PipelineX
Pipeline for eXperimentation
Overview
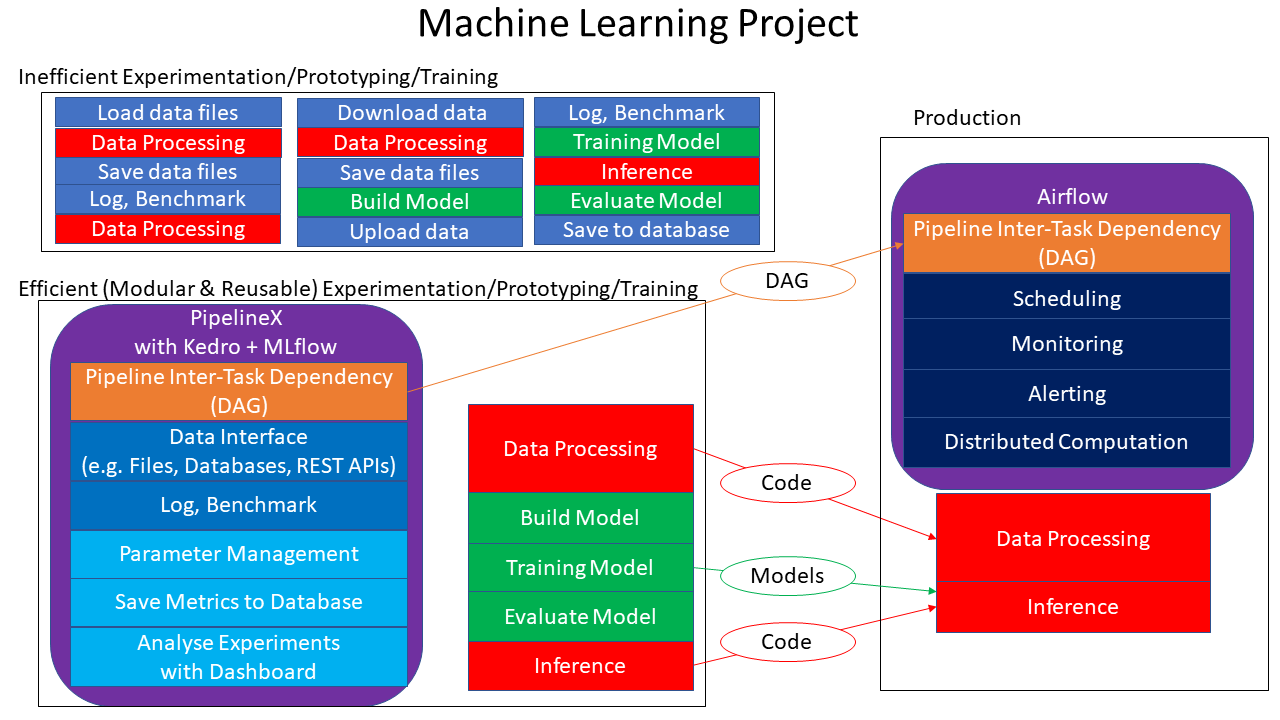
PipelineX is a Python package designed to make Machine Learning projects efficient with modular, reusable, and easy-to-use features for experimentation.
Please refer here to find out how PipelineX differs from other pipeline/workflow packages: Airflow, Luigi, Gokart, Metaflow, and Kedro.
PipelineX provides enhancements for YAML/JSON useful for parameter management summarized as follows.
- Import-less Python object: Include (nested) Python classes and functions in a YAML/JSON file
- Anchor-less aliasing: Look up another key in the same YAML/JSON file
- Python expression in YAML/JSON files
PipelineX provides enhancements for Kedro summarized as follows.
- Kedro pipeline/DAG definition in a YAML/JSON file with more options
- Additional Kedro-compatible data interface sets ("DataSets") for Computer Vision applications
- Additional decorators for benchmarking
- Integration with MLflow that enables to save metrics to a database supported by SQLAlchemy (SQLite, PostgreSQL, etc.)
PipelineX includes integration with the following Python packages.
- Kedro
- MLflow
- PyTorch
- Ignite
- Pandas
- OpenCV
- Memory Profiler
- Python bindings to the NVIDIA Management Library
- Shap
These wrappers are all independent and optional. You do not need to install all of these Python packages.
PipelineX shares similar philosophy/concepts with:
-
Pipeline/workflow packages: Apache Beam, Argo, Kubeflow, Apache Airflow, Luigi, Gokart, Metaflow
-
Parameter/Config management packages: Hydra, Jsonnet, Helm, ytt
PipelineX shares similar API styles with
-
Pipeline/workflow package: Kubeflow
-
Domain-specific packages: Allennlp, Ludwig, Detectron2
Installation
- [Option 1] To install the latest release from the PyPI:
$ pip install pipelinex- [Option 2] To install the latest pre-release:
$ pip install git+https://github.com/Minyus/pipelinex.git- [Option 3] To install the latest pre-release without need to reinstall even after modifying the source code:
$ git clone https://github.com/Minyus/pipelinex.git
$ cd pipelinex
$ python setup.py developExample/Demo Projects
-
Entry example using Scikit-learn to demonstrate PipelineX's Kedro-MLflow integration
-
parameters.ymlat conf/base/parameters.yml -
Essential packages: Scikit-learn, pandas, Kedro, MLflow
-
Application: Kaggle's exercise competition to predict which Titanic's passengers survived
-
Data: Kaggle's Titanic
-
Model: Logistic Regression
-
-
Entry example using Scikit-learn to demonstrate more PipelineX's options
- Adds more PipelineX's options, such as declaring Python objects in YAML, to the previous example.
-
-
parameters.ymlat conf/base/parameters.yml -
Essential packages: PyTorch, Ignite, Shap, Kedro, MLflow
-
Application: Image classification
-
Data: MNIST images
-
Model: CNN (Convolutional Neural Network)
-
Loss: Cross-entropy
-
-
Kaggle competition using PyTorch
-
parameters.ymlat kaggle/conf/base/parameters.yml -
Essential packages: PyTorch, Ignite, pandas, numpy, Kedro, MLflow
-
Application: Kaggle competition to predict the results of American Football plays
-
Data: Sparse heatmap-like field images and tabular data
-
Model: Combination of CNN and MLP
-
Loss: Continuous Rank Probability Score (CRPS)
-
-
parameters.ymlat conf/base/parameters.yml- Essential packages: OpenCV, Scikit-image, numpy, TensorFlow (pretrained model), Kedro, MLflow
- Application: Image processing to estimate the empty area ratio of cuboid container on a truck
- Data: container images
-
Uplift Modeling using CausalLift
parameters.ymlat conf/base/parameters.yml- Essential packages: CausalLift, Scikit-learn, XGBoost, pandas, Kedro
- Application: Uplift Modeling to find which customers should be targeted and which customers should not for a marketing campaign (treatment)
- Data: generated by simulation
Template repositories
The following 2 template repositories for PipelineX are available.
