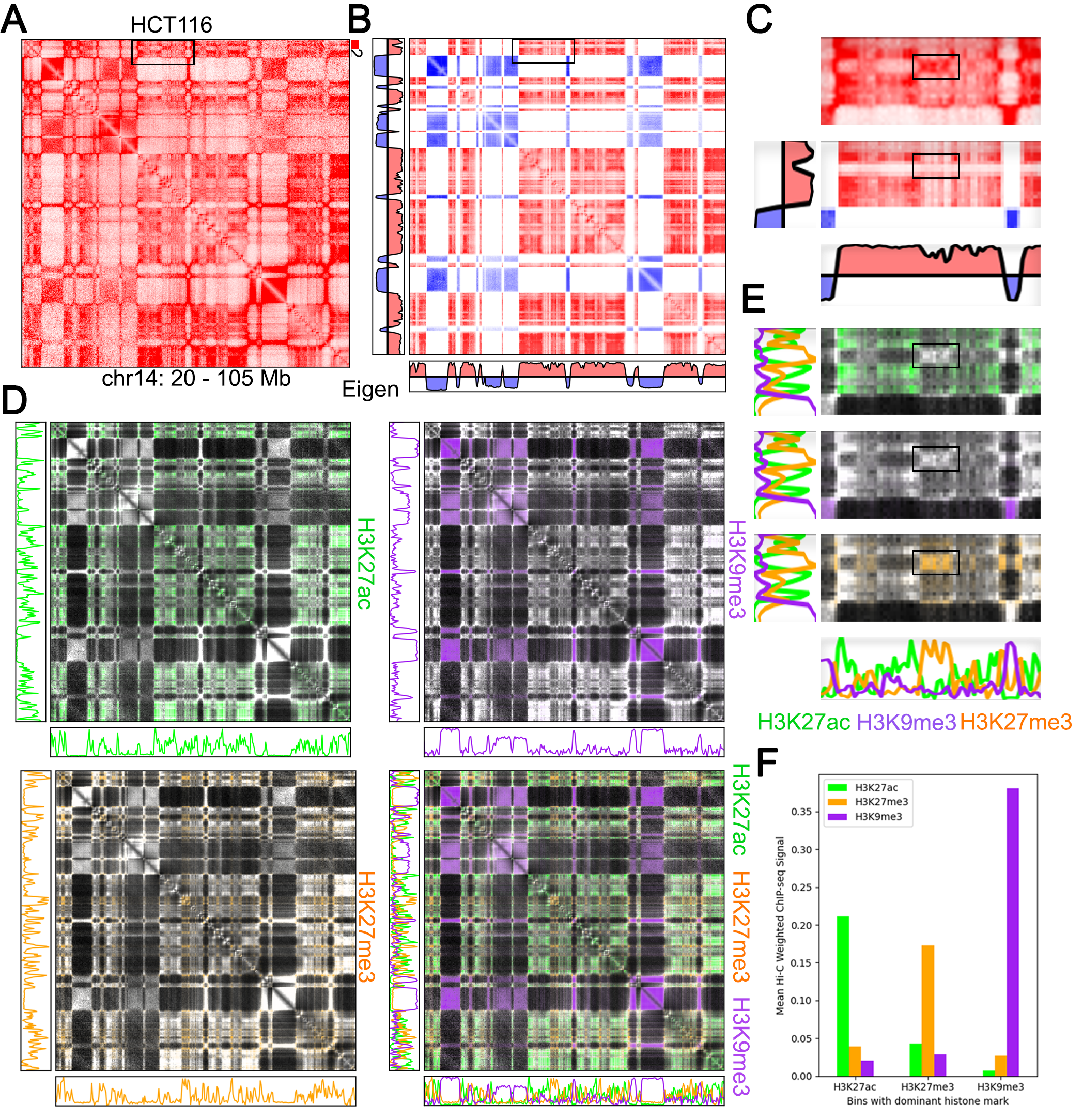
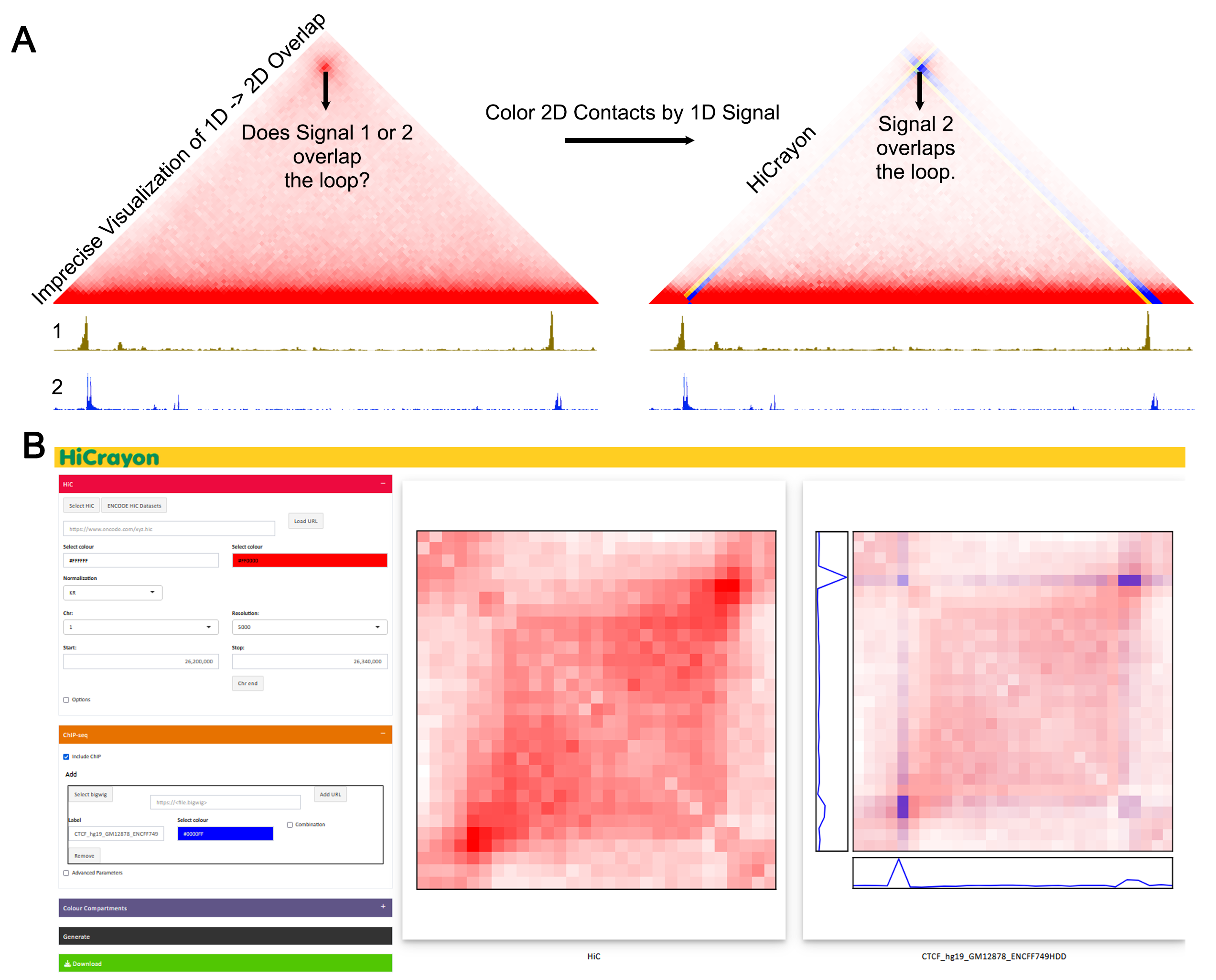
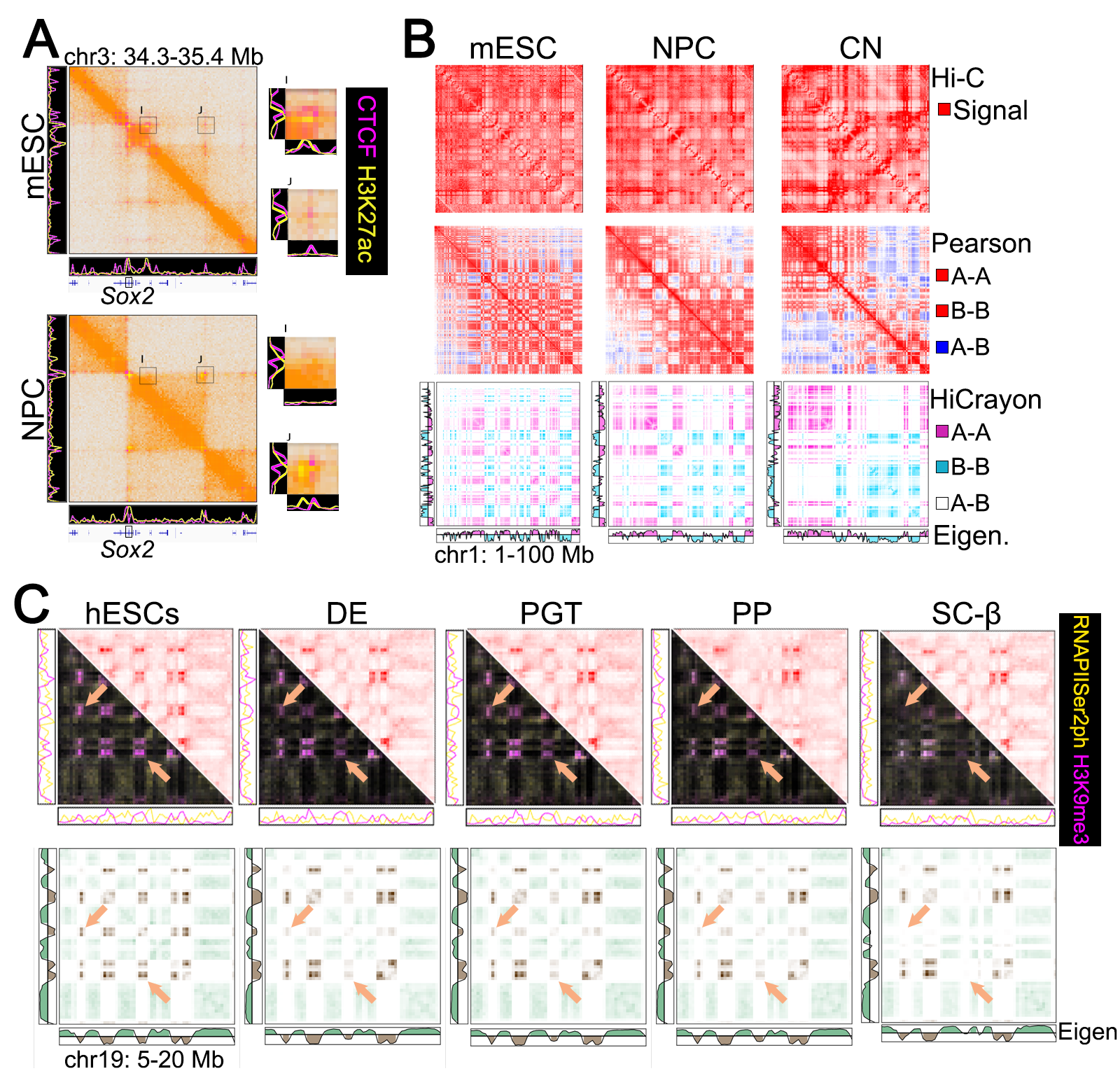
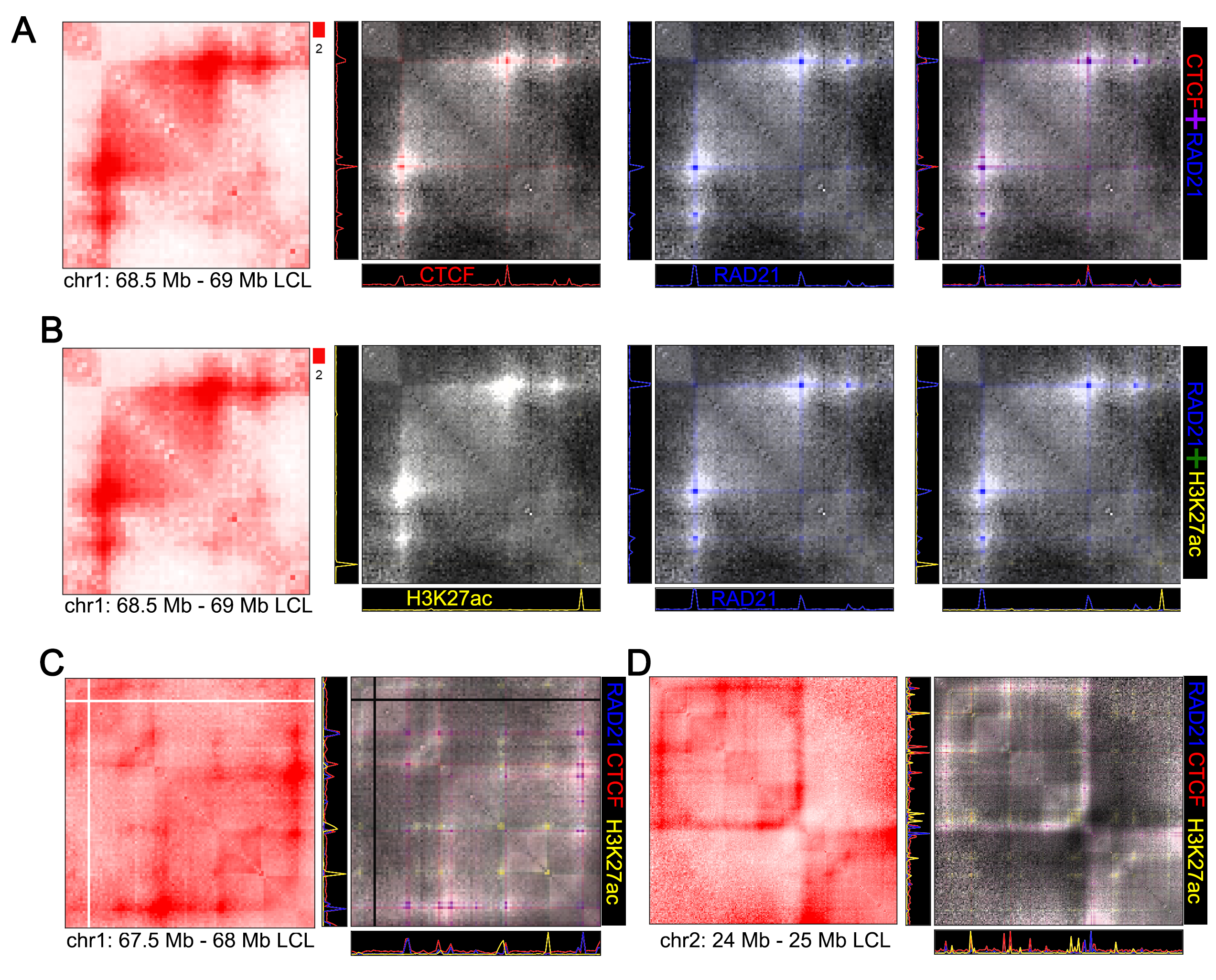
Visualize Hi-C with any bigwig or bedGraph to combine both forms of data in a single image.
- Values > 0 (ChIP-seq, RNA-seq, etc.)
- chr | start | stop | value
- Values below and above 0 (Hi-C Compartment calls)
- chr | start | stop | value
