On this page we summarize our efforts in the CZI funded project Seamless Integration of Quantitative Bio-Image Analysis Plugins. The original plan was to support the napari plugin community by providing guidance in making napari plugins interoperable and to produce user-documentation across plugins.
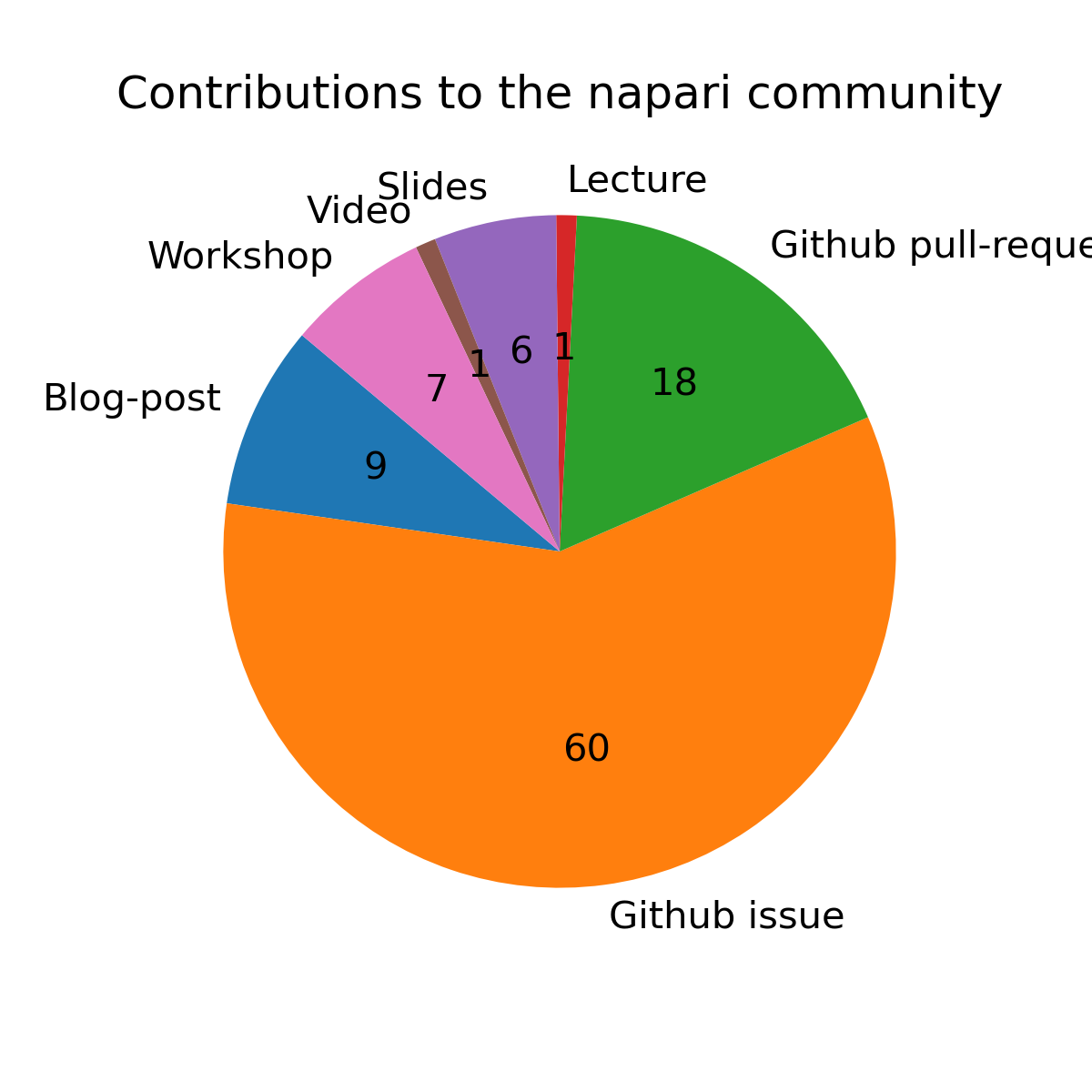
Our project team consisted of Mara Lampert and Robert Haase. We produced various materials as visualized in this overview pie chart.
We opened about 60 Github issues and 18 pull-requests, commonly supporting napari-plugin developers in enriching their documentation and making their napari-plugins work on other computers. Also, we assisted plugin developers in making their plugins interoperable with each other. In this way we contributed to about 30 napari plugins, the napari-core and the napari-hub website.