-
Template repository to use YAML less than Kedro: likely preferable for first-time users.
-
Template repository to use YAML more than Kedro: potentially preferable for users who are not satisfied with Kedro as is.
These were simplified versions of the template project created by kedro new command which uses Cookiecutter.
To use for a new project, fork the template repository and hit Use this template button next to Clone or download.
Pythonic enhanced YAML/JSON (HatchDict)
Import-less Python object (class and function)
YAML is a common text format used for application config files.
YAML's most notable advantage is allowing users to mix 2 styles, block style and flow style.
Example:
import yaml
from pprint import pprint # pretty-print for clearer look
# Read parameters dict from a YAML file in actual use
params_yaml="""
block_style_demo:
key1: value1
key2: value2
flow_style_demo: {key1: value1, key2: value2}
"""
parameters = yaml.safe_load(params_yaml)
print("### 2 styles in YAML ###")
pprint(parameters)### 2 styles in YAML ###
{'block_style_demo': {'key1': 'value1', 'key2': 'value2'},
'flow_style_demo': {'key1': 'value1', 'key2': 'value2'}}
To store highly nested (hierarchical) dict or list, YAML is more conveinient than hard-coding in Python code.
-
YAML's block style, which uses indentation, allows users to omit opening and closing symbols to specify a Python dict or list (
{}or[]). -
YAML's flow style, which uses opening and closing symbols, allows users to specify a Python dict or list within a single line.
So simply using YAML with Python will be the best way for Machine Learning experimentation?
Let's check out the next example.
Example:
import yaml
from pprint import pprint # pretty-print for clearer look
# Read parameters dict from a YAML file in actual use
params_yaml = """
model_kind: LogisticRegression
model_params:
C: 1.23456
max_iter: 987
random_state: 42
"""
parameters = yaml.safe_load(params_yaml)
print("### Before ###")
pprint(parameters)
model_kind = parameters.get("model_kind")
model_params_dict = parameters.get("model_params")
if model_kind == "LogisticRegression":
from sklearn.linear_model import LogisticRegression
model = LogisticRegression(**model_params_dict)
elif model_kind == "DecisionTree":
from sklearn.tree import DecisionTreeClassifier
model = DecisionTreeClassifier(**model_params_dict)
elif model_kind == "RandomForest":
from sklearn.ensemble import RandomForestClassifier
model = RandomForestClassifier(**model_params_dict)
else:
raise ValueError("Unsupported model_kind.")
print("\n### After ###")
print(model)### Before ###
{'model_kind': 'LogisticRegression',
'model_params': {'C': 1.23456, 'max_iter': 987, 'random_state': 42}}
### After ###
LogisticRegression(C=1.23456, class_weight=None, dual=False, fit_intercept=True,
intercept_scaling=1, l1_ratio=None, max_iter=987,
multi_class='warn', n_jobs=None, penalty='l2',
random_state=42, solver='warn', tol=0.0001, verbose=0,
warm_start=False)
This way is inefficient as we need to add import and if statements for the options in the Python code in addition to modifying the YAML config file.
Any better way?
PyYAML provides UnsafeLoader which can load Python objects without import.
import yaml
# You do not need `import sklearn.linear_model` using PyYAML's UnsafeLoader
# Read parameters dict from a YAML file in actual use
params_yaml = """
model:
!!python/object:sklearn.linear_model.LogisticRegression
C: 1.23456
max_iter: 987
random_state: 42
"""
parameters = yaml.unsafe_load(params_yaml) # unsafe_load required
model = parameters.get("model")
print("### model object by PyYAML's UnsafeLoader ###")
print(model)### model object by PyYAML's UnsafeLoader ###
LogisticRegression(C=1.23456, class_weight=None, dual=None, fit_intercept=None,
intercept_scaling=None, l1_ratio=None, max_iter=987,
multi_class=None, n_jobs=None, penalty=None, random_state=42,
solver=None, tol=None, verbose=None, warm_start=None)
PyYAML's !!python/object and !!python/name, however, has the following problems.
!!python/objector!!python/nameare too long to write.- Positional (non-keyword) arguments are apparently not supported.
Any better way?
PipelineX provides the solution.
PipelineX's HatchDict provides an easier syntax, as follows, to convert Python dictionaries read from YAML or JSON files to Python objects without import.
- Use
=key to specify the package, module, and class/function with.separator infoo_package.bar_module.baz_classformat. - [Optional] Use
_key to specify (list of) positional arguments (args) if any. - [Optional] Add keyword arguments (kwargs) if any.
To return an object instance like PyYAML's !!python/object, feed positional and/or keyword arguments. If there is no arguments, just feed null (known as None in Python) to _ key.
To return an uninstantiated (raw) object like PyYAML's !!python/name, just feed = key without positional nor keyword arugments.
Example:
from pipelinex import HatchDict
import yaml
from pprint import pprint # pretty-print for clearer look
# You do not need `import sklearn.linear_model` using PipelineX's HatchDict
# Read parameters dict from a YAML file in actual use
params_yaml="""
model:
=: sklearn.linear_model.LogisticRegression
C: 1.23456
max_iter: 987
random_state: 42
"""
parameters = yaml.safe_load(params_yaml)
model_dict = parameters.get("model")
print("### Before ###")
pprint(model_dict)
model = HatchDict(parameters).get("model")
print("\n### After ###")
print(model)### Before ###
{'=': 'sklearn.linear_model.LogisticRegression',
'C': 1.23456,
'max_iter': 987,
'random_state': 42}
### After ###
LogisticRegression(C=1.23456, class_weight=None, dual=False, fit_intercept=True,
intercept_scaling=1, l1_ratio=None, max_iter=987,
multi_class='warn', n_jobs=None, penalty='l2',
random_state=42, solver='warn', tol=0.0001, verbose=0,
warm_start=False)
This import-less Python object supports nested objects (objects that receives object arguments) by recursive depth-first search.
For more examples, please see Use with PyTorch and parameters.yml in example/demo projects .
This import-less Python object feature, inspired by the fact that Kedro uses load_obj for file I/O (DataSet), uses load_obj copied from kedro.utils which dynamically imports Python objects using importlib, a Python standard library.
Anchor-less aliasing (self-lookup)
To avoid repeating, YAML natively provides Anchor&Alias Anchor&Alias feature, and Jsonnet provides Variable feature to JSON.
