This package provides a minimal wrapper for the ISRIC Soilgrids API, allowing users to query soil properties by latitude/longitude and perform basic analyses on the returned data.
Functions:
get_soilgrids(): Provides a simple wrapper for the /properties/query API endpoint, parsing the geojson response into a pandas DataFrame.
Classes:
SoilGrids(): Provides methods for querying data from Soilgrids and performing aggregation, analysis, and visualisation.
Useful links:
- Documentation for the API: https://rest.isric.org/soilgrids/v2.0/docs
- ISRIC REST entry page, including fair use policy: https://rest.isric.org
- ISRIC data and software policy: https://www.isric.org/about/data-policy
- Soilgrids FAQ: https://www.isric.org/explore/soilgrids/faq-soilgrids
Contents:
- Introduction
- Installation
- Querying data using
SoilGrids - Getting the property (clay, sand, silt) with the highest value for each point
- Analysing the relationship between clay, sand, silt and organic carbon stock
- Additional features
- Testing
- Disclaimer/licensing
This package can be installed from GitHub using pip:
python -m pip install git+https://github.com/wurli/soilgridsThe following code reads in the mean values for clay, sand, silt, and organic carbon stock (OCS) in the top 30cm of soil for a random set of 50 points within roughly 25km of Herning, Denmark. Points can be queried at a maximum rate of 5/minute, so the following code takes about 10 minutes to run:
from IPython.display import Markdown, Image
from soilgrids import SoilGrids
import logging
import plotly.io as pio
import plotly.offline as po
# Turn off console logs for cleaner notebook output
logging.getLogger('soilgrids').setLevel(logging.ERROR)
# Helper for displaying tables as markdown
show = lambda df: display(Markdown(df.to_markdown(index=False)))
sg = SoilGrids()
# get_points_sample() reads in a uniformly distributed random sample of
# points within specified bounds. If you wanted to query an exact set of
# points, you could use `sg.get_points()`.
sg.get_points_sample(
50,
lat_min=55.958103, lat_max=56.225297,
lon_min=8.662215, lon_max=9.354390,
soil_property=['clay', 'sand', 'silt', 'ocs'],
depth=['0-5cm', '5-15cm', '15-30cm', '0-30cm'],
value='mean'
)
# Once Soilgrids has been queried, the results are accessible using the
# `data` property. For brevity, only a subset of the data is shown here.
show(sg.data[0:15].filter([
'lat', 'lon', 'soil_property', 'mapped_units',
'target_units', 'depth', 'mean'
]))| lat | lon | soil_property | mapped_units | target_units | depth | mean |
|---|---|---|---|---|---|---|
| 56.0024 | 9.17168 | clay | g/kg | % | 0-5cm | 93 |
| 56.0024 | 9.17168 | clay | g/kg | % | 5-15cm | 90 |
| 56.0024 | 9.17168 | clay | g/kg | % | 15-30cm | 85 |
| 56.0024 | 9.17168 | ocs | t/ha | kg/m² | 0-30cm | 69 |
| 56.0024 | 9.17168 | sand | g/kg | % | 0-5cm | 802 |
| 56.0024 | 9.17168 | sand | g/kg | % | 5-15cm | 804 |
| 56.0024 | 9.17168 | sand | g/kg | % | 15-30cm | 820 |
| 56.0024 | 9.17168 | silt | g/kg | % | 0-5cm | 105 |
| 56.0024 | 9.17168 | silt | g/kg | % | 5-15cm | 106 |
