dbd is a database prototyping tool that enables data analysts and engineers to quickly load and transform data in SQL databases.
dbd helps you with following tasks:
- Loading CSV, JSON, Excel, and Parquet data to database. It supports both local and online files (HTTP URLs). Data can be loaded incrementally or in full.
- Transforming data in existing database tables using insert-from-sql statements.
- Executing DDL (Data Definition Language) SQL scripts (statements like
CREATE SCHEMA, etc.).
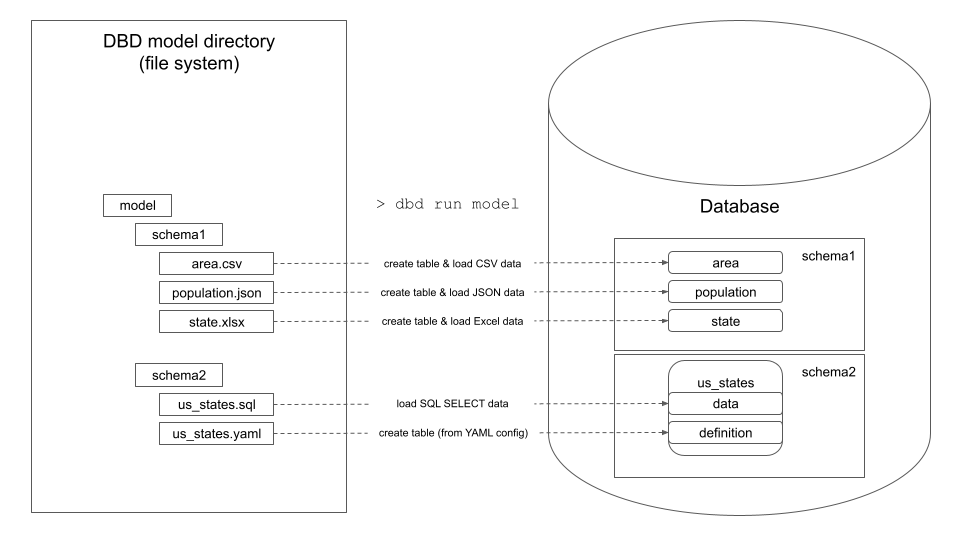
dbd processes a model directory that contains directories and files:
- Directories create new database schemas.
- Files create new database table or view. The new table's or view's name is the same as the data file name.
.csv,.json,.xlsx, and.parquetdata files are introspected and loaded to database as tables..sqlfiles that contain SQL SELECT statements are executed and the result is loaded to database as table or view..reffiles contain one or more local paths or URLs pointing to supported data files. The referenced files are loaded to database as tables..yamlfiles contain metadata for the files above. The.yamlfile has the same name as a data,.sql, or.reffile and specifies details of target table's columns (data types, constraints, indexes, etc.)..yamlfiles are optional. If not specified, dbd uses defaults (e.g.TEXTdata types for CSV columns).ddlfiles contain multiple SQL statements separated by semicolon that are executed against the database.
dbd knows the correct order in which to process files in the model directory to respect mutual dependencies between created objects.
dbd currently supports Postgres, MySQL/MariaDB, SQLite, Snowflake, BigQuery, and Redshift databases.
A short 5-minute getting started tutorial is available here.
You can also check out dbd's examples here. The easiest way how to execute them is to either clone or download dbd's github repository and start with the SQLite examples.
python3 -m venv dbd-env
source dbd-env/bin/activate
pip3 install dbd
git clone https://github.com/zsvoboda/dbd.git
cd dbd/examples/sqlite/basic
dbd run . These commands should create a new basic.db SQLite database with area, population, and state tables that are created and loaded from the corresponding files in the model directory.
dbd requires Python 3.8 or higher.
Check that you have a recent version of Python 3.8 or higher.
python3 -Vif not use a package manager to install the latest python:
On Fedora run:
sudo yum install python3On Ubuntu run:
sudo apt install python3Install Python virtual environment:
On Fedora run:
sudo yum install python3-virtualenvOn Ubuntu run:
sudo apt install python3-venvOn Windows just install Python 3.8 or higher from the Store.
Then activate the virtual environment:
On Linux run:
python3 -m venv dbd-env
source dbd-env/bin/activateOn Windows run:
python3 -m venv dbd-env
call dbd-env\Scripts\activate.batpip3 install dbdOR
git clone https://github.com/zsvoboda/dbd.git
cd dbd
pip3 install .dbd installs a command line executable that must reside on your path. Sometimes Python places the executable
(called dbd) outside of your PATH. Try to execute dbd after the installation. If the command cannot be found,
try to execute
export PATH=~/.local/bin:$PATHand run dbd again. pip3 usually complains about the fact that the directory where it is placing the executable is
not in PATH. You need to take the scripts directory that it suggests and put it on your PATH.
