Begin by loading the pre-processed data:
load('./data/ST_heart_spot_data.RData')Lets see what it contains:
names(heart)## [1] "atlas" "genes"
There are two data frames one with the RNAseq data and one with the atlas data. Every row is a spot and the two data frames are matched so heart$atlas[123,] is the same spot as heart$genes[123,].
head(heart$atlas)## x y intensity area id color color2 acronym
## 537 17044.09 14752.31 0 31934 17 #0B61A4 <NA> LV
## 550 15557.39 14725.60 0 30502 17 #0B61A4 #41b6c4 LV
## 556 16338.52 14749.41 0 36820 17 #0B61A4 #41b6c4 LV
## 565 17027.89 14057.10 0 32274 17 #0B61A4 #41b6c4 LV
## 566 14857.41 14019.45 0 20654 17 #0B61A4 #41b6c4 LV
## 570 19129.29 14026.82 0 33556 19 #FF4900 <NA> RV
## name right.left rostral.caudal anterior.posterior spot.id
## 537 left ventricle 171.67656 486.1625 192 537
## 550 left ventricle 87.43070 447.6606 192 550
## 556 left ventricle 132.92846 471.6618 192 556
## 565 left ventricle 177.67626 451.1608 192 565
## 566 left ventricle 32.18342 373.6570 192 566
## 570 right ventricle 290.67070 462.4113 192 570
## nuclei image spot.pos
## 537 30 1_CN73_E1_HE 15x16
## 550 34 1_CN73_E1_HE 17x16
## 556 38 1_CN73_E1_HE 16x16
## 565 28 1_CN73_E1_HE 15x17
## 566 20 1_CN73_E1_HE 18x17
## 570 20 1_CN73_E1_HE 12x17
heart$genes[1:10,1:4]## ENSG00000000003 ENSG00000000419 ENSG00000001036 ENSG00000001167
## 15x16 0 0 0 0
## 17x16 0 0 0 0
## 16x16 0 0 1 0
## 15x17 0 0 0 0
## 18x17 0 1 0 0
## 12x17 0 0 1 0
## 14x17 0 0 0 0
## 17x17 0 1 0 0
## 16x17 0 0 0 0
## 13x17 0 0 1 0
So lets explain the variables in heart$atlas:
names(heart$atlas)## [1] "x" "y" "intensity"
## [4] "area" "id" "color"
## [7] "color2" "acronym" "name"
## [10] "right.left" "rostral.caudal" "anterior.posterior"
## [13] "spot.id" "nuclei" "image"
## [16] "spot.pos"
| Variable | Explanation |
|---|---|
x |
X centroid of spot in original pixel image |
y |
Y centroid of spot in original pixel image |
intensity |
not really used here. |
area |
area in pixels of spot. |
id |
unique integer value for the region the spot is in. |
color |
hex color code for the region the spot is in. |
color2 |
Color code corrected for spots that couldn't be assigned |
acronym |
region acronym for where the spot is in. |
name |
region name for where the spot is in. |
right.left |
X coordinate in pixels of the reference atlas |
rostral.caudal |
Y coordinate in pixels of the reference atlas. |
anterior.posterior |
Z oordinate in pixels of the reference atlas |
spot.id |
just 1:now(heart$atlas) |
nuclei |
number of nuclei in spot |
image |
The image the spots came from |
spot.pos |
The array position of the spot in 33x35 |
RGL is a package that uses OpPenGL as backend for 3D visualization in R. misc3d is a package that we will use for drawing scenes in 3d.
library(rgl)
library(misc3d)Load the 3D volume heart atlas.
load('./data/heart.RData')We begin by defining the perspective we want to plot from (this doesn't have to make sense now I'll show later how to get these parameters):
perspective<-list(FOV = 30, ignoreExtent = FALSE, listeners = 1L,
mouseMode = structure(c("trackball", "zoom", "fov", "pull"
), .Names = c("left", "right", "middle", "wheel")), skipRedraw = FALSE,
userMatrix = structure(c(-0.0108720660209656, 0.899227440357208,
0.437346190214157, 0, 0.955604612827301, -0.119448974728584,
0.269354522228241, 0, 0.2944515645504, 0.420858442783356,
-0.858007192611694, 0, 0, 0, 0, 1), .Dim = c(4L, 4L)), scale = c(1,
1, 1), viewport = structure(c(0L, 0L, 1280L, 720L), .Names = c("x",
"y", "width", "height")), zoom = 1, windowRect = c(0L, 45L,
1280L, 765L), family = "sans", font = 1L, cex = 1, useFreeType = TRUE)Lets plot the entire heart and then regions outlines.