| 56.0024 | 9.17168 | silt | g/kg | % | 15-30cm | 95 |
| 56.1016 | 8.93631 | clay | g/kg | % | 0-5cm | 63 |
| 56.1016 | 8.93631 | clay | g/kg | % | 5-15cm | 58 |
| 56.1016 | 8.93631 | clay | g/kg | % | 15-30cm | 83 |
| 56.1016 | 8.93631 | ocs | t/ha | kg/m² | 0-30cm | 59 |
| 56.1016 | 8.93631 | sand | g/kg | % | 0-5cm | 805 |
The SoilGrids class provides a handy utility rank_properties() for finding
the most abundant soil types (i.e. properties) for each point. We see that
throughout the Herning region, the main soil properties are, in order, sand,
silt, and clay:
show(sg.rank_properties(
subset=['clay', 'sand', 'silt'],
top_depth=0,
bottom_depth=30
))| lat | lon | depth | mapped_units | property_no1 | property_no2 | property_no3 |
|---|---|---|---|---|---|---|
| 55.9681 | 9.19413 | 0-30cm | g/kg | sand: 863 | silt: 75 | clay: 62 |
| 55.969 | 8.84612 | 0-30cm | g/kg | sand: 858 | silt: 89 | clay: 52 |
| 55.9736 | 9.26458 | 0-30cm | g/kg | sand: 846 | silt: 92 | clay: 63 |
| 55.988 | 9.06893 | 0-30cm | g/kg | sand: 852 | silt: 96 | clay: 51 |
| 56.0024 | 9.17168 | 0-30cm | g/kg | sand: 812 | silt: 101 | clay: 88 |
| 56.0071 | 8.73347 | 0-30cm | g/kg | sand: 846 | silt: 98 | clay: 55 |
| 56.0079 | 9.1555 | 0-30cm | g/kg | sand: 847 | silt: 101 | clay: 52 |
| 56.0115 | 9.19478 | 0-30cm | g/kg | sand: 876 | silt: 73 | clay: 50 |
| 56.0116 | 9.11962 | 0-30cm | g/kg | sand: 854 | silt: 95 | clay: 51 |
| 56.0129 | 9.06321 | 0-30cm | g/kg | sand: 865 | silt: 81 | clay: 54 |
| 56.0152 | 9.09177 | 0-30cm | g/kg | sand: 855 | silt: 93 | clay: 52 |
| 56.0209 | 9.04625 | 0-30cm | g/kg | sand: 868 | silt: 75 | clay: 57 |
| 56.0215 | 9.01066 | 0-30cm | g/kg | sand: 870 | silt: 92 | clay: 37 |
| 56.0218 | 9.32185 | 0-30cm | g/kg | sand: 793 | silt: 129 | clay: 78 |
| 56.0227 | 8.77494 | 0-30cm | g/kg | sand: 659 | silt: 263 | clay: 77 |
| 56.0245 | 8.98394 | 0-30cm | g/kg | sand: 858 | silt: 96 | clay: 45 |
| 56.0293 | 8.80021 | 0-30cm | g/kg | sand: 786 | silt: 134 | clay: 80 |
| 56.0312 | 9.20504 | 0-30cm | g/kg | sand: 838 | silt: 102 | clay: 60 |
| 56.0312 | 9.33958 | 0-30cm | g/kg | sand: 807 | silt: 121 | clay: 71 |
| 56.0322 | 9.13423 | 0-30cm | g/kg | sand: 864 | silt: 83 | clay: 55 |
| 56.0327 | 8.69833 | 0-30cm | g/kg | sand: 857 | silt: 93 | clay: 50 |
| 56.0335 | 8.82781 | 0-30cm | g/kg | sand: 820 | silt: 112 | clay: 67 |
| 56.0421 | 8.89465 | 0-30cm | g/kg | sand: 819 | silt: 106 | clay: 74 |
| 56.0457 | 9.2178 | 0-30cm | g/kg | sand: 848 | silt: 101 | clay: 50 |
| 56.0524 | 9.05007 | 0-30cm | g/kg | sand: 846 | silt: 98 | clay: 57 |
| 56.0591 | 9.0177 | 0-30cm | g/kg | sand: 815 | silt: 112 | clay: 73 |
| 56.0706 | 9.04175 | 0-30cm | g/kg | sand: 840 | silt: 108 | clay: 53 |
| 56.0731 | 9.1996 | 0-30cm | g/kg | sand: 845 | silt: 93 | clay: 61 |
| 56.0768 | 9.11822 | 0-30cm | g/kg | sand: 857 | silt: 93 | clay: 50 |
| 56.0801 | 8.89161 | 0-30cm | g/kg | sand: 830 | silt: 106 | clay: 62 |
| 56.0936 | 8.88005 | 0-30cm | g/kg | sand: 841 | silt: 92 | clay: 65 |
| 56.1016 | 8.93631 | 0-30cm | g/kg | sand: 791 | silt: 138 | clay: 71 |