Example:
import yaml
from pprint import pprint # pretty-print for clearer look
# Read parameters dict from a YAML file in actual use
params_yaml="""
train_params:
train_batch_size: &batch_size 32
val_batch_size: *batch_size
"""
parameters = yaml.safe_load(params_yaml)
train_params_dict = parameters.get("train_params")
print("### Conversion by YAML's Anchor&Alias feature ###")
pprint(train_params_dict)### Conversion by YAML's Anchor&Alias feature ###
{'train_batch_size': 32, 'val_batch_size': 32}
Unfortunately, YAML and Jsonnet require a medium to share the same value.
This is why PipelineX provides Anchor-less aliasing feature.
You can directly look up another value in the same YAML/JSON file using $ key without an anchor nor variable.
To specify the nested key (key in a dict of dict), use . as the separator.
Example:
from pipelinex import HatchDict
import yaml
from pprint import pprint # pretty-print for clearer look
# Read parameters dict from a YAML file in actual use
params_yaml="""
train_params:
train_batch_size: 32
val_batch_size: {$: train_params.train_batch_size}
"""
parameters = yaml.safe_load(params_yaml)
train_params_dict = parameters.get("train_params")
print("### Before ###")
pprint(train_params_dict)
train_params = HatchDict(parameters).get("train_params")
print("\n### After ###")
pprint(train_params)### Before ###
{'train_batch_size': 32,
'val_batch_size': {'$': 'train_params.train_batch_size'}}
### After ###
{'train_batch_size': 32, 'val_batch_size': 32}
Python expression
Strings wrapped in parentheses are evaluated as a Python expression.
from pipelinex import HatchDict
import yaml
from pprint import pprint # pretty-print for clearer look
# Read parameters dict from a YAML file in actual use
params_yaml = """
train_params:
param1_tuple_python: (1, 2, 3)
param1_tuple_yaml: !!python/tuple [1, 2, 3]
param2_formula_python: (2 + 3)
param3_neg_inf_python: (float("-Inf"))
param3_neg_inf_yaml: -.Inf
param4_float_1e9_python: (1e9)
param4_float_1e9_yaml: 1.0e+09
param5_int_1e9_python: (int(1e9))
"""
parameters = yaml.load(params_yaml)
train_params_raw = parameters.get("train_params")
print("### Before ###")
pprint(train_params_raw)
train_params_converted = HatchDict(parameters).get("train_params")
print("\n### After ###")
pprint(train_params_converted)### Before ###
{'param1_tuple_python': '(1, 2, 3)',
'param1_tuple_yaml': (1, 2, 3),
'param2_formula_python': '(2 + 3)',
'param3_neg_inf_python': '(float("-Inf"))',
'param3_neg_inf_yaml': -inf,
'param4_float_1e9_python': '(1e9)',
'param4_float_1e9_yaml': 1000000000.0,
'param5_int_1e9_python': '(int(1e9))'}
### After ###
{'param1_tuple_python': (1, 2, 3),
'param1_tuple_yaml': (1, 2, 3),
'param2_formula_python': 5,
'param3_neg_inf_python': -inf,
'param3_neg_inf_yaml': -inf,
'param4_float_1e9_python': 1000000000.0,
'param4_float_1e9_yaml': 1000000000.0,
'param5_int_1e9_python': 1000000000}
The unified data interface framework
Machine Learning projects involves with loading and saving various data in various ways such as:
- files in local/network file system, Hadoop File System (HDFS), Amazon S3, Google Cloud Storage
- e.g. CSV, JSON, YAML, pickle, images, models, etc.
- databases
- Postgresql, MySQL etc.
- Spark
- REST API (HTTP(S) requests)
It is often the case that many Machine Learning Engineers code both data loading/saving and data transformation mixed in the same Python module or Jupyter notebook during experimentation/prototyping phase and suffer later on because:
- During experimentation/prototyping, we often want to save the intermediate data after each transformation.
- In production environments, we often want to skip saving data to minimize latency and storage space.
- To benchmark the performance or troubleshoot, we often want to switch the data source.
- e.g. read image files in local storage or download images through REST API
The proposed solution is the unified data interface.
Here is a simple demo example to predict survival on the Titanic.
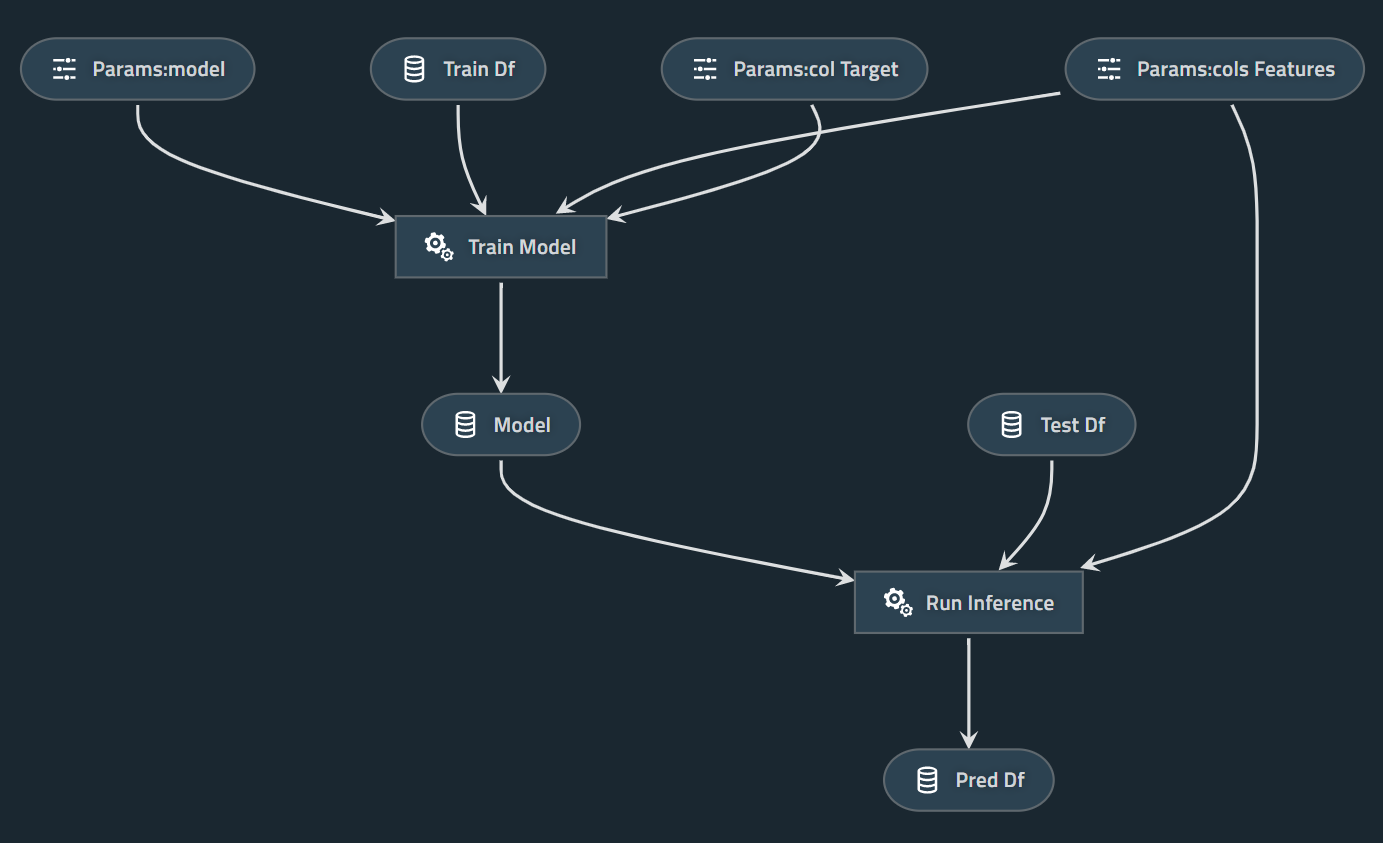
Common code to define the tasks/operations/transformations:
# Define tasks
def train_model(model, df, cols_features, col_target):
# train a model here
return model
def run_inference(model, df, cols_features):
# run inference here
return dfIt is notable that you do not need to add any Kedro-related code here to use Kedro later on.
