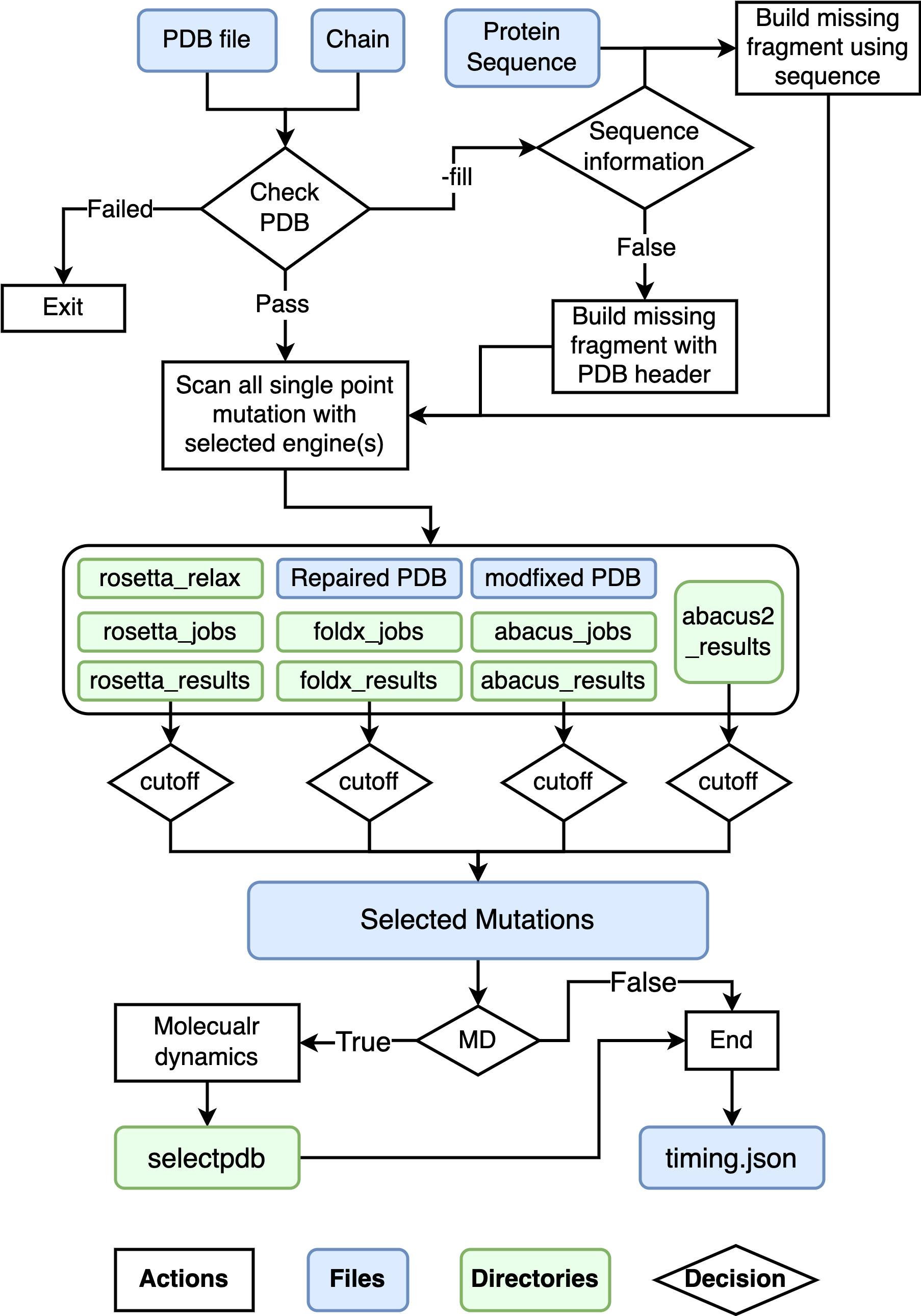
I am testing this repo with some different input structures, if you encountered any failure please post a issue.
GUI only work for FoldX.
First of all, please make sure you have added the FoldX executable to your environment! Secondly, Rosetta
(a mpi build is necessary otherwise the relax step will be skiped) is
required for cartesian_ddg (-mode slow) calculation or ddg_monomer row1 protocol(-mode fast).