#open 3D plot window
open3d(windowRect = c(0, 0, 1280, 720))
#use the perspective
par3d(perspective)
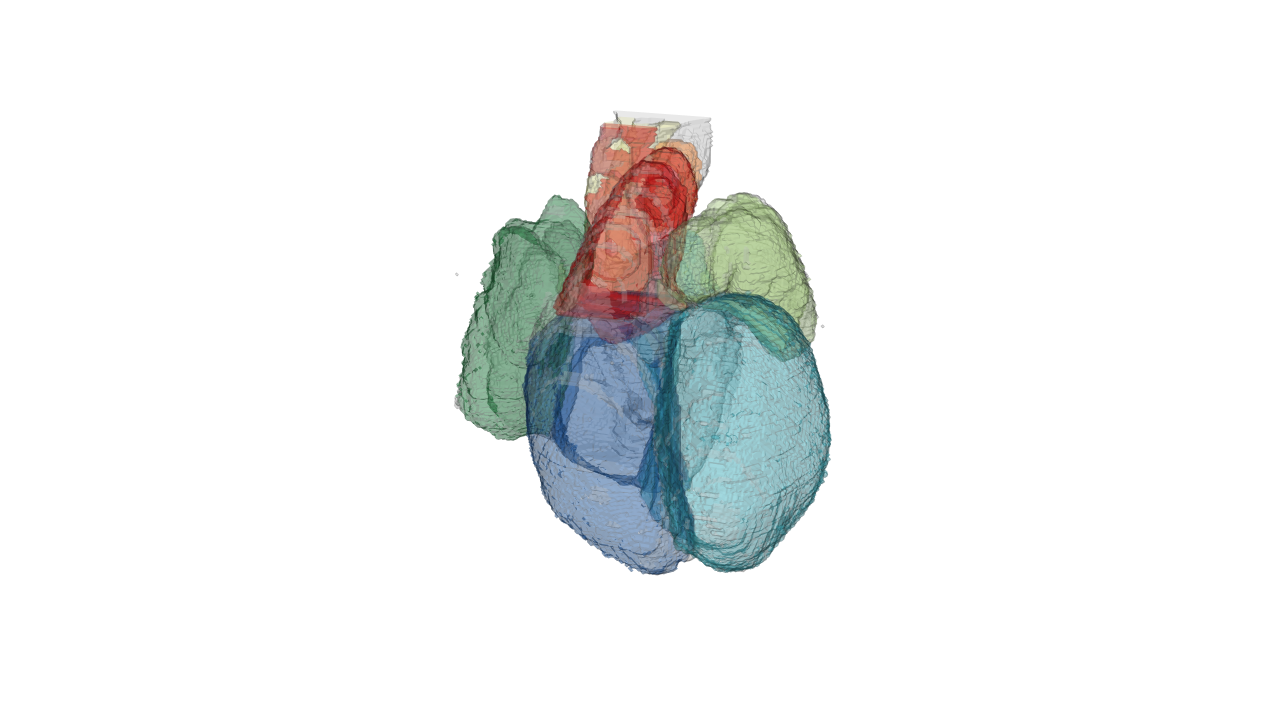
#draw low-resolution heart with color coding
drawScene.rgl(organ.dwnsmp[which(names(organ.dwnsmp)%in%c('WH', 'RA', 'RV', 'LA', 'LV', 'P', 'A', 'OT'))])
rgl.snapshot(file='./images/lowresolution.png')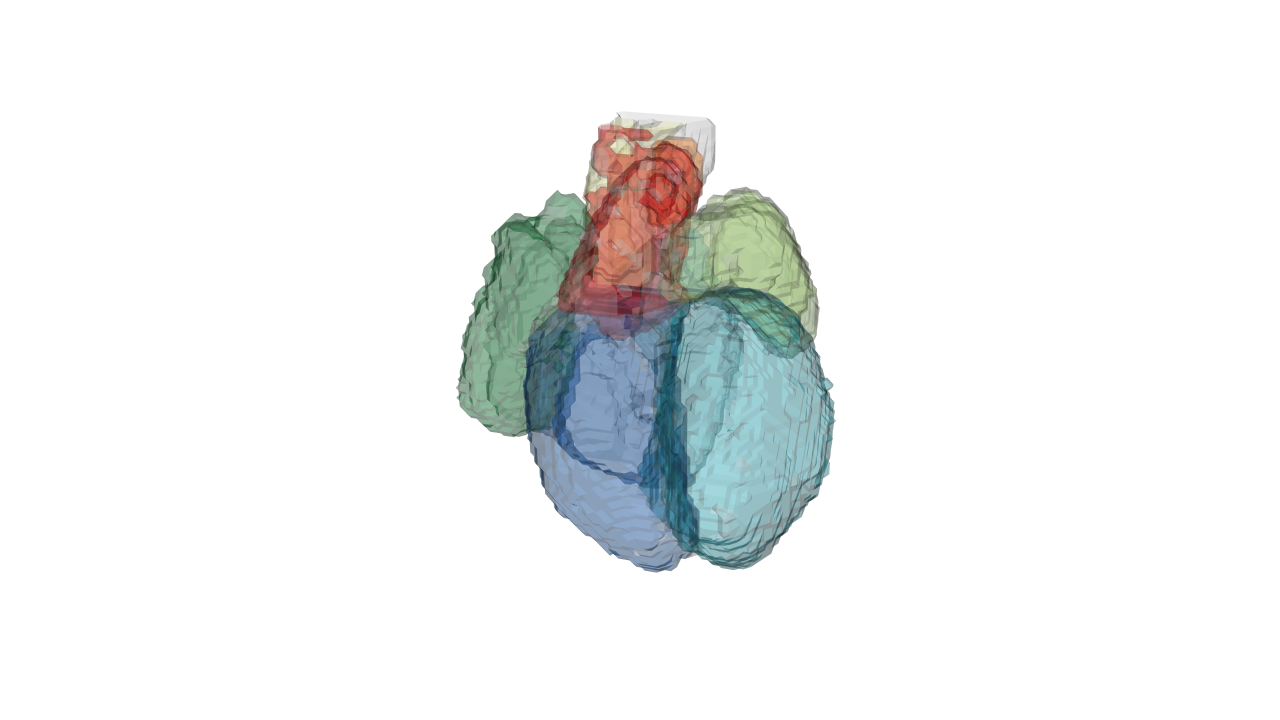 This is a low resolution of the heart that you can rotate and zoom in with the ouse in realtime.
This is a low resolution of the heart that you can rotate and zoom in with the ouse in realtime.
If you have rotated the heart and want to save the perspective parameters into a R object to use later simply run:
pp<- par3d(no.readonly = TRUE)And the parameters are now saved into pp and you can set the perspective in the 3D plot by par3d(pp) before plotting with drawScene.rgl().