| 56.1029 | 9.29378 | 0-30cm | g/kg | sand: 759 | silt: 132 | clay: 111 |
| 56.1034 | 9.1627 | 0-30cm | g/kg | sand: 864 | silt: 87 | clay: 49 |
| 56.1084 | 9.21805 | 0-30cm | g/kg | sand: 846 | silt: 84 | clay: 71 |
| 56.1126 | 8.80794 | 0-30cm | g/kg | sand: 836 | silt: 98 | clay: 65 |
| 56.1153 | 8.81545 | 0-30cm | g/kg | sand: 833 | silt: 107 | clay: 60 |
| 56.1223 | 9.24917 | 0-30cm | g/kg | sand: 838 | silt: 95 | clay: 67 |
| 56.1239 | 8.72794 | 0-30cm | g/kg | sand: 826 | silt: 109 | clay: 65 |
| 56.124 | 8.96254 | 0-30cm | g/kg | sand: 810 | silt: 125 | clay: 66 |
| 56.127 | 8.91474 | 0-30cm | g/kg | sand: 649 | silt: 252 | clay: 100 |
| 56.1365 | 9.0808 | 0-30cm | g/kg | sand: 786 | silt: 111 | clay: 104 |
| 56.1644 | 9.26652 | 0-30cm | g/kg | sand: 801 | silt: 117 | clay: 84 |
| 56.1707 | 8.92046 | 0-30cm | g/kg | sand: 802 | silt: 128 | clay: 70 |
| 56.177 | 8.82983 | 0-30cm | g/kg | sand: 807 | silt: 136 | clay: 57 |
| 56.1878 | 9.00154 | 0-30cm | g/kg | sand: 870 | silt: 86 | clay: 44 |
| 56.2074 | 9.23711 | 0-30cm | g/kg | sand: 807 | silt: 118 | clay: 75 |
| 56.2128 | 9.33269 | 0-30cm | g/kg | sand: 729 | silt: 183 | clay: 87 |
| 56.2192 | 8.77554 | 0-30cm | g/kg | sand: 820 | silt: 115 | clay: 64 |
You may notice that the above table shows only 49 points; it is missing (56.1462, 8.93131). This is because Soilgrids returns null values if there is no data available for a given point, and such cases must necessarily be excluded from the ranking calculation. In this case, the missing data may be explained by the point's close proximity to the Herningsholm Stream:
agg = sg.aggregate_means(0, 30)
missing = agg[agg['mean'].isna()]
show(missing.filter(['lat', 'lon', 'soil_property', 'mean']))| lat | lon | soil_property | mean |
|---|---|---|---|
| 56.1462 | 8.93131 | clay | nan |
| 56.1462 | 8.93131 | sand | nan |
| 56.1462 | 8.93131 | silt | nan |
| 56.1462 | 8.93131 | ocs | nan |
The ocs_correlation() method fits and displays summary statistics for a linear
model with clay, sand, and silt as predictors, and OCS as the response variable.
Based on the R-squared values returned in the summary, it doesn't look like
these soil properties are particularly good predictors for OCS in this case:
print(sg.ocs_correlation(capture_output=True))Call:
lm(formula = clay + sand + silt ~ ocs, data = soilgrids_summary,
na.action = na.omit)
Residuals:
Min 1Q Median 3Q Max
-1.90210 -0.83803 0.07654 0.14061 2.11926
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 1000.55348 1.25964 794.314 <2e-16 ***
ocs -0.01068 0.02040 -0.523 0.603
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Residual standard error: 0.948 on 47 degrees of freedom
Multiple R-squared: 0.005796, Adjusted R-squared: -0.01536
F-statistic: 0.274 on 1 and 47 DF, p-value: 0.6031
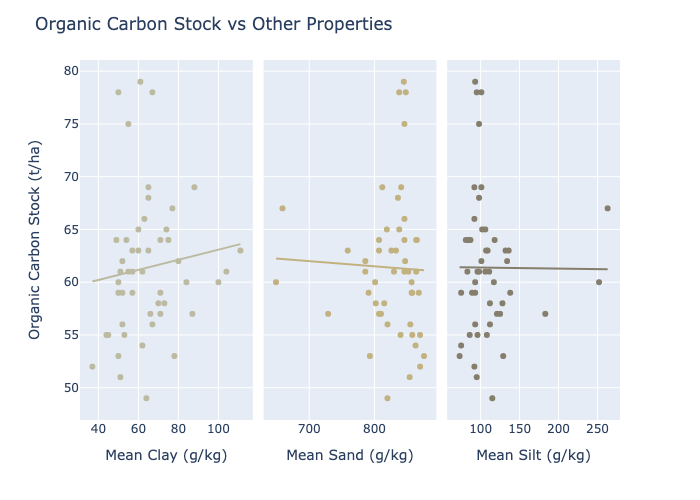
The plot_ocs_property_relationships() method can be used to obtain a graphical
representation of the relationships between OCS and the other soil properties
present in the data. These are displayed as scatterplots with overlayed lines
of best fit, i.e. the values predicted by a fitted linear regression.