Furthermore, you do not need to add any MLflow-related code here to use MLflow later on as Kedro hooks provided by PipelineX can handle behind the scenes.
This advantage enables you to keep your pipelines for experimentation/prototyping/benchmarking production-ready.
- Plain code:
# Configure: can be written in a config file (YAML, JSON, etc.)
train_data_filepath = "data/input/train.csv"
train_data_load_args = {"float_precision": "high"}
test_data_filepath = "data/input/test.csv"
test_data_load_args = {"float_precision": "high"}
pred_data_filepath = "data/load/pred.csv"
pred_data_save_args = {"index": False, "float_format": "%.16e"}
model_kind = "LogisticRegression"
model_params_dict = {
"C": 1.23456
"max_iter": 987
"random_state": 42
}
# Run tasks
import pandas as pd
if model_kind == "LogisticRegression":
from sklearn.linear_model import LogisticRegression
model = LogisticRegression(**model_params_dict)
train_df = pd.read_csv(train_data_filepath, **train_data_load_args)
model = train_model(model, train_df)
test_df = pd.read_csv(test_data_filepath, **test_data_load_args)
pred_df = run_inference(model, test_df)
pred_df.to_csv(pred_data_filepath, **pred_data_save_args)- Following the data interface framework, objects with
_load, and_savemethods, proposed by Kedro and supported by PipelineX:
# Define a data interface: better ones such as "CSVDataSet" are provided by Kedro
import pandas as pd
from pathlib import Path
class CSVDataSet:
def __init__(self, filepath, load_args={}, save_args={}):
self._filepath = filepath
self._load_args = {}
self._load_args.update(load_args)
self._save_args = {"index": False}
self._save_args.update(save_args)
def _load(self) -> pd.DataFrame:
return pd.read_csv(self._filepath, **self._load_args)
def _save(self, data: pd.DataFrame) -> None:
save_path = Path(self._filepath)
save_path.parent.mkdir(parents=True, exist_ok=True)
data.to_csv(str(save_path), **self._save_args)
# Configure data interface: can be written in catalog config file using Kedro
train_dataset = CSVDataSet(
filepath="data/input/train.csv",
load_args={"float_precision": "high"},
# save_args={"float_format": "%.16e"}, # You can set save_args for future use
)
test_dataset = CSVDataSet(
filepath="data/input/test.csv",
load_args={"float_precision": "high"},
# save_args={"float_format": "%.16e"}, # You can set save_args for future use
)
pred_dataset = CSVDataSet(
filepath="data/load/pred.csv",
# load_args={"float_precision": "high"}, # You can set load_args for future use
save_args={"float_format": "%.16e"},
)
model_kind = "LogisticRegression"
model_params_dict = {
"C": 1.23456
"max_iter": 987
"random_state": 42
}
cols_features = [
"Pclass", # The passenger's ticket class
"Parch", # # of parents / children aboard the Titanic
]
col_target = "Survived" # Column used as the target: whether the passenger survived or not
# Run tasks: can be configured as a pipeline using Kedro
# and can be written in parameters config file using PipelineX
if model_kind == "LogisticRegression":
from sklearn.linear_model import LogisticRegression
model = LogisticRegression(**model_params_dict)
train_df = train_dataset._load()
model = train_model(model, train_df, cols_features, col_target)
test_df = test_dataset._load()
pred_df = run_inference(model, test_df, cols_features)
pred_dataset._save(pred_df)Just following the data interface framework might be somewhat beneficial in the long run, but not enough.
Let's see what Kedro and PipelineX can do.
Kedro
Kedro is a Python package to develop pipelines consisting of:
-
data interface sets (data loading/saving wrappers, called "DataSets", that follows the unified data interface framework) such as:
pandas.CSVDataSet: a CSV file in local or cloud (Amazon S3, Google Cloud Storage) utilizing filesystem_spec (fsspec)pickle.PickleDataSet: a pickle file in local or cloud (Amazon S3, Google Cloud Storage) utilizing filesystem_spec (fsspec)pandas.SQLTableDataSet: a table data in an SQL database supported by SQLAlchemy- data interface sets for Spark, Google BigQuery, Feather, HDF, Parquet, Matplotlib, NetworkX, Excel, and more provided by Kedro
- Custom data interface sets provided by Kedro users
-
tasks/operations/transformations (called "Nodes") provided by Kedro users such as:
- data pre-processing
- training a model
- inference using a model
-
inter-task dependency provided by Kedro users
Kedro pipelines can be run sequentially or in parallel.
Regarding Kedro, please see:
- Kedro's document
- YouTube playlist: Writing Data Pipelines with Kedro
- Python Packages for Pipeline/Workflow
Here is a simple example Kedro project.
