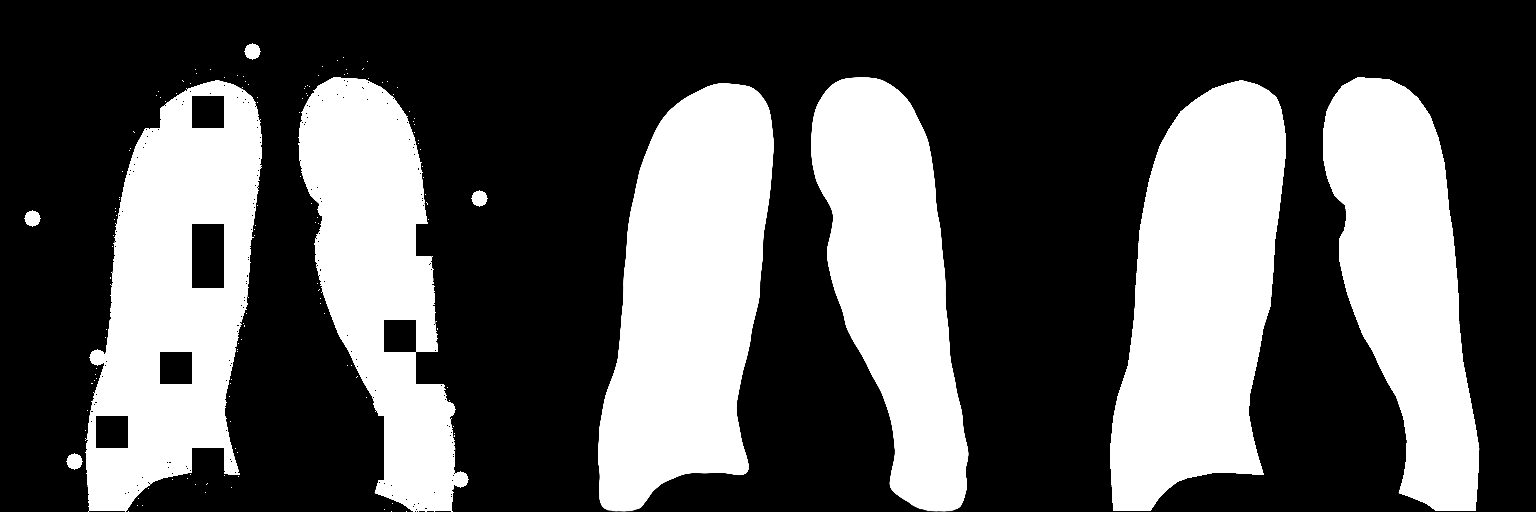
This is an unofficial PyTorch implementation of Anatomical Priors for Image Segmentation via Post-processing with Denoising Autoencoders
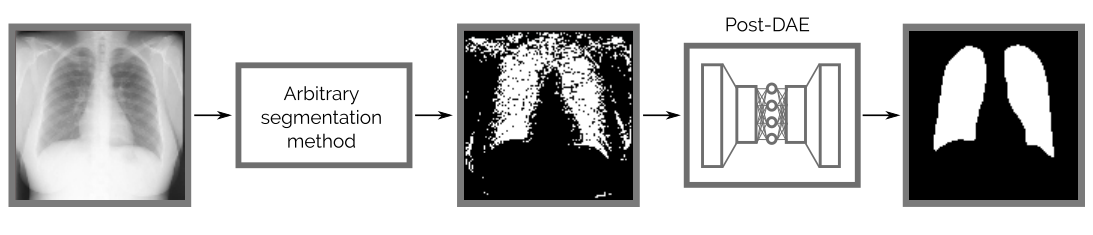 Post-DAE works as a post-processing step and improves the anatomical plausibility of segmentation masks obtained with arbitrary methods.
Post-DAE works as a post-processing step and improves the anatomical plausibility of segmentation masks obtained with arbitrary methods.
- We verify, that DAE can be used as an independent post-processing step to correct problematic and non-anatomically plausible masks produced by arbitrary segmentation methods.
