A One-Stop Large-Scale Graph Computing System from Alibaba
GraphScope is a unified distributed graph computing platform that provides a one-stop environment for performing diverse graph operations on a cluster of computers through a user-friendly Python interface. GraphScope makes multi-staged processing of large-scale graph data on compute clusters simple by combining several important pieces of Alibaba technology: including GRAPE, MaxGraph, and Graph-Learn (GL) for analytics, interactive, and graph neural networks (GNN) computation, respectively, and the vineyard store that offers efficient in-memory data transfers.
Visit our website at graphscope.io to learn more.
We provide a Playground with a managed JupyterLab. Try GraphScope straight away in your browser!
GraphScope can run on clusters managed by Kubernetes within containers. For quickly getting started, we can set up a local Kubernetes cluster and take advantage of pre-built Docker images as follows.
To run GraphScope on your local computer, the following dependencies or tools are required.
- Docker
- Python >= 3.6 (with pip)
- Local Kubernetes cluster set-up tool (e.g. Kind)
On macOS, you can follow the official guides to install them and enable Kubernetes in Docker. For Ubuntu/CentOS Linux distributions, we provide a script to install the above dependencies and prepare the environment. On Windows, you may want to install Ubuntu on WSL2 to use the script.
# run the environment preparing script.
./scripts/prepare_env.shGraphScope client is distributed as a python package and can be easily installed with pip.
pip3 install graphscopeNext, we will walk you through a concrete example to illustrate how GraphScope can be used by data scientists to effectively analyze large graphs.
Please note that we have not hardened this release for production use and it lacks important security features such as authentication and encryption, and therefore it is NOT recommended for production use (yet)!
ogbn-mag is a heterogeneous network composed of a subset of the Microsoft Academic Graph. It contains 4 types of entities(i.e., papers, authors, institutions, and fields of study), as well as four types of directed relations connecting two entities.
Given the heterogeneous ogbn-mag data, the task is to predict the class of each paper. Node classification can identify papers in multiple venues, which represent different groups of scientific work on different topics. We apply both the attribute and structural information to classify papers. In the graph, each paper node contains a 128-dimensional word2vec vector representing its content, which is obtained by averaging the embeddings of words in its title and abstract. The embeddings of individual words are pre-trained. The structural information is computed on-the-fly.
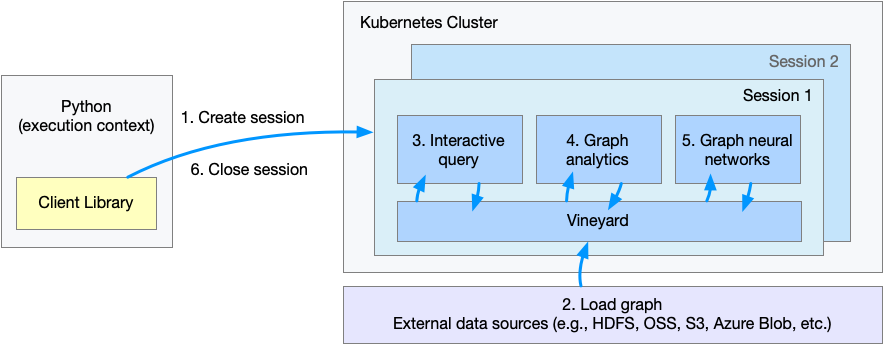
The figure shows the flow of execution when a client Python program is executed.
- Step 1. Create a session or workspace in GraphScope.
- Step 2. Define schema and load the graph.
- Step 3. Query graph data.
- Step 4. Run graph algorithms.
- Step 5. Run graph-based machine learning tasks.
- Step 6. Close the session.
To use GraphScope, we need to establish a session in a python interpreter.
import os
import graphscope
# assume we mount `~/test_data` to `/testingdata` in pods.
k8s_volumes = {
"data": {
"type": "hostPath",
"field": {
"path": os.path.expanduser("~/test_data/"),
"type": "Directory"
},
"mounts": {
"mountPath": "/testingdata"
}
}
}
sess = graphscope.session(k8s_volumes=k8s_volumes)For macOS, the session needs to establish with the LoadBalancer service type (which is NodePort by default).
sess = graphscope.session(k8s_volumes=k8s_volumes, k8s_service_type="LoadBalancer")A session tries to launch a coordinator, which is the entry for the back-end engines.
The coordinator manages a cluster of resources (k8s pods),
and the interactive/analytical/learning engines ran on them.
For each pod in the cluster, there is a vineyard instance at service for distributed data in memory.
GraphScope models graph data as property graph, in which the edges/vertices are labeled and have many properties.