# catalog.yml
train_df:
type: pandas.CSVDataSet # short for kedro.extras.datasets.pandas.CSVDataSet
filepath: data/input/train.csv
load_args:
float_precision: high
# save_args: # You can set save_args for future use
# float_format": "%.16e"
test_df:
type: pandas.CSVDataSet # short for kedro.extras.datasets.pandas.CSVDataSet
filepath: data/input/test.csv
load_args:
float_precision: high
# save_args: # You can set save_args for future use
# float_format": "%.16e"
pred_df:
type: pandas.CSVDataSet # short for kedro.extras.datasets.pandas.CSVDataSet
filepath: data/load/pred.csv
# load_args: # You can set load_args for future use
# float_precision: high
save_args:
float_format: "%.16e"# parameters.yml
model:
!!python/object:sklearn.linear_model.LogisticRegression
C: 1.23456
max_iter: 987
random_state: 42
cols_features: # Columns used as features in the Titanic data table
- Pclass # The passenger's ticket class
- Parch # # of parents / children aboard the Titanic
col_target: Survived # Column used as the target: whether the passenger survived or not# pipeline.py
from kedro.pipeline import Pipeline, node
from my_module import train_model, run_inference
def create_pipeline(**kwargs):
return Pipeline(
[
node(
func=train_model,
inputs=["params:model", "train_df", "params:cols_features", "params:col_target"],
outputs="model",
),
node(
func=run_inference,
inputs=["model", "test_df", "params:cols_features"],
outputs="pred_df",
),
]
)# run.py
from kedro.runner import SequntialRunner
# Set up ProjectContext here
context = ProjectContext()
context.run(pipeline_name="__default__", runner=SequentialRunner())Kedro pipelines can be visualized using kedro-viz.
Kedro pipelines can be productionized using:
- kedro-airflow: converts a Kedro pipeline into Airflow Python operators.
- kedro-docker: builds a Docker image that can run a Kedro pipeline
- kedro-argo: converts a Kedro pipeline into an Argo (backend of Kubeflow) pipeline
Supplements for Kedro
pipelinex.extras provides Kedro hooks, data interface sets, and decorators to supplement kedro.extras as follows.
Integration with MLflow by Kedro hooks (callbacks)
pipelinex.extras.hooks provides Kedro hooks (callbacks) to use MLflow without adding any MLflow-related code in the node (task) functions.
-
pipelinex.MLflowBasicLoggerHook: Configures and logs duration time for the pipeline to MLflow with args:- enable_mlflow: Enable configuring and logging to MLflow.
- uri:
uriarg fed to: https://www.mlflow.org/docs/latest/python_api/mlflow.html#mlflow.set_tracking_uri as the MLflow tracking server URI. Local file path, databases supported by SQLAlchemy (sqlite, mysql, mssql, and postgresql), HTTP server, Databricks workspace are supported. See MLflow's document at: https://mlflow.org/docs/latest/tracking.html#where-runs-are-recorded - experiment_name:
namearg fed to: https://www.mlflow.org/docs/latest/python_api/mlflow.html#mlflow.create_experiment as the MLflow experiment name. - artifact_location:
artifact_locationarg fed to: https://www.mlflow.org/docs/latest/python_api/mlflow.html#mlflow.create_experiment as the URI to store the artifacts. Local file paths, Amazon S3, Azure Blob Storage, Google Cloud Storage, SFTP server, NFS, and HDFS are supported. See MLflow's document at: https://mlflow.org/docs/latest/tracking.html#id10 - run_name: Shown as 'Run Name' in the MLflow UI.
- offset_hours: The offset hour (e.g. 0 for UTC+00:00) used for
__time_beginand__time_endparameters.
-
pipelinex.MLflowArtifactsLoggerHook: Logs artifacts of specified file paths and dataset names to MLflow with args:- enable_mlflow: Enable logging to MLflow.
- filepaths_before_pipeline_run: The file paths of artifacts to log before the pipeline is run.
- datasets_after_node_run: The dataset names to log after the node is run.
- filepaths_after_pipeline_run: The file paths of artifacts to log after the pipeline is run.
-
pipelinex.MLflowDataSetsLoggerHook: Logs datasets of (list of) float/int and str classes to MLflow with arg:- enable_mlflow: Enable logging to MLflow.
-
pipelinex.MLflowTimeLoggerHook: Logs duration time for each node (task) to MLflow and optionally visualizes the execution logs as a Gantt chart byplotly.figure_factory.create_ganttifplotlyis installed, with args:- enable_mlflow: Enable logging to MLflow.
- enable_plotly: Enable visualization of logged time as a gantt chart.
- gantt_filepath: File path to save the generated gantt chart.
- gantt_params: Args fed to: https://plotly.github.io/plotly.py-docs/generated/plotly.figure_factory.create_gantt.html
- metric_name_prefix: Prefix for the metric names. The metric names are
metric_name_prefixconcatenated with the string returned bytask_name_func.
- task_name_func: Callable to return the task name using
kedro.pipeline.node.Node- object.
-
pipelinex.AddTransformersHook: Adds Kedro transformers such as:pipelinex.MLflowIOTimeLoggerTransformer: Logs duration time to load and save each dataset with args:- enable_mlflow: Enable logging to MLflow.
