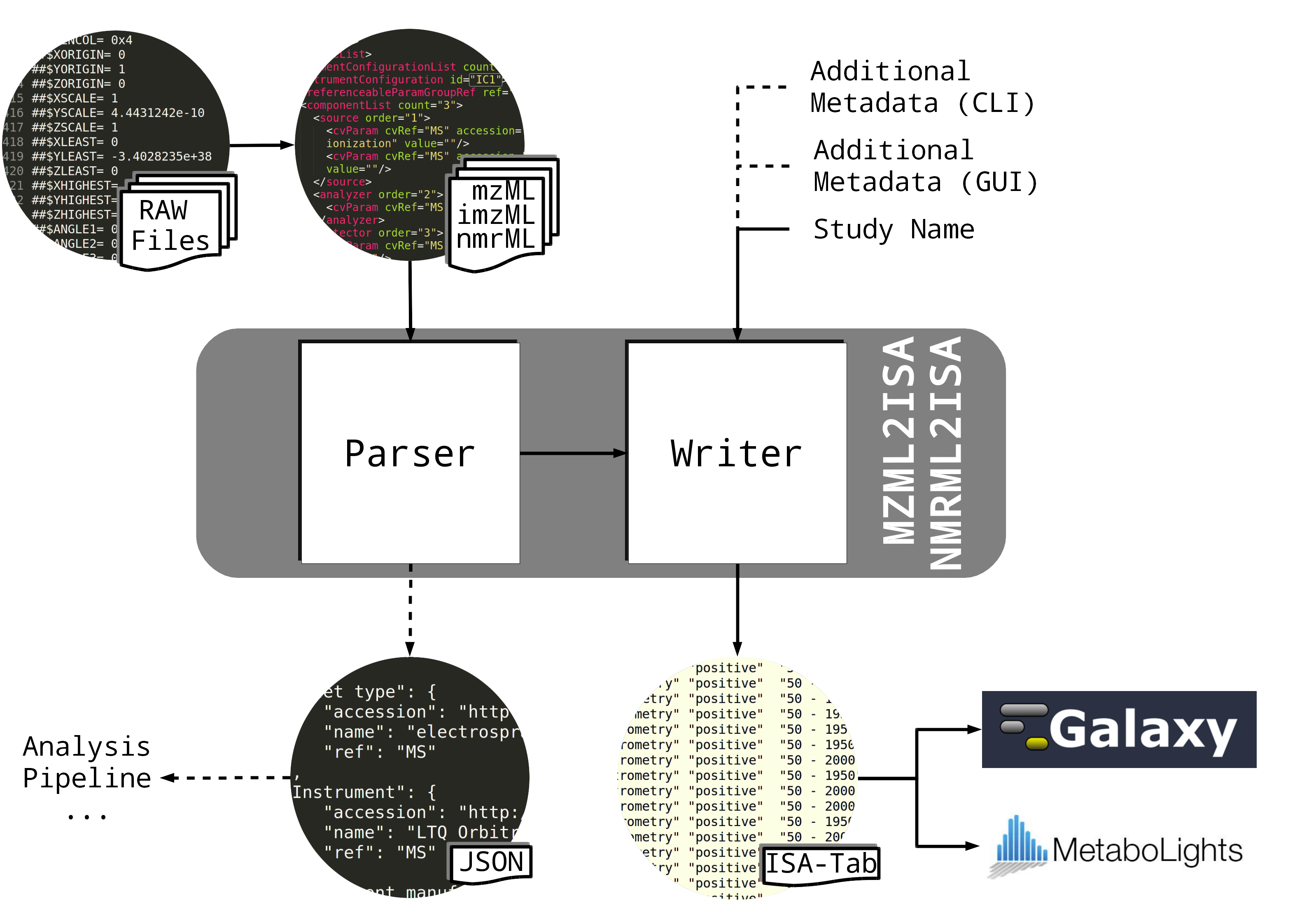
Parser to extract meta information from mzML mass spectrometry files and parse relevant information to a ISA-Tab structure
mzml2isa is a Python3 program that can automatically generate ISA-Tab document structure metadata files from raw XML metabolomics data files (mzML open access data format). The mzml2ISA tool provides the backbone of ISA-Tab metabolomics study which can then be edited with an ISA editing tool, ISAcreator (see MetaboLights pre-packaged ISA Creator)
- Currently the program does the following:
- Extract meta information from mzML files and stores it as either python dictionary or JSON format
- Creates an ISA-Tab file structure with relevant meta information filled in.
- Add additional metadata that cannot be parsed from mzML files to the ISA-Tab files through a JSON formatted dictionnary.
See the Installation page of the online documentation.
mzml2isa -i /path/to/mzml_files/ -o /path/to/out_folder/ -s name_of_studySee the Usage page and the Examples page for more information.
To download some real data from MetaboLights studies to test the converter with, run
python scripts/metabolights-dl.py <size>from inside the repository, where size is the maximum size in GiB you
can allocate to download files. The script will download the files to
the example_files/metabolights folder and then run mzml2isa on
those files..
If you use a *NIX machine with curlftpfs and bash available, you can also run
scripts/metabolights.shto mount the database to the example directory and start converting mzML studies.