Also, ABACUS is an outstanding software with great statistical energy function for protein design.
Structures downloaded from RCSB could be erroneous. One of the biggest problems that will directly affect energy calculation is breaks in chains.
Here I implemented a loop closure module using modeller, a great software with a very long history, as backend.
Due to their licenses, I cannot redistribute them here 😟 !
To our glad, openmm is open source! So the glass is half full 😃 .
Here is a good news, the ABACUS2 database is now available at https://zenodo.org/record/4533424. However, the necessary
module is not available in the zenodo version, you may use the online server at https://biocomp.ustc.edu.cn/servers/abacus-design.php to run ABACUS2.
Install DDGScan:
To avoid possilbe confilcts, create a new conda environment, and using mamba will be faster
conda create -n ddgscan python=3.9
conda activate ddgscan
conda install -c conda-forge mamba
mamba install pytorch torchvision torchaudio pytorch-cuda=11.6 -c pytorch -c nvidia
mamba install -c conda-forge openmm pdbfixer
git clone https://github.com/JinyuanSun/DDGScan.git
pip install pandas numpy joblib seaborn matplotlib venn logomaker mdtraj bio scikit-learn
python setup.py install
DDGScan -h # you should see help messageFoldX:
Register and download the executable.
Rosetta:
Follow the Rosetta document
I will recommend that users export ROSETTADB before runing grape-fast.py by appending this into ~/.bashrc:
export ROSETTADB="/path/to/rosetta/database"ABACUS1/2:
Send email to the authors for source code.
Modeller:
Get the Modeller license key at https://salilab.org/modeller/registration.html
export KEY_MODELLER=<your_key>
conda config --add channels salilab
conda install modellerI provide many options for users especially those know what they want. Here are some quick walk-through. pdb and chain are positional but you need to set
-E according to the software you have in your OS. -seq are strongly recommended to be set by the user.
Also, I highly recommend adding the -MD flag and using -P CUDA if a good gpu is available (better
than RTX2060 well be much faster than 48 core cpu). Also, I did not test how much precision dropped to use the -S fast
preset, but I do know it can be faster in about two orders of magnitude.
If using -fill flag, input structure will be automatically fixed using information from SEQRES record in native PDB
downloaded from RCSB using modeller. Model with lowest molpdf energy will be subjected to following step.
usage: DDGScan grape_phaseI [-h] [-fill] [-seq SEQUENCE] [-T THREADS] [-fc FOLDX_CUTOFF] [-rc ROSETTA_CUTOFF] [-ac ABACUS_CUTOFF] [-a2c ABACUS2_CUTOFF] [-nstruct RELAX_NUMBER]
[-nruns NUMOFRUNS] [-E {abacus,foldx,rosetta,abacus2} [{abacus,foldx,rosetta,abacus2} ...]] [-M {run,rerun,analysis,test}] [-S {fast,slow}] [-MD] [-P {CUDA,CPU}]
[-fix_mm]
pdb chain
positional arguments:
pdb Input PDB
chain Input PDB Chain to do in silico DMS
optional arguments:
-h, --help show this help message and exit
-fill, --fill_break_in_pdb
Use modeller to fill missing residues in your pdb file. Use this option with caution!
-seq SEQUENCE, --sequence SEQUENCE
The exact sequence of protein you want to design. All mutation will be named according to this sequence.
-T THREADS, --threads THREADS
Number of threads to run FoldX, Rosetta
-fc FOLDX_CUTOFF, --foldx_cutoff FOLDX_CUTOFF
Cutoff of FoldX ddg(kcal/mol)
-rc ROSETTA_CUTOFF, --rosetta_cutoff ROSETTA_CUTOFF
Cutoff of Rosetta ddg(R.E.U.)
-ac ABACUS_CUTOFF, --abacus_cutoff ABACUS_CUTOFF
Cutoff of ABACUS SEF(A.E.U.)
-a2c ABACUS2_CUTOFF, --abacus2_cutoff ABACUS2_CUTOFF
Cutoff of ABACUS2 SEF(A.E.U.)