- metric_name_prefix: Prefix for the metric names. The metric names are
metric_name_prefixconcatenated with 'load <data_set_name>' or 'save <data_set_name>'
To use these hooks, please see example projects at kedro_mlflow or pipelinex_sklearn
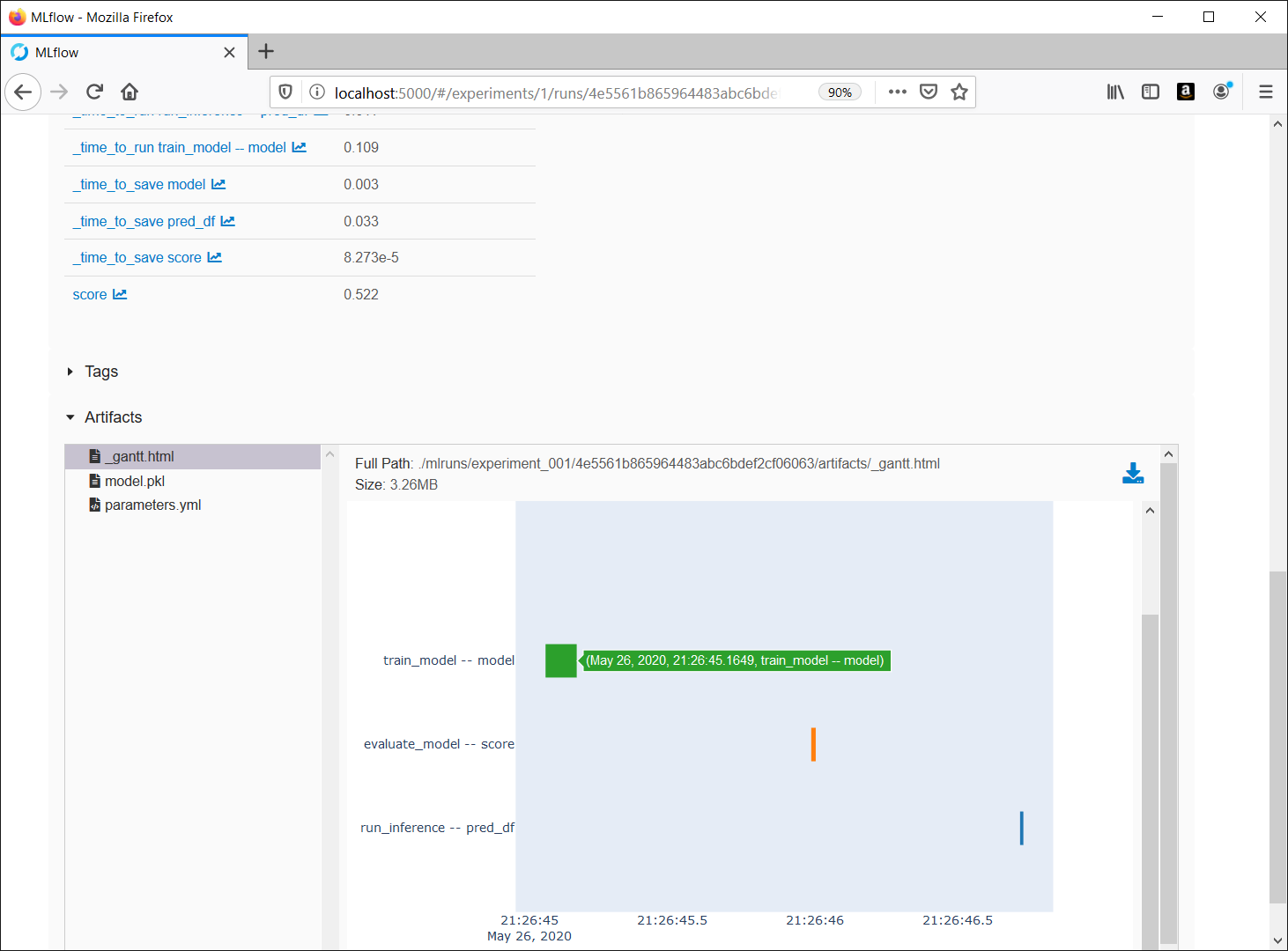 Experiment logs in MLflow's UI
Experiment logs in MLflow's UI
Additional Kedro data interface sets
pipelinex.extras.datasets provides the following data interface sets mainly for Computer Vision applications using OpenCV, Scikit-image, PyTorch/torchvision, and TensorFlow/Keras.
-
- loads/saves multiple numpy arrays (RGB, BGR, or monochrome image) from/to a folder in local storage using
pillowpackage, working likekedro.extras.datasets.pillow.ImageDataSetandkedro.io.PartitionedDataSetwith conversion between numpy arrays and Pillow images. - an example project is at pipelinex_image_processing
- loads/saves multiple numpy arrays (RGB, BGR, or monochrome image) from/to a folder in local storage using
-
- downloads multiple contents (such as images and json) by HTTP requests using
requestspackage - an example project is at pipelinex_image_processing
- downloads multiple contents (such as images and json) by HTTP requests using
-
- downloads multiple contents (such as images and json) by asynchronous HTTP requests using
httpxpackage - an example project is at pipelinex_image_processing
- downloads multiple contents (such as images and json) by asynchronous HTTP requests using
-
pipelinex.IterableImagesDataSet
- wrapper of
torchvision.datasets.ImageFolderthat loads images in a folder as an iterable data loader to use with PyTorch.
- wrapper of
-
pipelinex.PandasProfilingDataSet
- generates a pandas dataframe summary report using pandas-profiling
-
more data interface sets for pandas dataframe summarization/visualization provided by PipelineX
Additional function decorators for benchmarking
pipelinex.extras.decorators provides Python decorators for benchmarking.
-
- logs the duration time of a function (difference of timestamp before and after running the function).
- Slightly modified version of Kedro's log_time
-
- logs the peak memory usage during running the function.
memory_profilerneeds to be installed.- Slightly modified version of Kedro's mem_profile
-
- logs the difference of NVIDIA GPU usage before and after running the function.
pynvmlorpy3nvmlneeds to be installed.
from pipelinex import log_time
from pipelinex import mem_profile # Need to install memory_profiler for memory profiling
from pipelinex import nvml_profile # Need to install pynvml for NVIDIA GPU profiling
from time import sleep
import logging
logging.basicConfig(level=logging.INFO)
@nvml_profile
@mem_profile
@log_time
def foo_func(i=1):
sleep(0.5) # Needed to avoid the bug reported at https://github.com/pythonprofilers/memory_profiler/issues/216
return "a" * i
output = foo_func(100_000_000)INFO:pipelinex.decorators.decorators:Running 'foo_func' took 549ms [0.549s]
INFO:pipelinex.decorators.memory_profiler:Running 'foo_func' consumed 579.02MiB memory at peak time
INFO:pipelinex.decorators.nvml_profiler:Ran: 'foo_func', NVML returned: {'_Driver_Version': '418.67', '_NVML_Version': '10.418.67', 'Device_Count': 1, 'Devices': [{'_Name': 'Tesla P100-PCIE-16GB', 'Total_Memory': 17071734784, 'Free_Memory': 17071669248, 'Used_Memory': 65536, 'GPU_Utilization_Rate': 0, 'Memory_Utilization_Rate': 0}]}, Used memory diff: [0]
Enhanced Kedro context: YAML interface for Kedro pipelines
PipelineX enables you to use Kedro in more convenient ways. Using pipelinex.FlexibleContext, you can define the inter-task dependency (DAG) for Kedro pipelines in YAML.
