| Author | Project | Build Status | License | Code Quality | Coverage |
|---|---|---|---|---|---|
| G. Filitto | MRI colorectal cancer segmentation | Linux :  |
 |
Codacy : Codebeat : |
This package allows to segment cancer regions on T2-weighted Magnetic Resonance Images (MRI) of patients affected by colorectal cancer. The segmentation approach is based on Convolutional Neural Networks (CNNs) like U-Net. This package provides a series of modules and scripts to visualize, pre-process the DICOM files and to train a U-Net model.
Colorectal cancer is a malignant neoplasm of the large intestine resulting from the uncontrolled proliferation of one of the cells making up the colorectal tract. Colorectal cancer is the second malignant tumor per number of deaths after the lung cancer and the third per number of new cases after the breast and lung cancer. Risk factors for this kind of cancer include colon polyps, long-standing ulcerative colitis, diabetes II and genetic history (HNPCC or Lynch syndrome). In order to get information about diagnosis, therapy evaluation on colorectal cancer, analysis on radiological images can be performed through the application of dedicated algorithms.
In this scenario, the correct and fast identification of the cancer regions is a fundamental task. Up to now this task is performed using manual or semi-automatic techniques, which are time-consuming and subjected to the operator expertise.
This project provides an automatic pipeline for the segmentation of cancer areas on T2-weighted Magnetic Resonance Images of patient affected by colorectal cancer. The segmentation is achieved with a Convolutional Neural Network like U-Net.
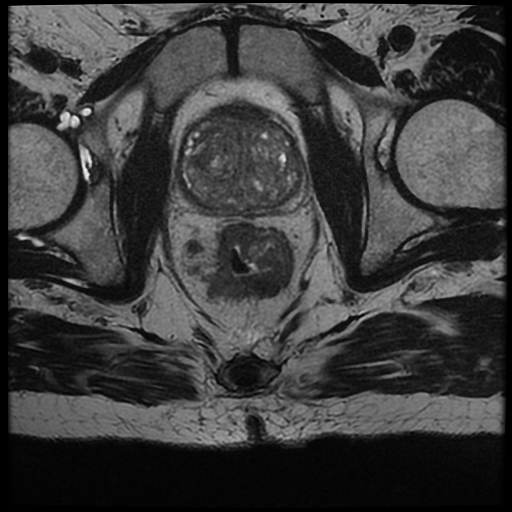
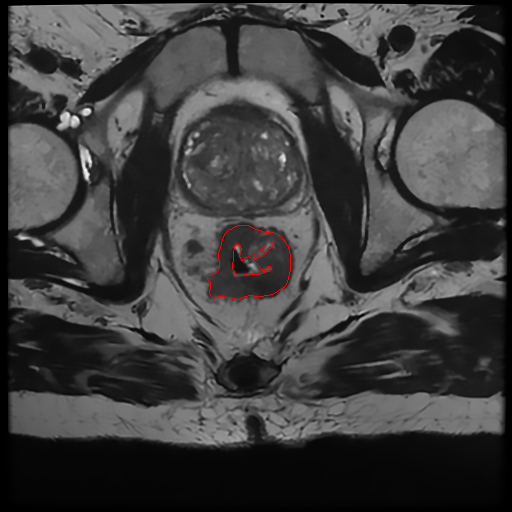
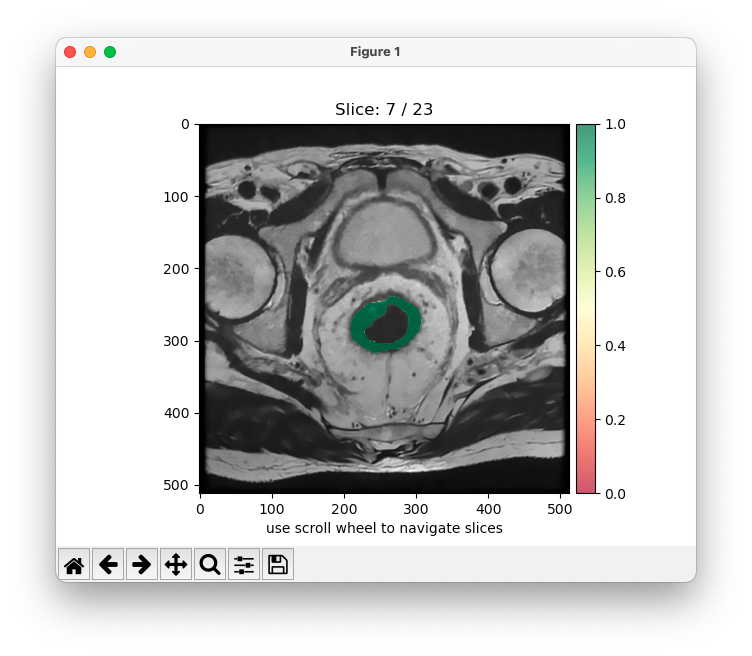
Example of segmentation . Left): Original MRI scan of a patient affected by colorectal cancer. Right): Original MRI scan of a patient affected by colorectal cancer with identified tumor area.