A light version of HiCrayon that allows import of publically availble Hi-C maps, bigwigs through URL eg. from ENCODE (https://www.encodeproject.org/), and locally stored small bedGraph files.
https://jrowleylab.com/HiCrayon/
To fully avail of the utility of HiCrayon, please proceed with installation below
Singularity allows deployment of HiCrayon using a container so you don't need to worry about software compatibility issues.
Build a singularity container from a docker image.
singularity build hicrayon.sif docker://nolandocker/hicrayon
Clone the hicrayon git repository.
git clone https://github.com/JRowleyLab/HiCrayon.git
cd into the HiCrayon directory and run the app inside the container
singularity exec ~/Containers/hicrayon.sif R -e "shiny::runApp('app.R', launch.browser=F, port = 3838)"
Additional parameters:
- if you'd like to attach a directory outside of your own:
-B /:/filesystem/ - host application to allow access on another device (beware security risks):
shiny::runApp(..., host="IPADDRESS")
- Create a conda environment with the hicrayon yaml file
conda create -n hicrayon hicrayon.yml
conda activate hicrayon
Clone the hicrayon git repo and cd into the repo
git clone https://github.com/JRowleyLab/HiCrayon.git
Run HiCrayon
R -e 'shiny::runApp("app.R", launch.browser=F)'