For more high resolution rendering you set the position you want of the heart and then rerun the code but not using the organ.dwnsmp but organ instead.
drawScene.rgl(organ[which(names(organ.dwnsmp)%in%c('WH', 'RA', 'RV', 'LA', 'LV', 'P', 'A', 'OT'))])
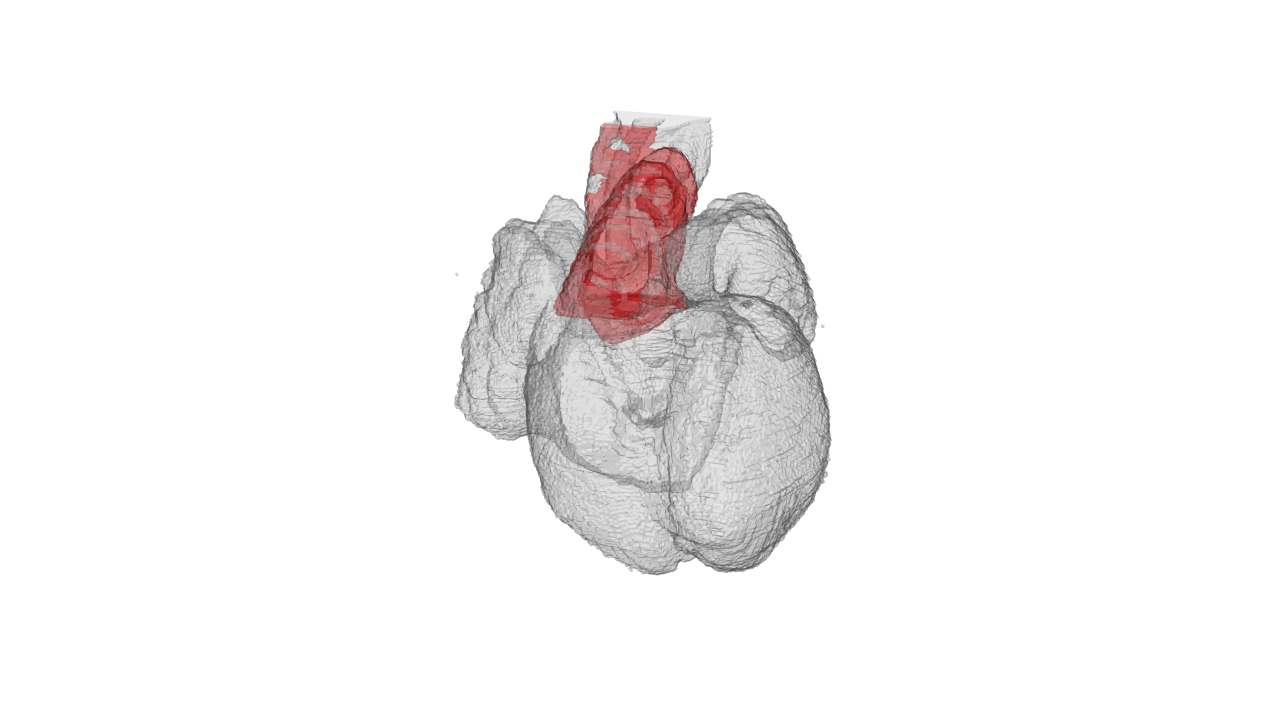
rgl.snapshot(file='./images/highresolution.png')You can select which region to highlight by setting the acronyms in the character vector. here I show WH whole heart as well as OT outflow tract:
drawScene.rgl(organ[which(names(organ.dwnsmp)%in%c('WH','OT'))])
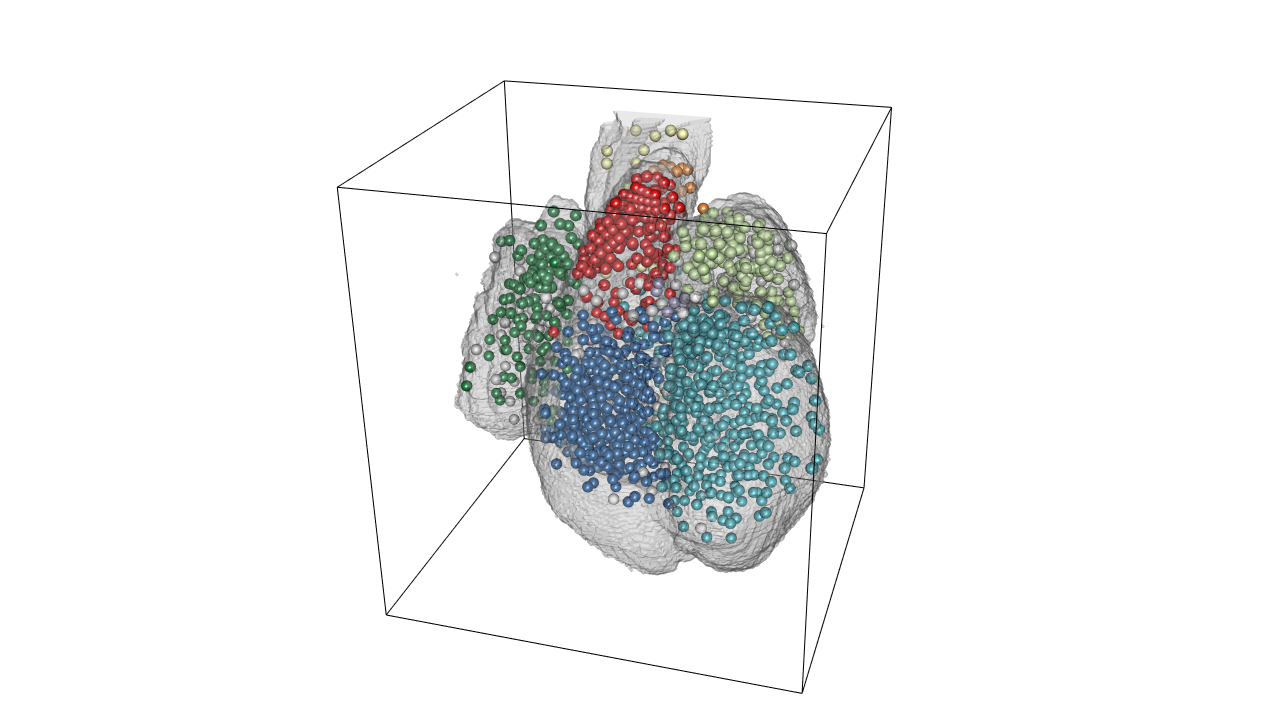
rgl.snapshot(file='./images/OT_highlight.png')Lets plot all the spots with the color according to their anatomical region:
