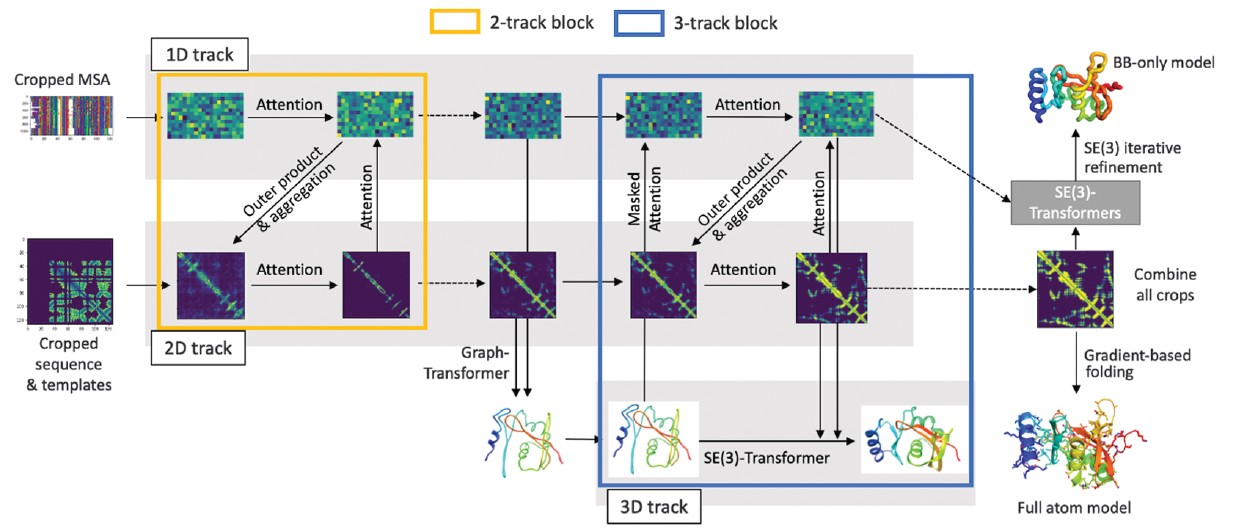
An unofficial re-implementation of RoseTTAFold, a three-track deep learning model for protein structure prediction.
pip install rosettafold-pytorchimport torch
from rosettafold-pytorch import RoseTTAFold
# dummy data
bsz, n_seq, max_len = 4, 8, 128
msa = torch.randint(0, 21, (bsz, n_seq, max_len))
seq = torch.randint(0, 21, (bsz, max_len))
aa_idx = torch.arange(0, max_len).unsqueeze(0).repeat(bsz, 1)
model = RoseTTAFold(
d_input=21,
d_msa=384,
d_pair=288
d_node=32,
d_edge=32,
d_state=32,
n_two_track_blocks=8,
n_three_track_blocks=5,
n_neighbors=[128, 128, 64, 64, 64],
n_encoder_layers=4,
p_dropout=0.1,
use_template=False
)
logits, xyz, plddt = model(msa, seq, aa_idx)
# logits for inter-residue geometries (6d coordinates)
theta = logits['theta'] # (bsz, max_len, max_len, 37)
phi = logits['phi'] # (bsz, max_len, max_len, 19)
omega = logits['omega'] # (bsz, max_len, max_len, 37)
dist = logits['dist'] # (bsz, max_len, max_len, 37)
xyz.shape # (bsz, max_len, 3, 3) : xyz coordinates for N, CA, C atoms
plddt.shape # (bsz, max_len) : predicted lDDT score for each residueThe official RoseTTAFold implementation : https://github.com/RosettaCommons/RoseTTAFold/tree/main
The official SE(3)-Transformer implementation : https://github.com/FabianFuchsML/se3-transformer-public
@article{baek2021accurate,
title={Accurate prediction of protein structures and interactions using a three-track neural network},
author={Baek, Minkyung and DiMaio, Frank and Anishchenko, Ivan and Dauparas,
Justas and Ovchinnikov, Sergey and Lee, Gyu Rie and Wang, Jue and Cong,
Qian and Kinch, Lisa N and Schaeffer, R Dustin and others
},
journal={Science},
volume={373},
number={6557},
pages={871--876},
year={2021},
publisher={American Association for the Advancement of Science}
}