Once you can execute the dbd command, clone the dbd repository and start with the SQLite examples:
git clone https://github.com/zsvoboda/dbd.git
cd dbd/examples/sqlite/basic
dbd run . You can also start with this tutorial.
You can generate dbd project initial layout by executing init command:
dbd init <new-project-name>The init command generates a new dbd project directory with the following content:
modeldirectory that contains the content files.dbd.profileconfiguration file that defines database connections. The profile file is usually shared by more dbd projects.dbd.projectproject configuration file references one of the connections from the profile file and define themodeldirectory location.
dbd stores database connections in the dbd.profile configuration file. dbd searches for it in the current directory or in your home directory. You can use --profile option to point it to a profile file in different location.
The profile file is YAML file with the following structure:
databases:
db1:
db.url: <sql-alchemy-database-url>
db2:
db.url: <sql-alchemy-database-url>
db3:
db.url: <sql-alchemy-database-url>Read this document for more details about specific SQLAlchemy database URL formats.
dbd stores project configuration in project configuration file that is usually stored in your dbd project directory. dbd searches for dbd.project file in your project's directory root. You can also use the --project option of the dbd command to specify a custom project configuration file.
The project configuration file also uses YAML format and references dbd model directory and databse connection from a profile config file. All paths in project file are either absolute or relative to the directory where the profile file is located.
For example:
model: ./model
database: db2Model directory contains directories and files. Directories represent database schemas. Files, in most cases, represent database tables.
For example, this model directory layout
dbd-model-directory
+- schema1
+-- us_states.csv
+- schema2
+-- us_counties.csv
creates two database schemas: schema1 and schema2 and two database tables: us_states in schema1 and us_counties in schema2. Both tables are populated with the data from the CSV files.
dbd supports following files located in the model directory:
- DATA files:
.csv,.json,.xls,.xlsx,.parquetfiles are loaded to database as tables - REF files:
.reffiles contain one or more absolute or relative paths to local files or URLs of online data files that are loaded to database as tables. All referenced files must have the same structure as they are loaded to the same table. - SQL files:
.sqlwith SQL SELECT statements are executed using insert-from-select SQL construct. The INSERT command is generated (the SQL file only contains a SQL SELECT statement) - DDL files: contain a sequence of SQL statements separated by semicolon. The DDL files can be named
prolog.ddlandepilog.ddl. Theprolog.ddlis executed before all other files in a specific schema. Theepilog.ddlis executed last. Theprolog.ddlandepilog.ddlin the top-level model directory are executed as the very first or the very last files in the model. - YAML files: specify additional configuration for the DATA, SQL, and REF files.
.ref file contains one or more references to files that dbd loads to the database as tables. The references can be URLs, absolute file paths or paths relative to the .ref file. All referenced data files must have the same structure as they are loaded to the same database table.
Here is an example of a .ref file:
https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_daily_reports/01-03-2022.csv
https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_daily_reports/01-04-2022.csv
../data/01-05-2022.csv
../data/01-06-2022.csv
The paths and URLs can point to data files with different formats (e.g. CSV or JSON) as long as the files have the same structure (number of columns and column types).
REF files support paths that reference files inside ZIP archives using the > path separator. For example:
../data/archive.zip>covid-variants.csv
OR
https://raw.githubusercontent.com/zsvoboda/dbd/master/tests/fixtures/capabilities/zip_local/data/archive.zip>covid-variants.csv
You can reference a kaggle dataset using the kaggle://kaggle-dataset-name>dataset-file url.
For example:
kaggle://kalilurrahman/new-york-times-covid19-dataset>us.csv
To use the Kaggle API, sign up for a Kaggle account at https://www.kaggle.com. Then go to the 'Account' tab
of your user profile (https://www.kaggle.com/<username>/account) and select 'Create API Token'.
This will trigger the download of kaggle.json, a file containing your API credentials.
Place this file in the location ~/.kaggle/kaggle.json (on Windows in the location
C:\Users\<Windows-username>\.kaggle\kaggle.json - you can check the exact location,
sans drive, with echo %HOMEPATH%).
You can define a shell environment variable KAGGLE_CONFIG_DIR to change this location to $KAGGLE_CONFIG_DIR/kaggle.json
(on Windows it will be %KAGGLE_CONFIG_DIR%\kaggle.json).
For your security, ensure that other users of your computer do not have read access to your credentials. On Unix-based systems you can do this with the following command:
chmod 600 ~/.kaggle/kaggle.json
You can also choose to export your Kaggle username and token to the environment:
export KAGGLE_USERNAME=datadinosaur
export KAGGLE_KEY=xxxxxxxxxxxxxx.sql file performs SQL data transformation in the target database. It contains a SQL SELECT statement that
dbd wraps in insert-from-select statement, executes it, and stores the result into a table or view that inherits its name from the SQL file name.