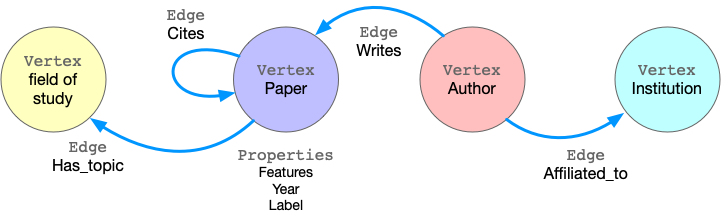
Taking ogbn-mag as example, the figure below shows the model of the property graph.
This graph has four kinds of vertices, labeled as paper, author, institution and field_of_study. There are four kinds of edges connecting them, each kind of edges has a label and specifies the vertex labels for its two ends. For example, cites edges connect two vertices labeled paper. Another example is writes, it requires the source vertex is labeled author and the destination is a paper vertex. All the vertices and edges may have properties. e.g., paper vertices have properties like features, publish year, subject label, etc.
To load this graph to GraphScope, one may use the code below with the data files. Please download and extract it to the mounted dir on local(in this case, ~/test_data).
g = sess.g()
g = g.add_vertices("/testingdata/ogbn_mag_small/paper.csv", label="paper")
g = g.add_vertices("/testingdata/ogbn_mag_small/author.csv", label="author")
g = g.add_vertices("/testingdata/ogbn_mag_small/institution.csv", label="institution")
g = g.add_vertices("/testingdata/ogbn_mag_small/field_of_study.csv", label="field_of_study")
g = g.add_edges(
"/testingdata/ogbn_mag_small/author_affiliated_with_institution.csv",
label="affiliated", src_label="author", dst_label="institution",
)
g = g.add_edges(
"/testingdata/ogbn_mag_small/paper_has_topic_field_of_study.csv",
label="hasTopic", src_label="paper", dst_label="field_of_study",
)
g = g.add_edges(
"/testingdata/ogbn_mag_small/paper_cites_paper.csv",
label="cites", src_label="paper", dst_label="paper",
)
g = g.add_edges(
"/testingdata/ogbn_mag_small/author_writes_paper.csv",
label="writes", src_label="author", dst_label="paper",
)Alternatively, we provide a function to load this graph for convenience.
from graphscope.dataset.ogbn_mag import load_ogbn_mag
g = load_ogbn_mag(sess, "/testingdata/ogbn_mag_small/")Here, the g is loaded in parallel via vineyard and stored in vineyard instances in the cluster managed by the session.
Interactive queries allow users to directly explore, examine, and present graph data in an exploratory manner in order to locate specific or in-depth information in time. GraphScope adopts a high-level language called Gremlin for graph traversal, and provides efficient execution at scale.
In this example, we use graph traversal to count the number of papers two given authors have co-authored. To simplify the query, we assume the authors can be uniquely identified by ID 2 and 4307, respectively.
# get the endpoint for submitting Gremlin queries on graph g.
interactive = sess.gremlin(g)
# count the number of papers two authors (with id 2 and 4307) have co-authored
papers = interactive.execute("g.V().has('author', 'id', 2).out('writes').where(__.in('writes').has('id', 4307)).count()").one()Graph analytics is widely used in real world. Many algorithms, like community detection, paths and connectivity, centrality are proven to be very useful in various businesses. GraphScope ships with a set of built-in algorithms, enables users easily analysis their graph data.
Continuing our example, below we first derive a subgraph by extracting publications in specific time out of the entire graph (using Gremlin!), and then run k-core decomposition and triangle counting to generate the structural features of each paper node.
Please note that many algorithms may only work on homogeneous graphs, and therefore, to evaluate these algorithms over a property graph, we need to project it into a simple graph at first.
# extract a subgraph of publication within a time range
sub_graph = interactive.subgraph("g.V().has('year', inside(2014, 2020)).outE('cites')")
# project the projected graph to simple graph.
simple_g = sub_graph.project(vertices={"paper": []}, edges={"cites": []})
ret1 = graphscope.k_core(simple_g, k=5)
ret2 = graphscope.triangles(simple_g)
# add the results as new columns to the citation graph
sub_graph = sub_graph.add_column(ret1, {"kcore": "r"})
sub_graph = sub_graph.add_column(ret2, {"tc": "r"})In addition, users can write their own algorithms in GraphScope. Currently, GraphScope support users to write their own algorithms in Pregel model and PIE model.
Graph neural networks (GNNs) combines superiority of both graph analytics and machine learning. GNN algorithms can compress both structural and attribute information in a graph into low-dimensional embedding vectors on each node. These embeddings can be further fed into downstream machine learning tasks.