Here are the options configurable in parameters.yml:
HatchDict features
# parameters.yml
model:
=: sklearn.linear_model.LogisticRegression
C: 1.23456
max_iter: 987
random_state: 42
cols_features: # Columns used as features in the Titanic data table
- Pclass # The passenger's ticket class
- Parch # # of parents / children aboard the Titanic
col_target: Survived # Column used as the target: whether the passenger survived or notDefine Kedro pipelines using PIPELINES key
- Optionally specify the default Python module (path of .py file) if you want to omit the module name
- Optionally specify the Python function decorator to apply to each node
- Specify
inputs,func, andoutputsfor each node- For sub-pipelines consisting of nodes of only single input and single output, you can optionally use Sequential API similar to PyTorch (
torch.nn.Sequential) and Keras (tf.keras.Sequential)
- For sub-pipelines consisting of nodes of only single input and single output, you can optionally use Sequential API similar to PyTorch (
# parameters.yml
PIPELINES:
__default__:
=: pipelinex.FlexiblePipeline
module: # Optionally specify the default Python module so you can omit the module name to which functions belongs
decorator: # Optionally specify function decorator(s) to apply to each node
nodes:
- inputs: ["params:model", train_df, "params:cols_features", "params:col_target"]
func: sklearn_demo.train_model
outputs: model
- inputs: [model, test_df, "params:cols_features"]
func: sklearn_demo.run_inference
outputs: pred_dfConfigure Kedro run config using RUN_CONFIG key
- Optionally run nodes in parallel
- Optionally run only missing nodes (skip tasks which have already been run to resume pipeline using the intermediate data files or databases.) Note: You can use Kedro CLI to overwrite these run configs.
# parameters.yml
RUN_CONFIG:
pipeline_name: __default__
runner: SequentialRunner # Set to "ParallelRunner" to run in parallel
only_missing: False # Set True to run only missing nodes
tags: # None
node_names: # None
from_nodes: # None
to_nodes: # None
from_inputs: # None
load_versions: # NoneDefine Kedro hooks using HOOKS key
# parameters.yml
HOOKS:
- =: pipelinex.MLflowBasicLoggerHook # Configure and log duration time for the pipeline
enable_mlflow: True # Enable configuring and logging to MLflow
uri: sqlite:///mlruns/sqlite.db
experiment_name: experiment_001
artifact_location: ./mlruns/experiment_001
offset_hours: 0 # Specify the offset hour (e.g. 0 for UTC/GMT +00:00) to log in MLflow
- =: pipelinex.MLflowArtifactsLoggerHook # Log artifacts of specified file paths and dataset names
enable_mlflow: True # Enable logging to MLflow
filepaths_before_pipeline_run: # Optionally specify the file paths to log before pipeline is run
- conf/base/parameters.yml
datasets_after_node_run: # Optionally specify the dataset names to log after the node is run
- model
filepaths_after_pipeline_run: # None # Optionally specify the file paths to log after pipeline is run
- =: pipelinex.MLflowOutputsLoggerHook # Log output datasets of (list of) float, int, and str classes
enable_mlflow: True # Enable logging to MLflow
- =: pipelinex.MLflowTimeLoggerHook # Log duration time to run each node (task)
enable_mlflow: True # Enable logging to MLflow
- =: pipelinex.AddTransformersHook # Add transformers
transformers:
=: pipelinex.MLflowIOTimeLoggerTransformer # Log duration time to load and save each dataset
enable_mlflow: TrueHere are the options configurable in catalog.yml:
HatchDictfeatures available- Optionally enable caching using
cachedkey set to True if you do not want Kedro to load the data from disk/database which were in the memory. (kedro.io.CachedDataSetis used under the hood.)
The complete example project is available here.
Use with PyTorch
To develop a simple neural network, it is convenient to use Sequential API
(e.g. torch.nn.Sequential, tf.keras.Sequential).
- Hardcoded:
from torch.nn import Sequential, Conv2d, ReLU
model = Sequential(
Conv2d(in_channels=3, out_channels=16, kernel_size=[3, 3]),
ReLU(),
)
print("### model object by hard-coding ###")
print(model)### model object by hard-coding ###
Sequential(
(0): Conv2d(3, 16, kernel_size=[3, 3], stride=(1, 1))
(1): ReLU()
)
- Using import-less Python object feature:
from pipelinex import HatchDict
import yaml
from pprint import pprint # pretty-print for clearer look
# Read parameters dict from a YAML file in actual use
params_yaml="""
model:
=: torch.nn.Sequential
_:
- {=: torch.nn.Conv2d, in_channels: 3, out_channels: 16, kernel_size: [3, 3]}
- {=: torch.nn.ReLU, _: }
"""
parameters = yaml.safe_load(params_yaml)
model_dict = parameters.get("model")
print("### Before ###")
pprint(model_dict)
model = HatchDict(parameters).get("model")
print("\n### After ###")
print(model)### Before ###
{'=': 'torch.nn.Sequential',
'_': [{'=': 'torch.nn.Conv2d',
'in_channels': 3,
'kernel_size': [3, 3],
'out_channels': 16},
{'=': 'torch.nn.ReLU', '_': None}]}
### After ###
Sequential(
(0): Conv2d(3, 16, kernel_size=[3, 3], stride=(1, 1))
(1): ReLU()
)
In addition to Sequential, TensorFLow/Keras provides modules to merge branches such as
tf.keras.layers.Concatenate, but PyTorch provides only functional interface such as torch.cat.
PipelineX provides modules to merge branches such as ModuleConcat, ModuleSum, and ModuleAvg.
- Hardcoded:
from torch.nn import Sequential, Conv2d, AvgPool2d, ReLU
from pipelinex import ModuleConcat
model = Sequential(
ModuleConcat(
Conv2d(in_channels=3, out_channels=16, kernel_size=[3, 3], stride=[2, 2], padding=[1, 1]),
AvgPool2d(kernel_size=[3, 3], stride=[2, 2], padding=[1, 1]),
),
ReLU(),
)
print("### model object by hard-coding ###")
print(model)### model object by hard-coding ###
Sequential(
(0): ModuleConcat(
(0): Conv2d(3, 16, kernel_size=[3, 3], stride=[2, 2], padding=[1, 1])
(1): AvgPool2d(kernel_size=[3, 3], stride=[2, 2], padding=[1, 1])
)
(1): ReLU()
)
- Using import-less Python object feature:
from pipelinex import HatchDict
import yaml
from pprint import pprint # pretty-print for clearer look
# Read parameters dict from a YAML file in actual use
params_yaml="""
model:
=: torch.nn.Sequential
_:
- =: pipelinex.ModuleConcat
_:
- {=: torch.nn.Conv2d, in_channels: 3, out_channels: 16, kernel_size: [3, 3], stride: [2, 2], padding: [1, 1]}
- {=: torch.nn.AvgPool2d, kernel_size: [3, 3], stride: [2, 2], padding: [1, 1]}
- {=: torch.nn.ReLU, _: }
"""
parameters = yaml.safe_load(params_yaml)
model_dict = parameters.get("model")
print("### Before ###")
pprint(model_dict)
model = HatchDict(parameters).get("model")
print("\n### After ###")
print(model)### Before ###
{'=': 'torch.nn.Sequential',
'_': [{'=': 'pipelinex.ModuleConcat',
'_': [{'=': 'torch.nn.Conv2d',
'in_channels': 3,
'kernel_size': [3, 3],
'out_channels': 16,
'padding': [1, 1],
'stride': [2, 2]},
{'=': 'torch.nn.AvgPool2d',
'kernel_size': [3, 3],
'padding': [1, 1],
'stride': [2, 2]}]},
{'=': 'torch.nn.ReLU', '_': None}]}
### After ###
Sequential(
(0): ModuleConcat(
(0): Conv2d(3, 16, kernel_size=[3, 3], stride=[2, 2], padding=[1, 1])
(1): AvgPool2d(kernel_size=[3, 3], stride=[2, 2], padding=[1, 1])
)
(1): ReLU()
)
Use with PyTorch Ignite
Wrappers of PyTorch Ignite provides most of features available in Ignite, including integration with MLflow, in an easy declarative way.