img-segm is composed of a series of modules contained in MRIsegm and scripts in utils:
- modules allow one to load, visualize, processing the DICOM series and to train a U-Net model.
- scripts provide a fast way to handle DICOM series and ROI from command line.
The documentation can be found here.
For a better description of each module:
| Module | Description |
|---|---|
| utils | methods to load and visualize the DICOM series and the relative Region Of Interest (ROI) |
| processing | methods to perform operations such as denoising, resizing, predict the images |
| models | contains the implementation of a U-net model |
| datagenerators | contains methods to generate data for training the model |
| metrics | contains useful metrics for training the model |
| losses | contains useful losses for training the model |
| graphics | methods to display the predictions |
notes:
- It is also possible to use models, metrics and losses from the segmentation-models library.
For a better description of each script:
| Script | Description |
|---|---|
| dcmsorter | move and sort DICOM files from nested dirs like patientID/EXAMINATION/DIRECTORY1/DIRECTORY2 to patientID/EXAMINATION |
| dcm2img | convert DICOM series to .png images |
| roimanager | show the ROI saved as .roi over the images |
For the usage of each script please refer to the documentation.
First of all ensure to have the right python version installed.
This project use Tensorflow, opencv-python, numpy: see the requirements for more information.
⚠️ Tensorflow v2.5 requires the minimum Python version to be 3.6!
First, clone the git repository and change directory:
git clone https://github.com/giuseppefilitto/img-segm.git
cd img-segmThen, pip-install the requirements and run the setup script:
pip install -r requirements.txt
⚠️ Apple Silicon: ensure to have installed a proper Conda env, then install tensorflow as explained here. Finally, install one by one the remaining dependencies using conda (if available) or pip.
python setup.py installTesting routines use PyTest and Hypothesis packages.
A full set of test is provided in testing directory.
You can run the full list of test with:
python -m pytestOnce you have installed it, you can start to segment the images directly from your bash.
The input --dir is the path of the dir containing the DICOM series.
Please ensure that the folder contains only one series.
If the directory is a nested dir, the script will find automatically the sub-dir containing the DICOM series.
python -m MRIsegm --dir='/path/to/input/series/' where:
--diris the path of the directory containing the DICOM series (required).
Name of the model's weights saved in the weights dir.
python -m MRIsegm --dir='/path/to/input/series/' --model='model_name'notes:
model_nameset as default:efficientnetb0_BTC=4_full_150E_OPT=adam_LOSS=dice_loss_plus_1binary_focal_loss- Remember to specify the name without
_weights.h5 - you can also use your own model's weight saving the weights in the
weightsdir asmodel_name_weights.h5.
⚠️ You need to save also the architecture asmodel_name.jsonfile in the same dir.