-nstruct RELAX_NUMBER, --relax_number RELAX_NUMBER
Number of how many relaxed structure
-nruns NUMOFRUNS, --numofruns NUMOFRUNS
Number of runs in FoldX BuildModel
-E {abacus,foldx,rosetta,abacus2} [{abacus,foldx,rosetta,abacus2} ...], --engine {abacus,foldx,rosetta,abacus2} [{abacus,foldx,rosetta,abacus2} ...]
-M {run,rerun,analysis,test}, --mode {run,rerun,analysis,test}
Run, Rerun or analysis
-S {fast,slow}, --preset {fast,slow}
Fast or Slow
-MD, --molecular_dynamics
Run 1ns molecular dynamics simulations for each mutation using openmm.
-P {CUDA,CPU}, --platform {CUDA,CPU}
CUDA or CPU
-fix_mm, --fix_mainchain_missing
fixing missing backbone bone using pdbfixer
usage: DDGScan list_distribute [-h] [-msaddg] [-fill] [-fix_mm] [-T THREADS] [-nstruct RELAX_NUMBER] [-nruns NUMOFRUNS] [-E {foldx,rosetta,abacus2,rosetta_fast} [{foldx,rosetta,abacus2,rosetta_fast} ...]]
[-repair] [-MD] [-P {CUDA,CPU}]
pdb mutation_list_file
positional arguments:
pdb Input PDB
mutation_list_file Mutation list file, see README for details
optional arguments:
-h, --help show this help message and exit
-msaddg, --output_of_MSAddg
The format of MSAddg *.scan.txt, and there may be mismatch between your pdb and sequence
-fill, --fill_break_in_pdb
Use modeller to fill missing residues in your pdb file. Use this option with caution!
-fix_mm, --fix_mainchain_missing
fixing missing backbone bone using pdbfixer
-T THREADS, --threads THREADS
Number of threads to run FoldX, Rosetta or ABACUS2
-nstruct RELAX_NUMBER, --relax_number RELAX_NUMBER
Number of how many relaxed structure
-nruns NUMOFRUNS, --numofruns NUMOFRUNS
Number of runs in FoldX BuildModel
-E {foldx,rosetta,abacus2,rosetta_fast} [{foldx,rosetta,abacus2,rosetta_fast} ...], --engine {foldx,rosetta,abacus2,rosetta_fast} [{foldx,rosetta,abacus2,rosetta_fast} ...]
-repair, --foldx_repair
Run Repair before ddG calculation
-MD, --molecular_dynamics
Run 1ns molecular dynamics simulations for each mutation using openmm.
-P {CUDA,CPU}, --platform {CUDA,CPU}
CUDA or CPUusage: DDGScan analysis_and_plot [-h] [--residue_position RESIDUE_POSITION]
[--plot_type {all,venn,residue_bar,heatmap,position_avg_boxplot,variance_lineplot,kde_plot,residue_logo} [{all,venn,residue_bar,heatmap,position_avg_boxplot,variance_lineplot,kde_plot,residue_logo} ...]]
pdb results_dir
positional arguments:
pdb your target pdb file
results_dir directory of results of grape_phase_I or list_distribute
optional arguments:
-h, --help show this help message and exit
--residue_position RESIDUE_POSITION
residue position, if you asked for a barplot at residue level
--plot_type {all,venn,residue_bar,heatmap,position_avg_boxplot,variance_lineplot,kde_plot,residue_logo} [{all,venn,residue_bar,heatmap,position_avg_boxplot,variance_lineplot,kde_plot,residue_logo} ...]
plots you want to make
You may want to try it out on a small protein like Gb1:
I will recommend using the -S fast with -MD flag, and using CUDA to accelerate molecular dynamics simulations.
This is a very good crystal structure solved by X-ray, so I did not pass any value about fixing the PDB file!
Using -S slow to get more accuracy!
wget https://files.rcsb.org/download/1PGA.pdb
DDGScan grape_phaseI 1PGA.pdb A -E foldx abacus rosetta -M run -T 40 -S slow -MD -P CUDAYou should expecting outputs like:
A folder named foldx_results containing:
All_FoldX.score
MutationsEnergies_BestPerPositionBelowCutOff_SortedByEnergy.tab
MutationsEnergies_BelowCutOff.tab
MutationsEnergies_BestPerPosition_SortedByEnergy.tab
MutationsEnergies_BelowCutOff_SortedByEnergy.tab
MutationsEnergies_CompleteList.tab
MutationsEnergies_BestPerPosition.tab
MutationsEnergies_CompleteList_SortedByEnergy.tab
MutationsEnergies_BestPerPositionBelowCutOff.tab
And another folder named foldx_jobs contains many subdirectories, in each subdirectory, containing raw output for
every mutation built by FoldX. Of course, there will be directories start with rosetta or abacus, depending on your choice!
