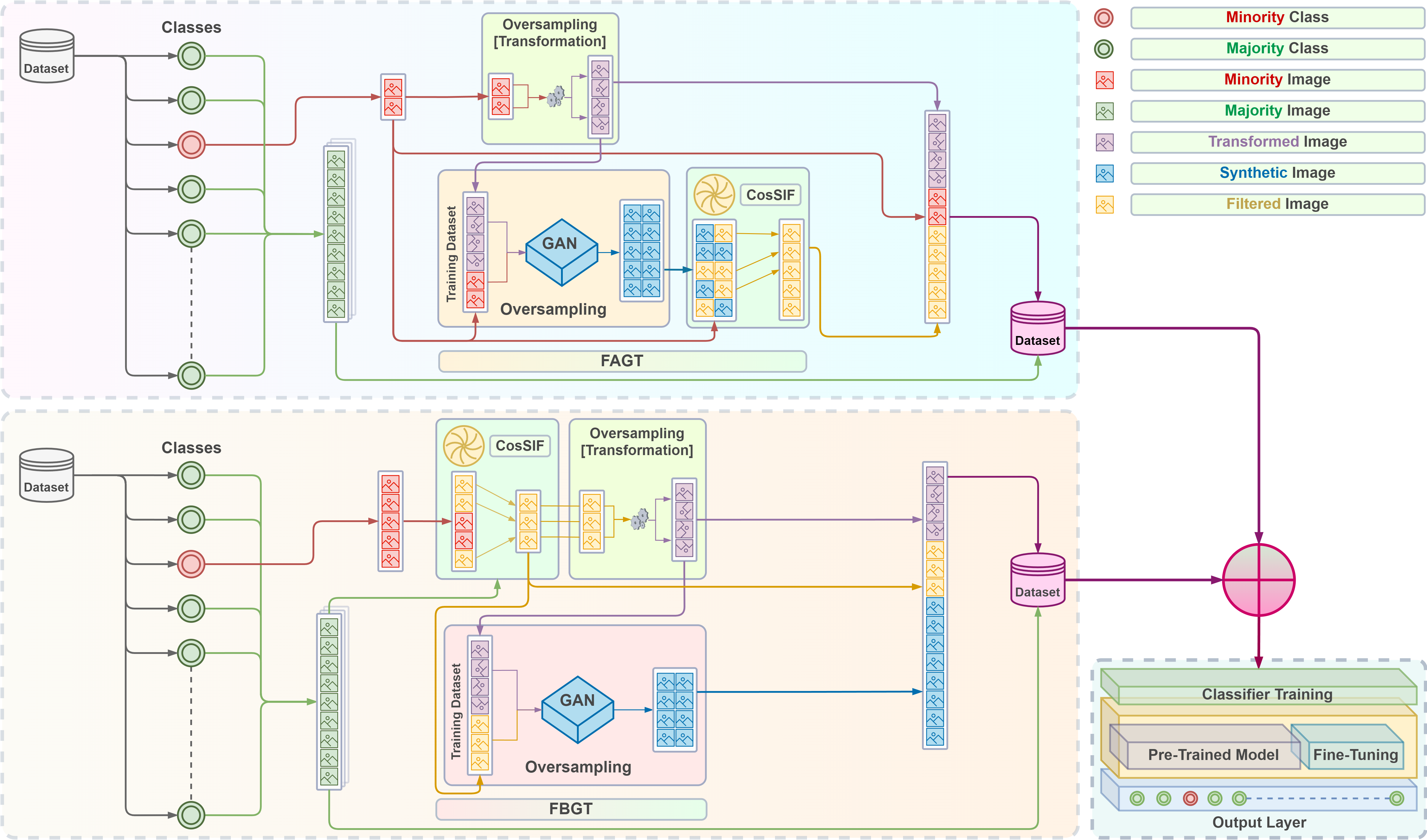
This is the official code for our CosSIF: Cosine similarity-based image filtering to overcome low inter-class variation in synthetic medical image datasets paper.
