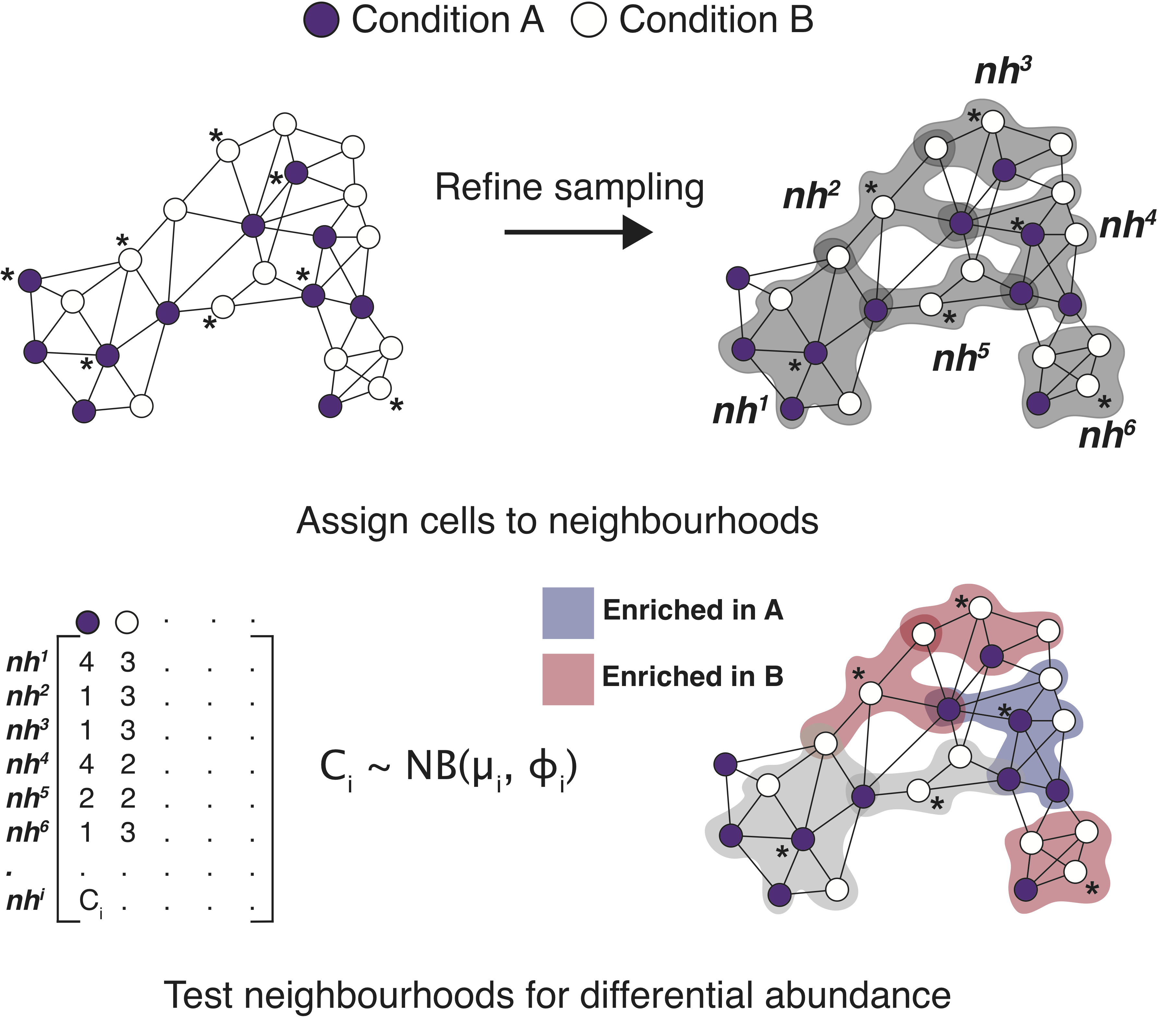
Milo is a method for differential abundance analysis on KNN graph from single-cell datasets. For more details, read our manuscript. If you use Milo in your study, please cite Dann, E., Henderson, N.C., Teichmann, S.A. et al. Differential abundance testing on single-cell data using k-nearest neighbor graphs. Nat Biotechnol (2021).
## Milo is available from Bioconductor (preferred stable installation)
if (!requireNamespace("BiocManager", quietly = TRUE))
install.packages("BiocManager")
BiocManager::install("miloR")
## Install development version
devtools::install_github("MarioniLab/miloR", ref="devel")
- Basic Milo example on simulated dataset
- Milo example on mouse gastrulation dataset: this includes a demo for downstream analysis functions.
- Integrating miloR in scanpy/anndata workflow (see also
milopyfor a full workflow in python) - Specifying contrasts of interest for differential abundance testing with Milo
- Using a mixed effect model for dependendent samples
An example of the Milo work flow to get started:
data(sim_trajectory)
milo.meta <- sim_trajectory$meta
milo.obj <- Milo(sim_trajectory$SCE)
milo.obj
Build a graph and neighbourhoods.
milo.obj <- buildGraph(milo.obj, k=20, d=30)
milo.obj <- makeNhoods(milo.obj, k=20, d=30, refined=TRUE, prop=0.2)
Calculate distances, count cells according to an experimental design and perform DA testing.
milo.obj <- calcNhoodDistance(milo.obj, d=30)
milo.obj <- countCells(milo.obj, samples="Sample", meta.data=milo.meta)
milo.design <- as.data.frame(xtabs(~ Condition + Sample, data=milo.meta))
milo.design <- milo.design[milo.design$Freq > 0, ]
rownames(milo.design) <- milo.design$Sample
milo.design <- milo.design[colnames(nhoodCounts(milo.obj)),]
milo.res <- testNhoods(milo.obj, design=~Condition, design.df=milo.design)
head(milo.res)
For any question, feature request or bug report please create a new issue in this repository. If you have an error or code-based query, please provide
the executed code and the preceding code from the point of constructing the Milo object, along with the output of your sessionInfo() - this will help
us immeasurably to diagnose the issue.
We welcome contributions and suggestions from the community (though we may not take them onboard if they don't align with our development roadmap - please don't be offended). Please submit the initial idea as an issue, which we will discuss and ask for refinements/clarifications. If we approve the idea, then please open a pull request onto the devel branch, from which we will begin a review process. To smooth the process, please note that code changes must be backwards compatible, and must include all relevant unit tests.
Milo is under (semi)continuous development, but it has also been around for a couple of years. That means (hopefully) most of the common bugs have been
identified. If you have an error or issue to raise, first make sure you are using the most up to date version, either via Bioconductor, or from the Main
branch of the repo. Please also make sure to check if your problem has arisen before by looking at both the open and closed issues. If neither of these
solve your problem, then please open a new issue describing the problem in full, with code and a minimally reproducible example. Don't forget to include
the output of your sessionInfo() as well. Thanks.