If -md was turned on, all produced snapshots can be found in selectpdb with afterMD as a suffix in the name of PDB files.
In some cases, one may need to redesign a multimer for biocatalysis. For homo-multimer, we need to manage mutations at symmetrical interfaces. To reduce the required computing resources, DDGScan treats homo-multimer as a monomer for 1st round prediction. And putative beneficial mutations located at the interface will be selected for further prediction, and multiple mutations will be made on the entire multimer.
multimer_scan.py multimer.pdbUsing scripts/inspectmutation.py to inspect mutations in pymol:
pymol inspectmutation.py $Wildtype_structure $Mutation_structure $Mutation_position $ChainAbout principles for protein physics, refer to this book.
For a given set of single-point mutations of a protein. This module distributes calculations to cores and can parse pre-defined special groups of mutations to make.
# followings are pre-defined groups:
_small: GAVSTC
_large: FYWKRHQE
_neg: DE
_pos: RK
_polar: YTSHKREDQN
_non_charged_polar: YTSNQH
_hydrophobic: FILVAGMW
_cys: C
_pro: P
_scan: ARNDCQEGHILKMFPSTWYV
Also, "dynamic selection" are supported.
# followings are dynamic selection:
@smaller: mutation to AA with smaller vdw
@bigger: mutation to AA with bigger vdw
@more_hydrophobic: mutation to AA more hydrophobic
@less_hydrophobic: mutation to AA more hydrophilic
@more_sheet_tendency: mutation to AA with higher sheet tendency
@less_sheet_tendency: mutation to AA with lower sheet tendency
@more_helix_tendency: mutation to AA with higher helix tendency
@less_helix_tendency: mutation to AA with lower helix tendency
@{random}: random is an integer in range 1 to 19 ,randomly select few mutations for you, good luck!
An example mutation list file is a plain text file seperated with space, looks like:
wildtype chain position mutation
A A 26 P
A A 26 ILV # make A -> I,L,V
A A 26 _polar # make A -> Y,T,S,H,K,R,E,D,Q,N
A A 26 @9
A A 26 @smaller # make A -> G
DDGScan also support MSAddg output, you need to add a -msaddg flag. The best 80 predictions made by MSAddg will be
selected.
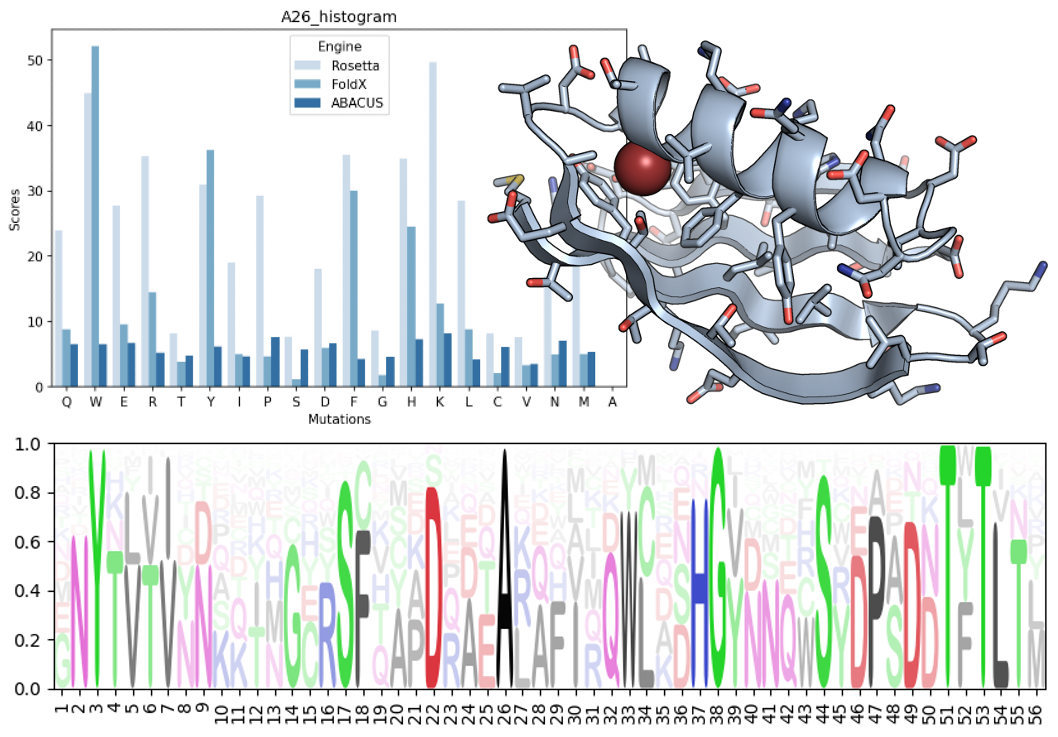
DDGScan list_distribute 1pga.pdb 1pga.fa.scan.txt -repair -msaddg -T 10 -E foldxPost analysis can help you to easily access to many kinds of plots. Here are two example of a bar-plot of a saturated single point mutation and mutation logo sequence.
