicregFinetune the resulting model either using
python genotype/regression.py --wandb_pretrained pretrainedor
python project/cfo/regression_vicreg.py --wandb_pretrained pretrained_vicregEvaluate using a frozen pretrained model using
python project/cfo/regression_sklearn.py --wandb_pretrained pretrained