Here is an example of us_states.sql file that creates a new us_states database table:
SELECT
state.abbrev AS state_code,
state.state AS state_name,
population.population AS state_population,
area.area_sq_mi AS state_area_sq_mi
FROM state
JOIN population ON population.state = state.abbrev
JOIN area on area.state_name = state.state
.yaml file specifies additional configuration for a corresponding DATA, REF or SQL file with the same base file name. Here is a YAML configuration example for the us_states.sql file above:
table:
columns:
state_code:
nullable: false
primary_key: true
type: CHAR(2)
state_name:
nullable: false
index: true
type: VARCHAR(50)
state_population:
nullable: false
type: INTEGER
state_area_sq_mi:
nullable: false
type: INTEGER
process:
materialization: table
mode: dropThis .yaml file re-types the state_population and the state_area_sq_mi columns to INTEGER, disallows NULL values in all columns, and makes the state_code column table's primary key.
You don't have to describe all table's columns. The columns that you leave out will have their types set to the default TEXT datatype in case of DATA files and is defined by the insert-from-select in case of SQL files.
The us_states.sql table is dropped and data are re-loaded in full everytime the dbd executes this model.
.yaml file's columns are mapped to a columns of the table that dbd creates from a corresponding DATA, REF or SQL file. For example, a CSV header columns or SQL SELECT column AS column clauses.
dbd supports following column's parameters:
- type: column's SQL type.
- primary_key: is the column part of table's primary key (true|false)?
- foreign_keys: all other database table columns that are referenced from a column in table (in format
foreign-table.referenced-column). - nullable: does column allow null values (true|false)?
- index: is column indexed (true|false)?
- unique: does column store unique values (true|false)?
The process section defines following processing options:
- materialization: specifies whether dbd creates a physical
tableor aviewwhen processing SQL file. The REF and DATA files always yield physical table. - mode: specifies what dbd does with table's data. You can specify values
drop,truncate, orkeep. The mode option is ignored for views.
The dbd tool's parameter --only helps with iterative development process by allowing you to specify a subset of the
tables to process. The --only parameter accepts a comma-separated list of fully qualified table names (schema.table-name).
For example:
dbd run --only stage.ext_country
only processes the ext_country table in the stage schema.
The --only parameter also processes all passed table's dependencies. You can skip the dependencies with --no-deps option.
Most of model files support Jinja2 templates. For example, this REF file loads 6 CSV files to database (4 online files from a URL and 2 from a local filesystem):
{% for n in range(4) %}
https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_daily_reports/01-0{{ n+1 }}-2022.csv
{% endfor %}
../data/01-05-2022.csv
../data/01-06-2022.csvProfile an project configuration files also us Jinja2 templates. You can expend any environment variable with the {{ environment-variable-name }} syntax.
For example, you can define your database connection parameters like username or password in environment variables and use this profile configuration file:
databases:
states_snowflake:
db.url: "snowflake://{{ SNOWFLAKE_USER }}:{{ SNOWFLAKE_PASSWORD }}@{{ SNOWFLAKE_ACCOUNT_IDENTIFIER }}/{{ SNOWFLAKE_DB }}/{{ SNOWFLAKE_SCHEMA }}?warehouse={{SNOWFLAKE_WAREHOUSE }}"
covid:
db.url: "snowflake://{{ SNOWFLAKE_USER }}:{{ SNOWFLAKE_PASSWORD }}@{{ SNOWFLAKE_ACCOUNT_IDENTIFIER }}/{{ SNOWFLAKE_DB }}/{{ SNOWFLAKE_SCHEMA }}?warehouse={{SNOWFLAKE_WAREHOUSE }}"All supported database engines except SQLite support fast data loading mode. In this mode, data are loaded to a database table using bulk load (SQL COPY) command instead of individual INSERT statements.
MySQL and Redshift require additional configuration to enable fast data loading mode. Without this extra configuration dbd reverts to slow inserting mode via INSERT statements.
To enable fast loading mode, you need specify local_infile=1 query parameter in the MySQL connection url.
You also must enable the LOCAL INFILE mode on your MySQL server. You can for example do this by executing this
SQL statement:
SET GLOBAL local_infile = trueTo enable fast loading mode, you need specify copy_stage parameter in the dbd.project configuration file.
The copy_stage parameter must reference a storage definition in your dbd.profile configuration file.
Check the example configuration files in the examples/redshift/covid_cz directory. Here are the example definitions of the
environment variables that these configuration files use:
export AWS_COVID_STAGE_S3_URL="s3://covid/stage"
export AWS_COVID_STAGE_S3_ACCESS_KEY="AKIA43SWERQGXMUYFIGMA"
export AWS_COVID_STAGE_S3_S3_SECRET_KEY="iujI78eDuFFGJF6PSjY/4CIhEJdMNkuS3g4t0BRwX"dbd code is open-sourced under BSD 3-clause license.