Journal: Computers in Biology and Medicine, Volume 172, April 2024, 108317
This code requires two separate conda environments. Run the following to install the required packages on Windows.
cd environments/
conda env create -f csf-ovs.yml
conda env create -f csf-model.ymlThe following notebooks require the csf-ovs environment to be activated.
scripts/
|-- HAM10000/
|---- HAM10000_filtering/
|------ HAM10000_FAGT.ipynb
|------ HAM10000_FBGT.ipynb
|---- HAM10000_GAN_train.ipynb
|---- HAM10000_augment_ds.ipynb
|---- HAM10000_oversampling.ipynb
|---- HAM10000_pre-processing.ipynb
|-- ISIC-2016/
|---- ISIC-2016_filtering/
|------ ISIC-2016_FAGT.ipynb
|------ ISIC-2016_FBGT.ipynb
|---- ISIC-2016_GAN_train.ipynb
|---- ISIC-2016_augment_ds.ipynb
|---- ISIC-2016_oversampling.ipynb
|---- ISIC-2016_pre-processing.ipynb
Run the following to activate the csf-ovs environment.
conda activate csf-ovsThe rest of the notebooks require the csf-model environment to be activated. Run the following to activate the csf-model environment.
conda activate csf-modelThe conda environments can also be setup manually. Install the packages mentioned packages.txt file.
Install Visual Studio 2019 along with the following workloads:
- Desktop development with C++
- Universal Windows Platform development
Download and extract the HAM10000 and ISIC-2016 (Task-3) datasets, along with their associated ground truth files. Then, create a folder named datasets in the root directory and organize the folders and ground truth files in the following order:
datasets/
|-- HAM10000/
|---- downloads/
|------ HAM10000_images/
|------ HAM10000_metadata.csv
|-- ISIC-2016/
|---- downloads/
|------ ISBI2016_ISIC_Part3_Training_Data/
|------ ISBI2016_ISIC_Part3_Test_Data/
|-------ISBI2016_ISIC_Part3_Training_GroundTruth.csv
|-------ISBI2016_ISIC_Part3_Test_GroundTruth.csv
Note that you may need to rename the file and folder names as required.
To begin the pre-processing, run the following scripts associated with each dataset:
ISIC-2016_pre-processing.ipynb
HAM10000_pre-processing.ipynb
For oversampling, run the following scripts associated with each dataset:
ISIC-2016_oversampling.ipynb
HAM10000_oversampling.ipynb
During the execution of the oversampling scripts, there comes a certain point where it becomes necessary to also execute the filtering and GAN training scripts.
For filtering, run the following scripts associated with each dataset:
ISIC-2016_FBGT.ipynb
ISIC-2016_FAGT.ipynb
HAM10000_FBGT.ipynb
HAM10000_FAGT.ipynb
For GAN training, run the following scripts associated with each dataset on Kaggle:
ISIC-2016_GAN_train.ipynb
HAM10000_GAN_train.ipynb
Note that the saving of the trained GANs on Kaggle must be conducted manually. The details are provided on the scripts and must be followed accordingly.
To achieve the final augmented datasets associated with each different experiment, run the following codes.
ISIC-2016_augment_ds.ipynb
HAM10000_augment_ds.ipynb
To train the models on the oversampled ISIC-2016 dataset and its filtered variants, navigate to the following directory and run the scripts.
scripts/
|-- ISIC-2016/
|---- ISIC-2016_model_train/
|------ ConvNeXt/
|------ Swin-Transformer/
|------ ViT/
To train the models on the oversampled HAM10000 dataset and its filtered variants, navigate to the following directory and upload the scripts on Kaggle along with its corresponding variants of the dataset.
scripts/
|-- HAM10000/
|---- HAM10000_model_train/
|------ ConvNeXt/
|------ Swin-Transformer/
|------ ViT/
Note that the saving of the trained models on Kaggle must be conducted manually. The details are provided on the scripts and must be followed accordingly.