- This is a method that can be trained using segmentation-only datasets or anatomical masks coming from arbitrary image modalities, since the DAE is trained using only segmentation masks, and no intensity information is required during learning. [1]
- We validate Post-DAE in the context of lung segmentation in X-ray images, showing its robustness by improving segmentation masks
- Clone this repo using :
git clone https://github.com/balazsborsos/dae_postprocessing
- Install the requirements using :
pip install -r requirements.txt
- To start model training run the main.py file with following arguments :
python train -t "PATH_TO_CONFIG" -i "PATH_TO_MASKS"
An example config yml can be found in the utils folder. The input is just the path to the masks of the dataset, corruptions take place in the training script.
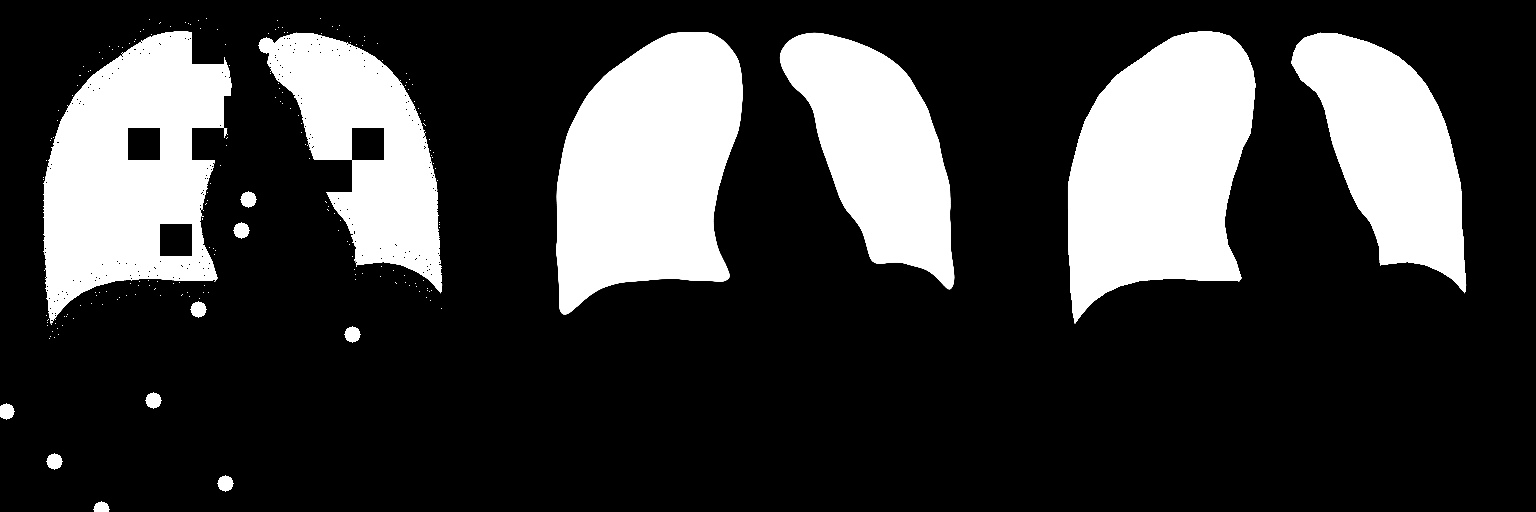
Input Image Predicted Ground Truth
- Chest Xray Masks and Labels [2] dataset was used for training.
- It freely accessible on Kaggle.
- The dataset was compiled from papers [3] and [4]
The arvix version of the paper can found at the following link.
If you find this repo useful please cite the original authors :
@inproceedings{larrazabal2019anatomical,
title={Anatomical priors for image segmentation via post-processing with denoising autoencoders},
author={Larrazabal, Agostina J and Martinez, Cesar and Ferrante, Enzo},
booktitle={Medical Image Computing and Computer Assisted Intervention--MICCAI 2019: 22nd International Conference, Shenzhen, China, October 13--17, 2019, Proceedings, Part VI 22},
pages={585--593},
year={2019},
organization={Springer}
}