The Biomaterials Annotator: a system for ontology-based concept annotation of biomaterials text.
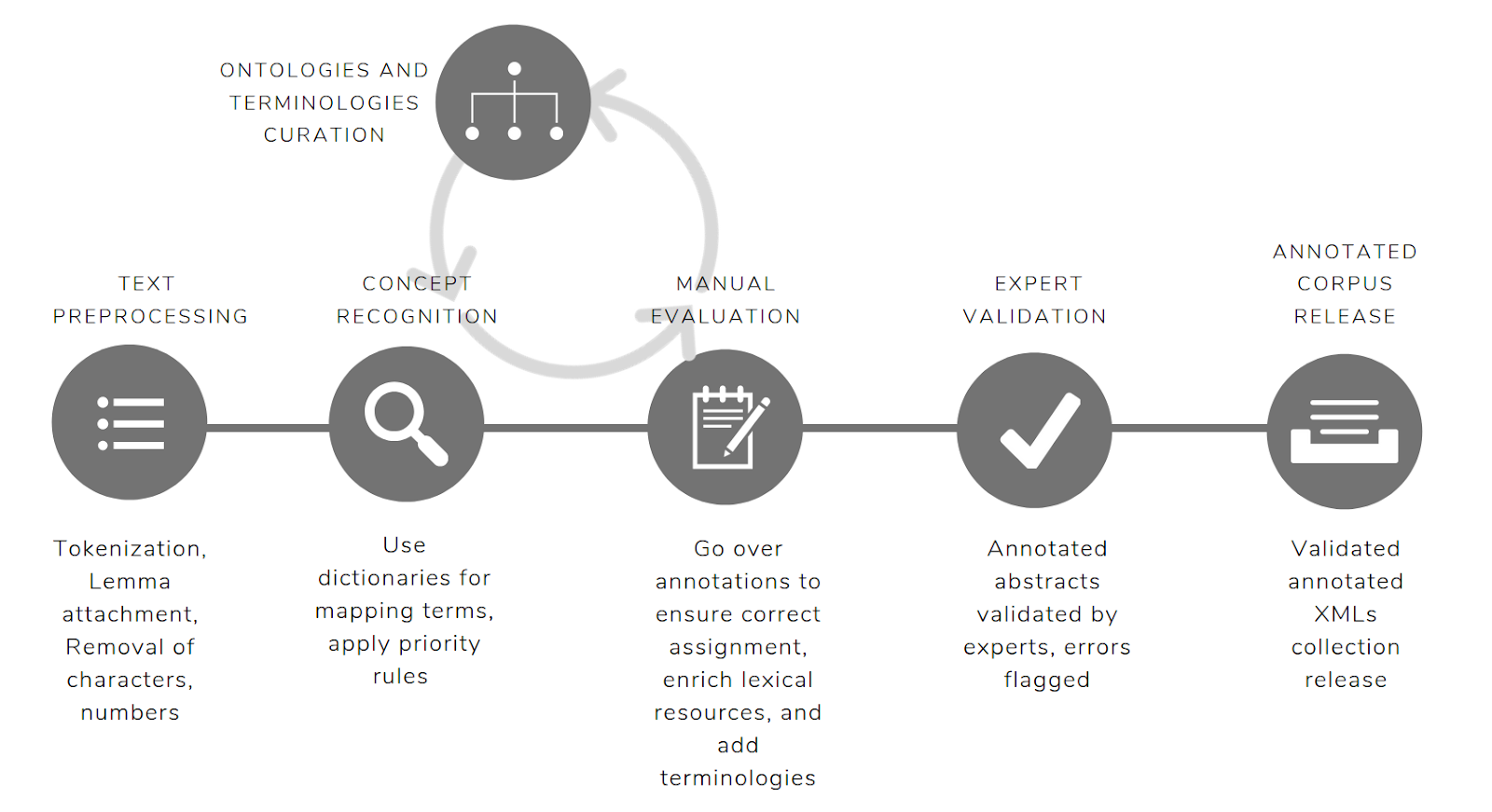
The Biomaterials Annotator is an ontology-based NER system that identifies biomaterial concepts. It provides a schema for combining terms from mutiple ontologies, vocabularies and nomenclutures. A full list of the type of concepts annotated are available here.
The global scores calculated for the system are: 0.75 strict F-score, 0.79 lenient F-Score and 0.77 average F-score. The full results including metrics by category are available here. Here are the validated set of abstracts.
The Biomaterials Annotator has been implemented following a modular organization, using software containers for the different components. The pipeline is orchestrated using Nextflow as workflow manager. Natural language processing (NLP) components were mainly developed in Java, and it relying on the Stanford CoreNLP Natural Language Processing open source toolkit.
A biomaterials annotated gold standard corpus of 1222 MEDLINE abstracts resulting from the execution of the Biomaterials Annotator is available and free to use at https://github.com/ProjectDebbie/Biomaterials_annotated_corpus. The corpus contains articles describing the evaluation of biomaterials and medical devices in either a laboratory or clinical setting, Each abstract is individually contained as a separate file under the GATE format.
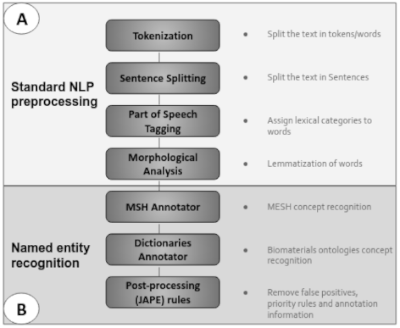
The Standard NLP preprocessing component is available at https://gitlab.bsc.es/inb/text-mining/generic-tools/nlp-standard-preprocessing The MSH Annotator annotates pre-selected categories from the MeSH terminology; and the Dictionary Annotator annotates does the same using manually collected ontologies and vocabularies. This is followed by execution of the Post-processing rules, including entity recognition based on lexical rules, removal of false positives and abbreviations concept recognition, among other tasks.
The MSH Annotator is available at https://github.com/ProjectDebbie/debbie_umls_annotations; and the Dictionary Annotator and Post-processing rules are available at https://github.com/ProjectDebbie/DEBBIE_dictionaries_annotations.
- nlp-standard-preprocessing: registry.hub.docker.com/javicorvi/nlp-standard-preprocessing:dev_1.6
- debbie-umls-annotation: registry.hub.docker.com/projectdebbie/debbie_umls_annotation:release-1.0.7
- debbie-dictionaries-annotations: registry.hub.docker.com/projectdebbie/debbie_dictionaries_annotations:release-2.0.0
- MESH (UMLS)
- DEB
- GMDN
- CHEBI
- IOBC
- NCIT
- NPO
- OBI
- ONTOTOXNUC
- UBERON
- PREMEDONTO
- EDAM Bioimaging Ontology
- CHMO
You need to have docker and nextflow installed, then configure and the run.sh file.
We use SemVer for versioning. For the versions available, see the tags on this repository.
Javier Corvi and Osnat Hakimi
Corvi, J., Fuenteslópez, C., Fernández, J., Gelpi, J., Ginebra, M.-P., Capella-Guitierrez, S., Hakimi, O.: The biomaterials annotator: a systemfor ontology-based concept annotation of biomaterials text. In:Proceedings of the Second Workshop on Scholarly DocumentProcessing, pp. 36–48. Association for Computational Linguistics,Online (2021). https://www.aclweb.org/anthology/2021.sdp-1.5
This project is licensed under the GNU License - see the LICENSE file for details
 This project has received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Sklodowska-Curie grant agreement No 751277
This project has received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Sklodowska-Curie grant agreement No 751277