The performance evaluation of all our pre-trained models can be done anytime, provided that the models reside in the following folder order.
models/
|-- HAM10000/
|---- ConvNeXt/
|------ ham10000-convnext-fagt-alpha-3
|------ ham10000-convnext-fbgt-alpha-1
|------ ham10000-convnext-no-filtering
|---- Swin-Transformer/
|------ ham10000-swin-fagt-alpha-1
|------ ham10000-swin-fagt-alpha-2
|------ ham10000-swin-fagt-alpha-3
|------ ham10000-swin-fbgt-alpha-1
|------ ham10000-swin-fbgt-alpha-2
|------ ham10000-swin-fbgt-alpha-3
|------ ham10000-swin-no-filtering
|---- ViT/
|------ ham10000-vit-fagt-alpha-3
|------ ham10000-vit-fbgt-alpha-1
|------ ham10000-vit-no-filtering
|-- ISIC-2016/
|---- ConvNeXt/
|------ ISIC-2016-convnext-fagt-alpha-3
|------ ISIC-2016-convnext-fbgt-alpha-1
|------ ISIC-2016-convnext-no-filtering
|---- Swin-Transformer/
|------ ISIC-2016-swin-fagt-alpha-1
|------ ISIC-2016-swin-fagt-alpha-2
|------ ISIC-2016-swin-fagt-alpha-3
|------ ISIC-2016-swin-fbgt-alpha-1
|------ ISIC-2016-swin-fbgt-alpha-2
|------ ISIC-2016-swin-fbgt-alpha-3
|------ ISIC-2016-swin-no-filtering
|---- ViT/
|------ ISIC-2016-vit-fagt-alpha-3
|------ ISIC-2016-vit-fbgt-alpha-1
|------ ISIC-2016-vit-no-filtering
To evaluate the performance of all trained models on the oversampled ISIC-2016 dataset and its variants, run the following script:
ISIC-2016_model_test.ipynb