In this case we can see that the plotted points never show particularly strong agreement with the fitted line, again providing evidence that clay, sand, and silt are not particularly good predictors for OCS:
# Note: statsmodels must be installed to fit the regression line
fig = sg.plot_ocs_property_relationships()
fig.write_image("README_files/ocs_property_relationships.png")
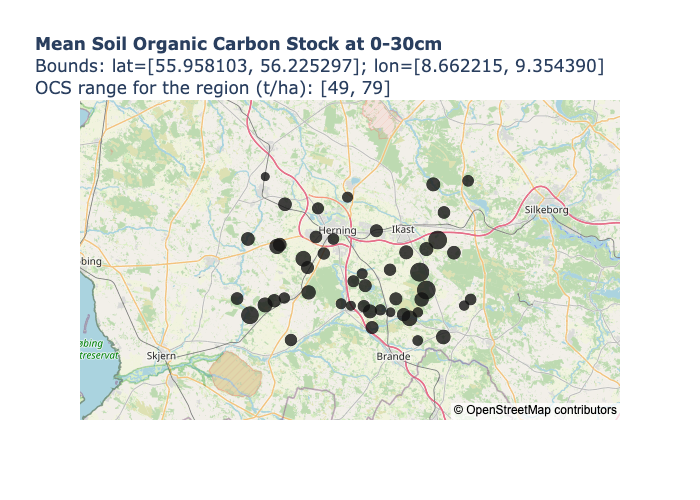
Image("README_files/ocs_property_relationships.png")The plot_property_map() method can display the points as they appear
geographically. The points are sized according to the value of the property
you choose to plot, and the tooltip also displays the values for other
properties present in the data:
fig = sg.plot_property_map('ocs', zoom=8)
fig.write_image("README_files/property_map_ocs.png")
Image("README_files/property_map_ocs.png")Working with data from SoilGrids poses a challenge since different soil properties are measured at different levels of granularity. For example, values for clay are provided at depths 0-5cm, 5-15cm, and 15-30cm, whereas OCS is provided as a single value for 0-30cm as a whole.
In order to compare OCS with clay, we need to aggregate the values for clay to get a representative mean value for the whole 0-30cm. However, since the 3 values for clay don't all cover the same amount of depth (they cover 5, 10 and 15 centimetres respectively), we can't simply take another average to obtain a figure.
SoilGrids.aggregate_means() is a utility for aggregating the mean values
provided by Soilgrids by weighting individual values according to the total
depth they represent. This method is crucial to most of the data analysis in
this package. The following shows how mean values are aggregated for a single
point:
small_datasets = [
dataset \
.filter(['lat', 'lon', 'soil_property', 'depth', 'mean']) \
.query(
"lat == 55.968112 & lon == 9.194132 &"
"soil_property in ['clay', 'ocs']"
)
for dataset in [sg.data, sg.aggregate_means(top_depth=0, bottom_depth=30)]
]
for dataset in small_datasets:
show(dataset)| lat | lon | soil_property | depth | mean |
|---|---|---|---|---|
| 55.9681 | 9.19413 | clay | 0-5cm | 58 |
| 55.9681 | 9.19413 | clay | 5-15cm | 49 |
| 55.9681 | 9.19413 | clay | 15-30cm | 72 |
| 55.9681 | 9.19413 | ocs | 0-30cm | 54 |
| lat | lon | soil_property | depth | mean |
|---|---|---|---|---|
| 55.9681 | 9.19413 | clay | 0-30cm | 62 |
| 55.9681 | 9.19413 | ocs | 0-30cm | 54 |
This package is thoroughly tested using pytest. To run the test suite, use:
python -m pytest tests- Use of Soilgrids data is subject to ISRIC data and software policy.
- This package is licensed as GPL-2.