#draw all the spots with region color
drawScene.rgl(organ[which(names(organ.dwnsmp)%in%c('WH'))])
radius.of.spots.in.atlas.pixels<- (100/(2383.36/532))/3
spheres3d(598-heart$atlas$rostral.caudal, 532-heart$atlas$right.left, heart$atlas$anterior.posterior, col=heart$atlas$color2, radius=radius.of.spots.in.atlas.pixels)
#bounding box
box3d()
rgl.snapshot(filename='./images/3d_heart_spots.png')Changeing colors depending on region can easily be done like this:
#create a matching index with regions accoridng to the order you want to change color
matching.index <- match(heart$atlas$acronym, c('RA', 'LA', 'RV', 'LV', 'P', 'A', 'OT', 'epc'))
#overwrite with the color vector
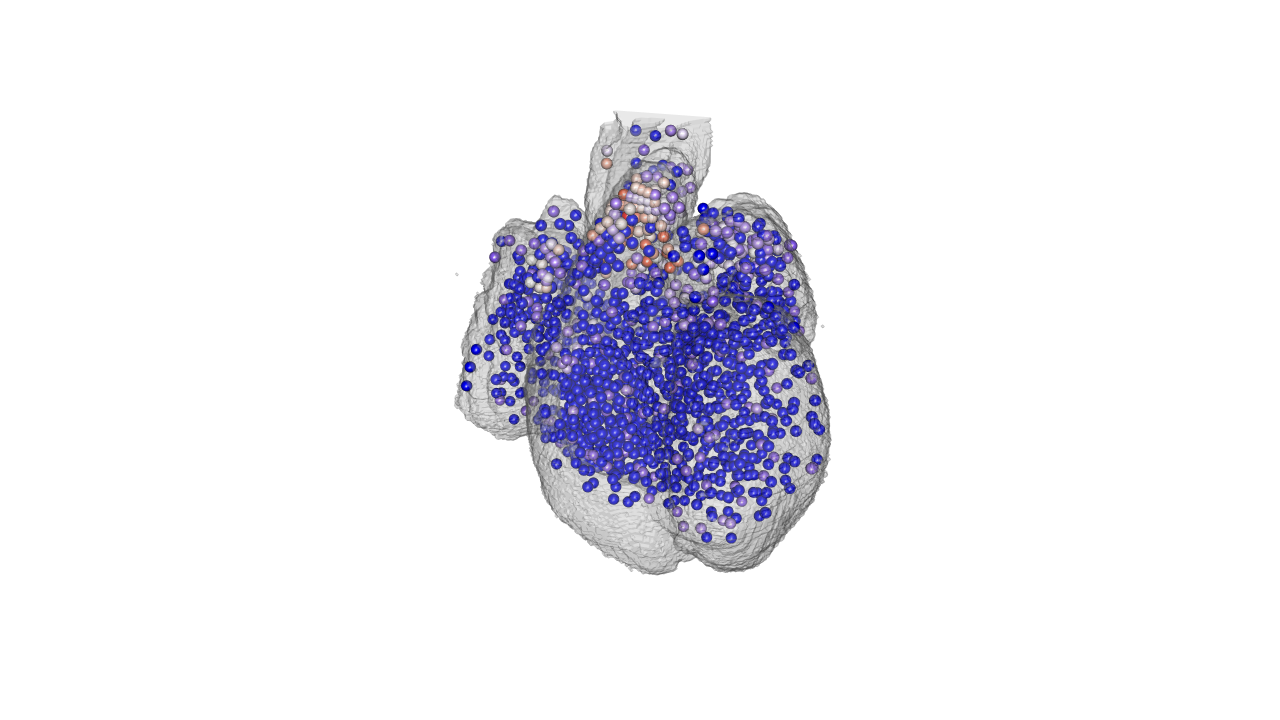
heart$atlas$color2[!is.na(matching.index)] <- c('#c2e699', "#238443", '#41b6c4', '#225ea8', '#ffffb2', "#fd8d3c", '#e31a1c', '#9e9ac8')[na.omit(matching.index)]Finally lets plot the gene expression of specific spots:
#plot gene of interest OGN
gene.of.interest<-heart$genes[,which(colnames(heart$genes)=='ENSG00000106809')]
#color ramp palette base don gene expression
PaletteFunction <- colorRampPalette(c("blue", "white", "red"), space = "Lab")
gene.expression<-PaletteFunction(100)[as.numeric(cut(scale(log2(gene.of.interest+1)), breaks = 100))]
drawScene.rgl(organ[which(names(organ.dwnsmp)%in%c('WH'))])
spheres3d(598-heart$atlas$rostral.caudal, 532-heart$atlas$right.left, heart$atlas$anterior.posterior, col=gene.expression, radius=radius.of.spots.in.atlas.pixels )
rgl.snapshot(filename='./images/OGN_3d_heart.png')