To evaluate the performance of all trained models on the oversampled HAM10000 dataset and its variants, run the following script:
HAM10000_model_test.ipynb
We provide our pre-trained models on GitHub Releases for reproducibility. Below are the best-performing pre-trained models.
| Dataset | Model | Filtering | Sensitivity (%) | False Negative | Download |
|---|---|---|---|---|---|
| ISIC-2016 | ViT | FAGT (α = 0.75) | 72.00 | 21 | download |
| Dataset | Model | Filtering | Accuracy (%) | F1-score (%) | Download |
|---|---|---|---|---|---|
| HAM10000 | ConvNeXt | FAGT (α = 0.85) | 94.44 | 84.06 | download |
We offer CosSIF as an end-to-end module, available on PyPI. To implement this filtering method, kindly adhere to the subsequent instructions:
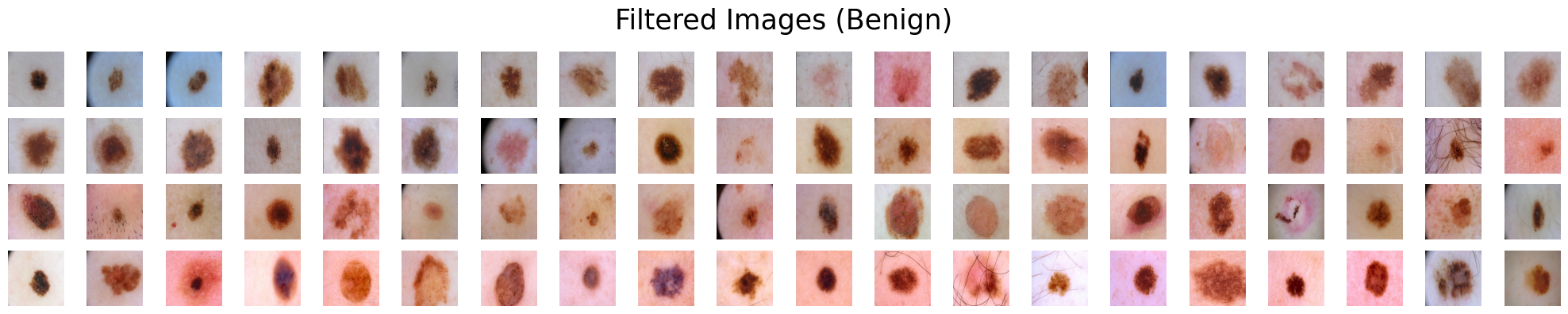
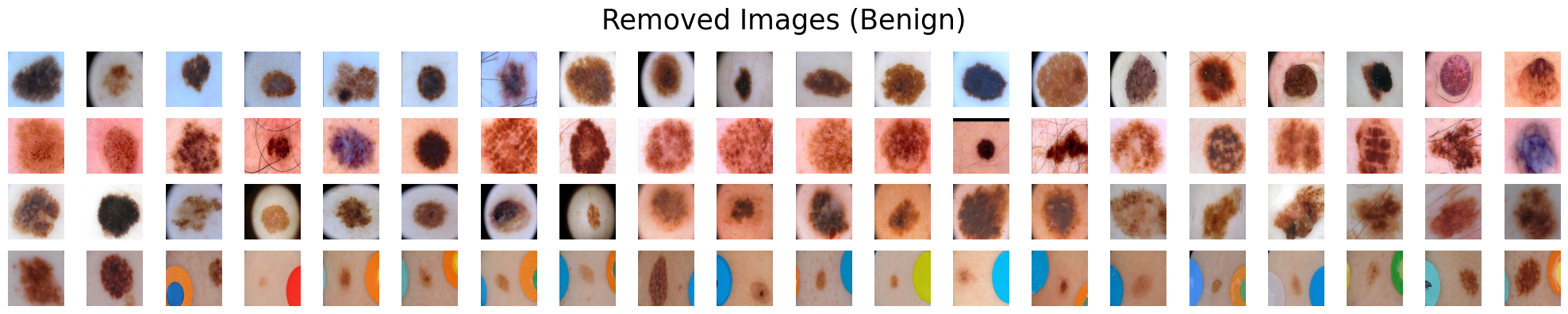
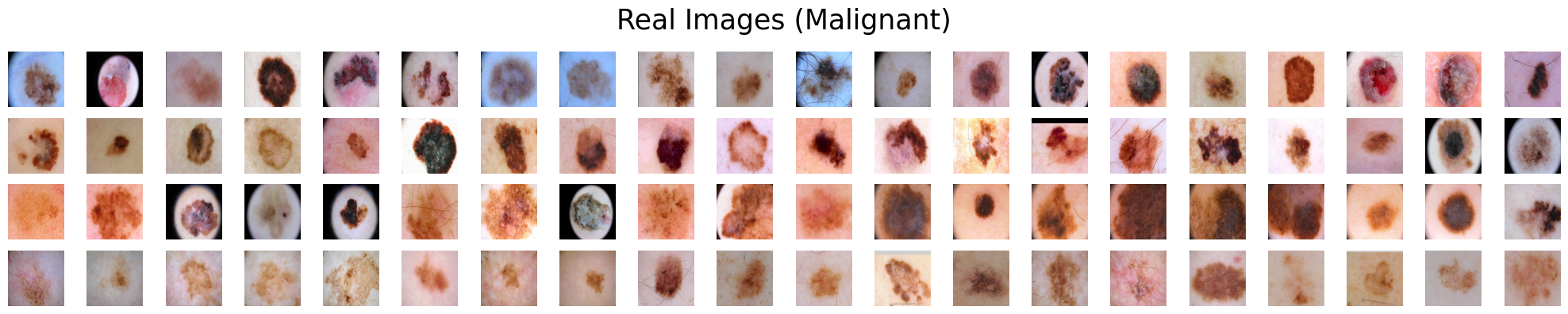
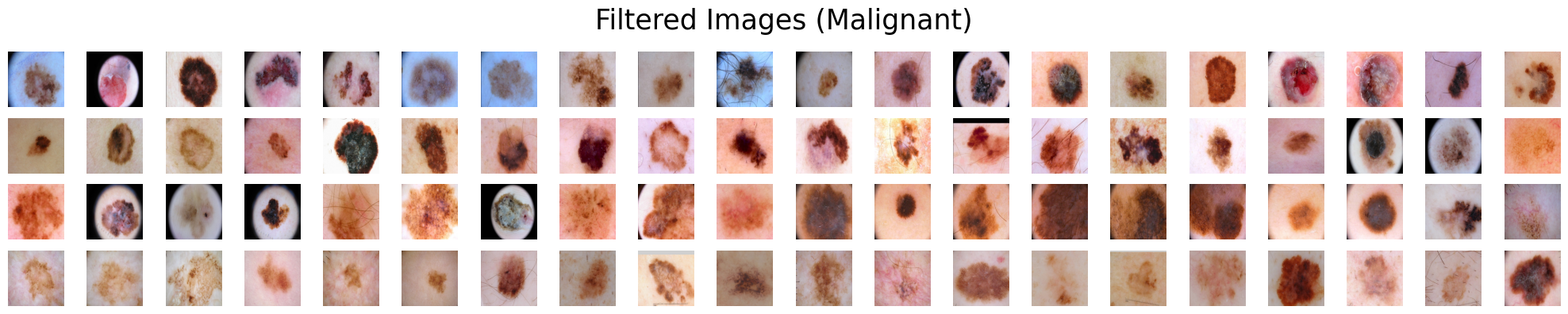
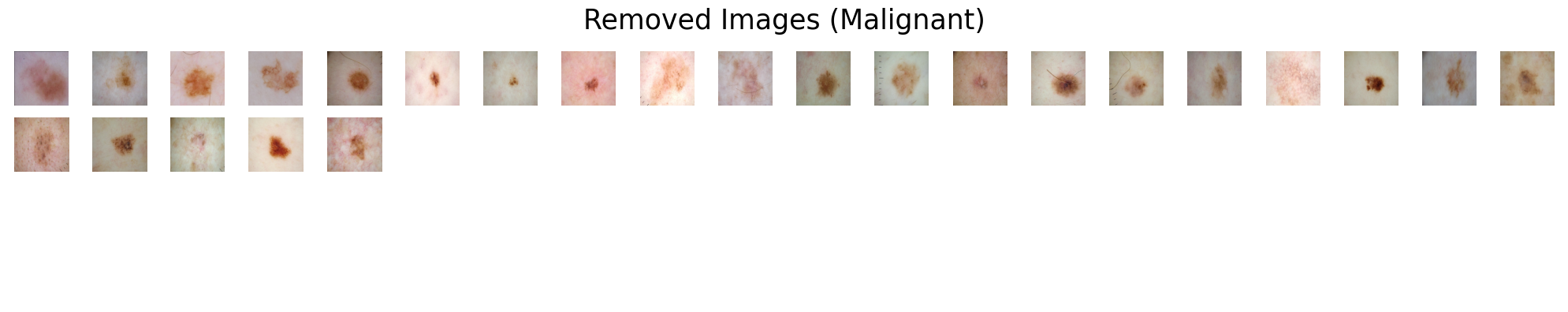
Suppose we are presented with two categories of skin lesions: benign and malignant. Our objective is to sift out benign images that bear resemblance to malignant ones, and similarly, malignant images that bear resemblance to benign ones.
from cossif import cossifcsf = cossif.CosSIF()# Filter benign images
csf.calculate_and_filter(
target_path=cls_benign, # Path to the folder that contains benign images.
secondary_path=cls_malignant, # Path to the folder that contains malignant images.