In addition, the following optional features are available in PipelineX.
- Use only partial samples in dataset (Useful for quick preliminary check before using the whole dataset)
- Time limit for training (Useful for code-only (Kernel-only) Kaggle competitions with time limit)
Here are the arguments for NetworkTrain:
loss_fn (callable): Loss function used to train.
Accepts an instance of loss functions at https://pytorch.org/docs/stable/nn.html#loss-functions
epochs (int, optional): Max epochs to train
seed (int, optional): Random seed for training.
optimizer (torch.optim, optional): Optimizer used to train.
Accepts optimizers at https://pytorch.org/docs/stable/optim.html
optimizer_params (dict, optional): Parameters for optimizer.
train_data_loader_params (dict, optional): Parameters for data loader for training.
Accepts args at https://pytorch.org/docs/stable/data.html#torch.utils.data.DataLoader
val_data_loader_params (dict, optional): Parameters for data loader for validation.
Accepts args at https://pytorch.org/docs/stable/data.html#torch.utils.data.DataLoader
evaluation_metrics (dict, optional): Metrics to compute for evaluation.
Accepts dict of metrics at https://pytorch.org/ignite/metrics.html
evaluate_train_data (str, optional): When to compute evaluation_metrics using training dataset.
Accepts events at https://pytorch.org/ignite/engine.html#ignite.engine.Events
evaluate_val_data (str, optional): When to compute evaluation_metrics using validation dataset.
Accepts events at https://pytorch.org/ignite/engine.html#ignite.engine.Events
progress_update (bool, optional): Whether to show progress bar using tqdm package
scheduler (ignite.contrib.handle.param_scheduler.ParamScheduler, optional): Param scheduler.
Accepts a ParamScheduler at
https://pytorch.org/ignite/contrib/handlers.html#module-ignite.contrib.handlers.param_scheduler
scheduler_params (dict, optional): Parameters for scheduler
model_checkpoint (ignite.handlers.ModelCheckpoint, optional): Model Checkpoint.
Accepts a ModelCheckpoint at https://pytorch.org/ignite/handlers.html#ignite.handlers.ModelCheckpoint
model_checkpoint_params (dict, optional): Parameters for ModelCheckpoint at
https://pytorch.org/ignite/handlers.html#ignite.handlers.ModelCheckpoint
early_stopping_params (dict, optional): Parameters for EarlyStopping at
https://pytorch.org/ignite/handlers.html#ignite.handlers.EarlyStopping
time_limit (int, optioinal): Time limit for training in seconds.
train_dataset_size_limit (int, optional): If specified, only the subset of training dataset is used.
Useful for quick preliminary check before using the whole dataset.
val_dataset_size_limit (int, optional): If specified, only the subset of validation dataset is used.
useful for qucik preliminary check before using the whole dataset.
cudnn_deterministic (bool, optional): Value for torch.backends.cudnn.deterministic.
See https://pytorch.org/docs/stable/notes/randomness.html for details.
cudnn_benchmark (bool, optional): Value for torch.backends.cudnn.benchmark.
See https://pytorch.org/docs/stable/notes/randomness.html for details.
mlflow_logging (bool, optional): If True and MLflow is installed, MLflow logging is enabled.
Please see the example code using MNIST dataset prepared based on the original code.
It is also possible to use:
- FlexibleModelCheckpoint handler which enables to use timestamp in the model checkpoint file name to clarify which one is the latest.
- CohenKappaScore metric which can compute Quadratic Weighted Kappa Metric used in some Kaggle competitions. See sklearn.metrics.cohen_kappa_score for details.
It is planned to port some code used with PyTorch Ignite to PyTorch Ignite repository once test and example codes are prepared.
Use with OpenCV
A challenge of image processing is that the parameters and algorithms that work with an image often do not work with another image. You will want to output intermediate images from each image processing pipeline step for visual check during development, but you will not want to output all the intermediate images to save time and disk space in production.
Wrappers of OpenCV and ImagesLocalDataSet are the solution. You can concentrate on developping your image processing pipeline for an image (3-D or 2-D numpy array), and it will run for all the images in a folder.
If you are devepping an image processing pipeline consisting of 5 steps and you have 10 images, for example, you can check 10 generated images in each of 5 folders, 50 images in total, during development.
Use with PyTorch Lightning
(To-do)
Use with TensorFlow/Keras
(To Do.)
Use with Docker container
Build Docker image
# docker build -t pipelinex:3.7.7-slim -f dockerfiles/dockerfile .Use with Docker container
# docker run -it --name pipelinex pipelinex:3.7.7-slim /bin/bashWhy and how PipelineX was born
When I was working on a Deep Learning project, it was very time-consuming to develop the pipeline for experimentation. I wanted 2 features.
First one was an option to resume the pipeline using the intermediate data files instead of running the whole pipeline. This was important for rapid Machine/Deep Learning experimentation.
Second one was modularity, which means keeping the 3 components, task processing, file/database access, and DAG definition, independent. This was important for efficient software engineering.
After this project, I explored for a long-term solution. I researched about 3 Python packages for pipeline development, Airflow, Luigi, and Kedro, but none of these could be a solution.
Luigi provided resuming feature, but did not offer modularity. Kedro offered modularity, but did not provide resuming feature.
After this research, I decided to develop my own package that works on top of Kedro. Besides, I added syntactic sugars including Sequential API similar to Keras and PyTorch to define DAG. Furthermore, I added integration with MLflow, PyTorch, Ignite, pandas, OpenCV, etc. while working on more Machine/Deep Learning projects.
After I confirmed my package worked well with the Kaggle competition, I released it as PipelineX.
Contributors wanted!
Please see CONTRIBUTING.md for details.
Author
Yusuke Minami