2019.04: Developed GUI and single mutation scan for FoldX.
2021.10: Restart this project for Rosetta and ABACUS supporting.
2021.11: Added openmm for MDs.
2021.12: Added modeller for loop modelling and args was rewritten.
2022.03: Released a few more codes on plotting and updateed the command line interface.
2022.04: Release v0.1.0!
2022.05: Fixed some minor bugs reported.
2022.06: Add multimer_scan.py for homo-multimer interface mutation analysis.
2023.01: Add support for ddg_monomer, about 80% accuracy of the catresian_ddg, 100 times faster.
2023.01: Release v0.1.1!
Continuing...
To avoid issues caused by pdb file, it is recommended to carefully exam your input file. One can
use /path/to/rosetta/main/tools/protein_tools/scripts/clean_pdb.py
to clean pdb. However, this script will also renumber pdb file.
During test, some cases failed because of the following problems:
- Non-canonical amino acid in pdb will cause failure due to lack parameters in all predictors, therefore is not accepted.
- Gaps in pdb introduce ugly energy, you may want to apply
-fillor use model predicted by AlphaFold. Based on some case study and reports from the community, the AF2 structrue is prefectly OK as a good start point.
For this software:
@software{sun_jinyuan_2022_1046990,
author = {Sun Jinyuan},
title = {{DDGScan: an integrated parallel workflow for the
in silico point mutation scan of protein}},
month = apr,
year = 2022,
publisher = {Zenodo},
version = {v0.1.0},
doi = {10.5072/zenodo.1046990},
url = {https://doi.org/10.5072/zenodo.1046990}
}For methodology:
@article{cui2021computational,
title={Computational redesign of a PETase for plastic biodegradation under ambient condition by the GRAPE strategy},
author={Cui, Yinglu and Chen, Yanchun and Liu, Xinyue and Dong, Saijun and Tian, Yu’e and Qiao, Yuxin and Mitra, Ruchira and Han, Jing and Li, Chunli and Han, Xu and others},
journal={ACS Catalysis},
volume={11},
number={3},
pages={1340--1350},
year={2021},
publisher={ACS Publications}
}
@incollection{sun2021grape,
title={GRAPE, a greedy accumulated strategy for computational protein engineering},
author={Sun, Jinyuan and Cui, Yinglu and Wu, Bian},
booktitle={Methods in Enzymology},
volume={648},
pages={207--230},
year={2021},
publisher={Elsevier}
}MIT License
Copyright (c) 2022 jinyuan sun
Permission is hereby granted, free of charge, to any person obtaining a copy
of this software and associated documentation files (the "Software"), to deal
in the Software without restriction, including without limitation the rights
to use, copy, modify, merge, publish, distribute, sublicense, and/or sell
copies of the Software, and to permit persons to whom the Software is
furnished to do so, subject to the following conditions:
The above copyright notice and this permission notice shall be included in all
copies or substantial portions of the Software.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE
SOFTWARE.
If you need any help like installing backend software or interpreting results, you may contact me to get help by filling this form.