filtered_path=cls_benign_filtered, # Path to the folder that will contain filtered benign images.
removed_path=cls_benign_removed, # Path to the folder that will contain removed benign images.
record_save_path=save_path, # Save path of the similarity calculation record.
record_keep=False, # Chose whether to keep the similarity calculation record or not.
file_name='t_benign_x_s_malignant', # Name of the similarity calculation record file.
filter_type='dissimilar', # Type of filtering. Can be either "similar" or "dissimilar".
filter_range=0.85, # The filtering ratio. A value of 0.85 means that 15% of the images will be removed.
)# Filter malignant images
csf.calculate_and_filter(
target_path = cls_malignant,
secondary_path = cls_benign,
filtered_path=cls_malignant_filtered,
removed_path=cls_malignant_removed,
record_save_path = save_path,
record_keep=False,
file_name = 't_malignant_x_s_benign',
filter_type = 'similar',
filter_range = 0.85,
)This code base is built on top of the following repositories:
This code utilizes the following models for fine-tuning:
- https://huggingface.co/microsoft/swin-tiny-patch4-window7-224
- https://huggingface.co/google/vit-base-patch16-224
- https://huggingface.co/facebook/convnext-tiny-224
- Copyright @ Mominul Islam.
- ORCID iD: https://orcid.org/0009-0001-6409-964X
- Copy or sharing of this code is strictly prohibited until further notice.