In our example, we train a GCN model to classify the nodes (papers) into 349 categories, each of which represents a venue (e.g. pre-print and conference). To achieve this, first we launch a learning engine and build a graph with features following the last step.
# define the features for learning
paper_features = []
for i in range(128):
paper_features.append("feat_" + str(i))
paper_features.append("kcore")
paper_features.append("tc")
# launch a learning engine.
lg = sess.learning(sub_graph, nodes=[("paper", paper_features)],
edges=[("paper", "cites", "paper")],
gen_labels=[
("train", "paper", 100, (0, 75)),
("val", "paper", 100, (75, 85)),
("test", "paper", 100, (85, 100))
])Then we define the training process, and run it.
# Note: Here we use tensorflow as NN backend to train GNN model. so please
# install tensorflow.
from graphscope.learning.examples import GCN
from graphscope.learning.graphlearn.python.model.tf.trainer import LocalTFTrainer
from graphscope.learning.graphlearn.python.model.tf.optimizer import get_tf_optimizer
# supervised GCN.
def train(config, graph):
def model_fn():
return GCN(
graph,
config["class_num"],
config["features_num"],
config["batch_size"],
val_batch_size=config["val_batch_size"],
test_batch_size=config["test_batch_size"],
categorical_attrs_desc=config["categorical_attrs_desc"],
hidden_dim=config["hidden_dim"],
in_drop_rate=config["in_drop_rate"],
neighs_num=config["neighs_num"],
hops_num=config["hops_num"],
node_type=config["node_type"],
edge_type=config["edge_type"],
full_graph_mode=config["full_graph_mode"],
)
trainer = LocalTFTrainer(
model_fn,
epoch=config["epoch"],
optimizer=get_tf_optimizer(
config["learning_algo"], config["learning_rate"], config["weight_decay"]
),
)
trainer.train_and_evaluate()
config = {
"class_num": 349, # output dimension
"features_num": 130, # 128 dimension + kcore + triangle count
"batch_size": 500,
"val_batch_size": 100,
"test_batch_size": 100,
"categorical_attrs_desc": "",
"hidden_dim": 256,
"in_drop_rate": 0.5,
"hops_num": 2,
"neighs_num": [5, 10],
"full_graph_mode": False,
"agg_type": "gcn", # mean, sum
"learning_algo": "adam",
"learning_rate": 0.0005,
"weight_decay": 0.000005,
"epoch": 20,
"node_type": "paper",
"edge_type": "cites",
}
train(config, lg)See more details in node_classification_on_citation.ipynb, with the running results. Please note that learning feature of GraphScope is not support on macOS yet.
At last, we close the session after processing all graph tasks.
sess.close()This operation will notify the backend engines and vineyard to safely unload graphs and their applications, Then, the coordinator will dealloc all the applied resources in the k8s cluster.
GraphScope ships with a Dockerfile that can build docker images for releasing. The images are built on a builder image with all dependencies installed and copied to
a runtime-base image. To build images with latest version of GraphScope, go to the root directory and run this command.
# for the first time, run this script to install make, doxygen for docs and java env for testing.
# ./scripts/prepare_dev.sh
make graphscope
make interactive_manager
# by default, the built image is tagged as graphscope/graphscope:SHORTSHA and graphscope/maxgraph_standalone_manager:SHORTSHAGraphScope python interface is separate with the engines image. If you are developing python client and not modifying the protobuf files, the engines image doesn't require to be rebuilt.
You may want to re-install the python client on local.
cd python
python3 setup.py installNote that the learning engine client has C/C++ extensions modules and setting up the build environment is a bit tedious. By default the locally-built client library doesn't include the support for learning engine. If you want to build client library with learning engine enabled, please refer Build Python Wheels.
To verify the correctness of your developed features, your code changes should pass our tests.
You may run the whole test suite with commands:
# for the first time, run this script to install make, doxygen for docs and java env for testing.
# ./scripts/prepare_dev.sh
# run all test cases
./scripts/test.sh --all
# run all test cases on your built image
./scripts/test.sh --all --gs_image graphscope/graphscope:SHORTSHA --gie_manager_image graphscope/maxgraph_standalone_manager:SHORTSHA
# or run the selected cases on a certain module. e.g.,
./scripts/test.sh --python
./scripts/test.sh --gieDocumentation can be generated using Sphinx. Users can build the documentation using:
# build the docs
make graphscope-docs
# to open preview on local
open docs/_build/html/index.htmlThe latest version of online documentation can be found at https://graphscope.io/docs
GraphScope is released under Apache License 2.0. Please note that third-party libraries may not have the same license as GraphScope.
Any contributions you make are greatly appreciated!
- Join in the Slack channel for discussion.
- Please report bugs by submitting a GitHub issue.
- Please submit contributions using pull requests.