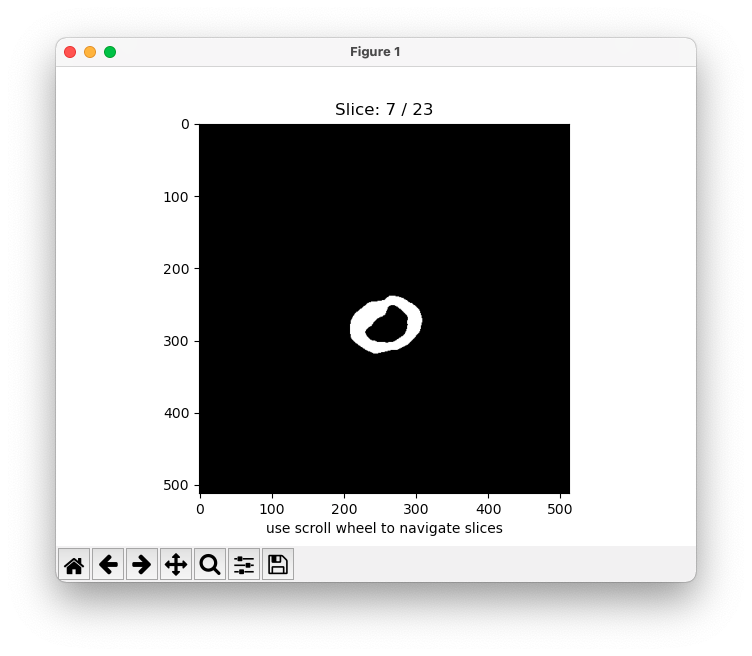
When enabled plot the predicted binary [0, 1] mask of each slice.
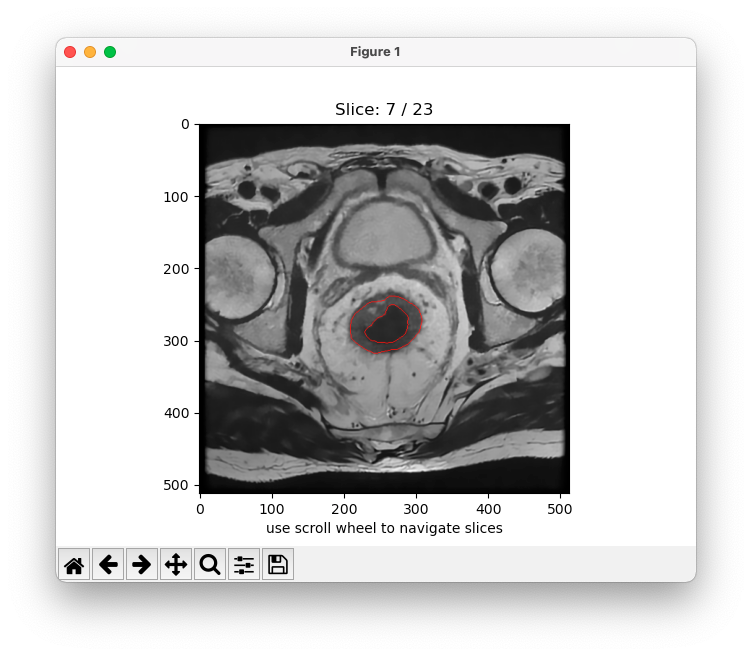
python -m MRIsegm --dir='/path/to/input/series/' --maskWhen enabled plot the predicted probability map between 0. and 1. of each slice over the original image.
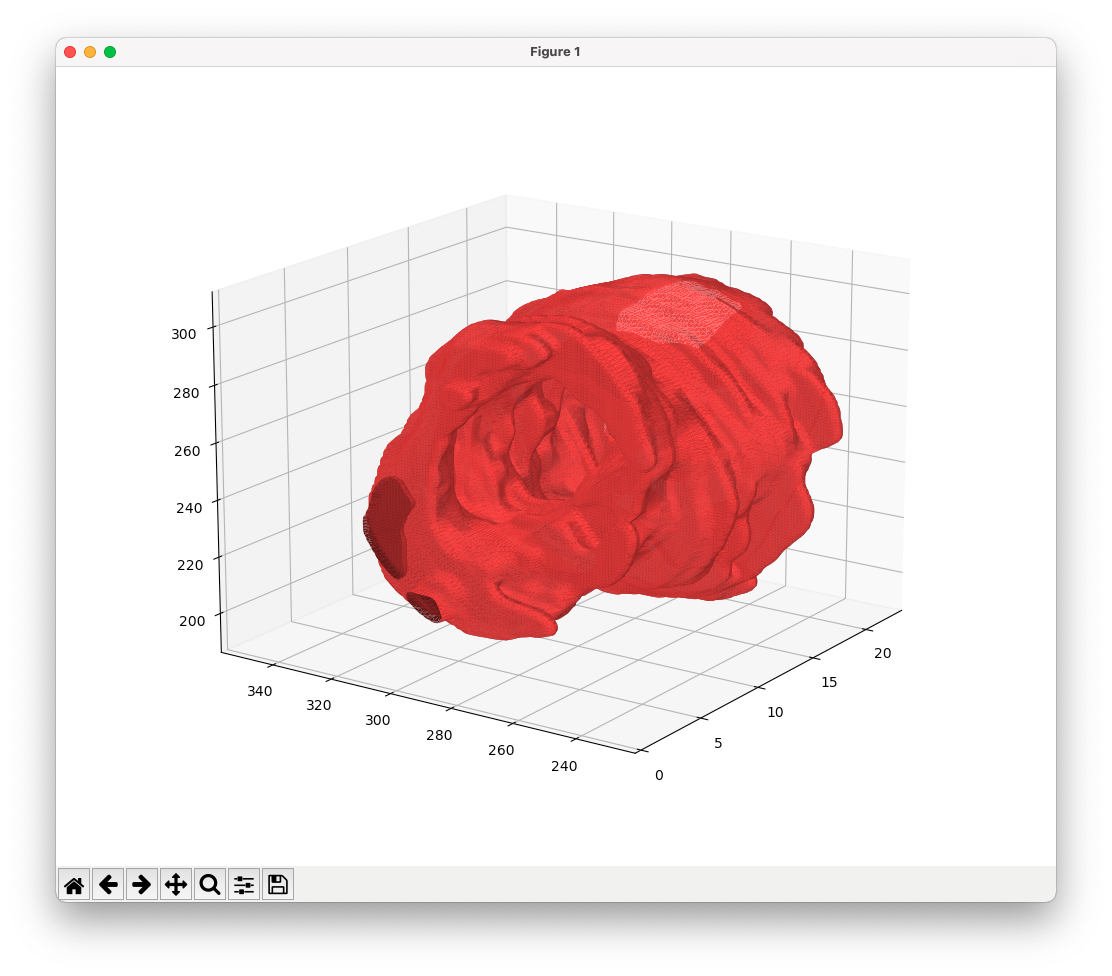
python -m MRIsegm --dir='/path/to/input/series/' --densityWhen enabled plot the a 3D mesh of the segmented areas.
python -m MRIsegm --dir='/path/to/input/series/' --3DSeveral examples are available under the notebooks dir.
Segmentation:
| Purpose | Example |
|---|---|
| load, visualize DICOM series | DicomExplorer |
| perform Image processing operations | ImageProcessing |
| Prepare the data for model training | PrepareData |
| train a U-net model | UNet-Segmentation |
| train a model from segmentation-models | Segmentation |
| Display the predictions | VisualizePredictions |
| Save the predicted image | SavePredictions |
| Convert model to model's weights | ConverterToWeights |
| plot 3D mesh | 3D-mesh |
features analysis:
| Purpose | Example |
|---|---|
| Feature Extraction | FeaturesExtraction |
| Features Analysis | Features Analysis |
| Write .dcm files | Writedicom |
| Write .nrrd files | sitkvolumeR2 |
This package is licensed under the MIT "Expat" License.
Giuseppe Filitto git
If you have found img-segm helpful in your research, please
consider citing this project
@misc{MRI colorectal cancer segmentation,
author = {Filitto, Giuseppe},
title = {MRI colorectal cancer segmentation},
year = {2021},
publisher = {GitHub},
howpublished = {\url{https://github.com/giuseppefilitto/img-segm}},
}