Following our original goal, we provided napari-plugin documentation in form of blog-posts and videos that show how to work with multiple napari-plugins together.
- Rescaling images and Pixel Anisotropy
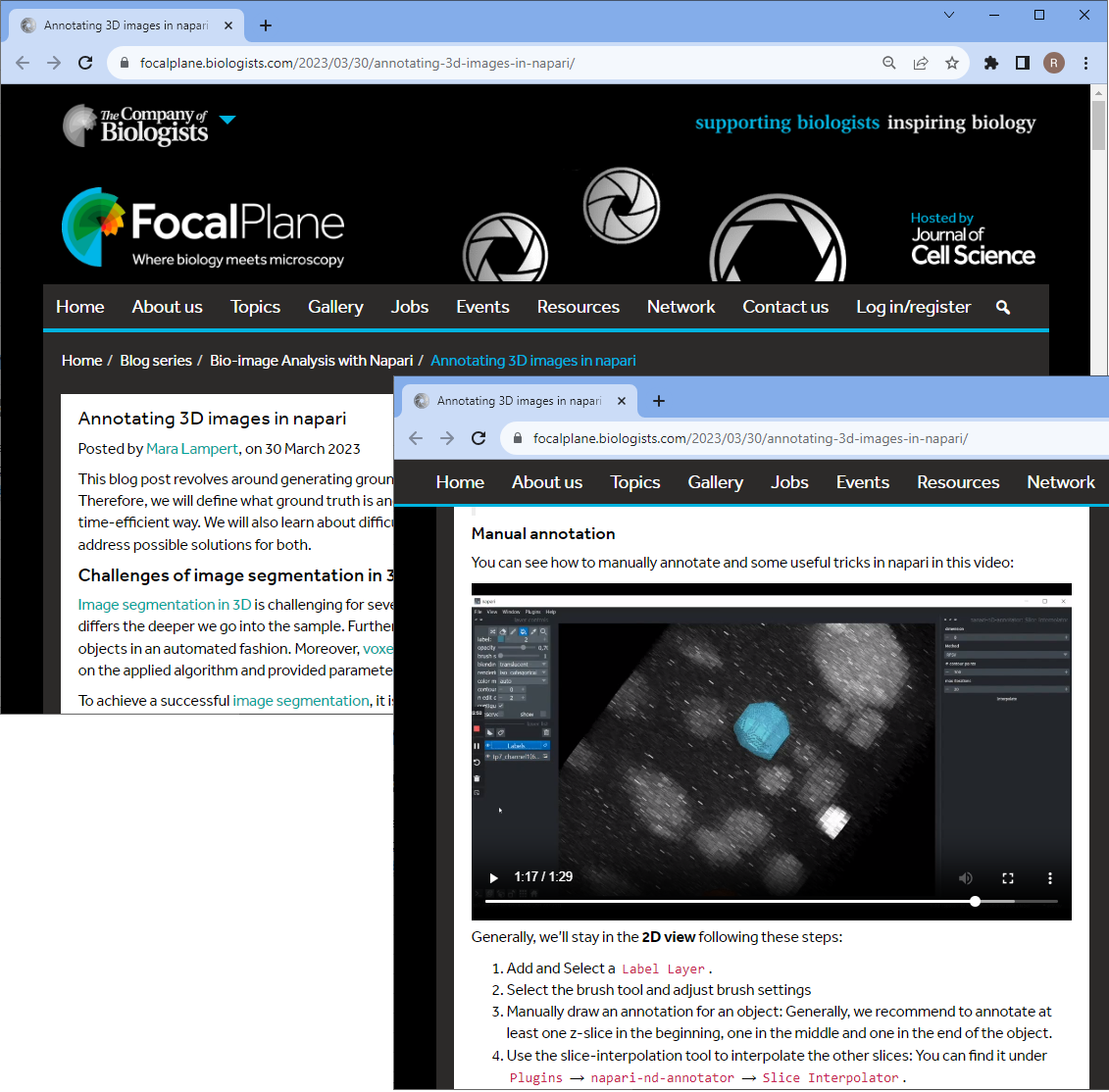
- Annotating 3D images in Napari
- Quality assurance of segmentation results
- Feature extraction in Napari
- Tracking in Napari
- Bio-imaGE Analysis using Napari and Python (EMBO Practical Course, Video)
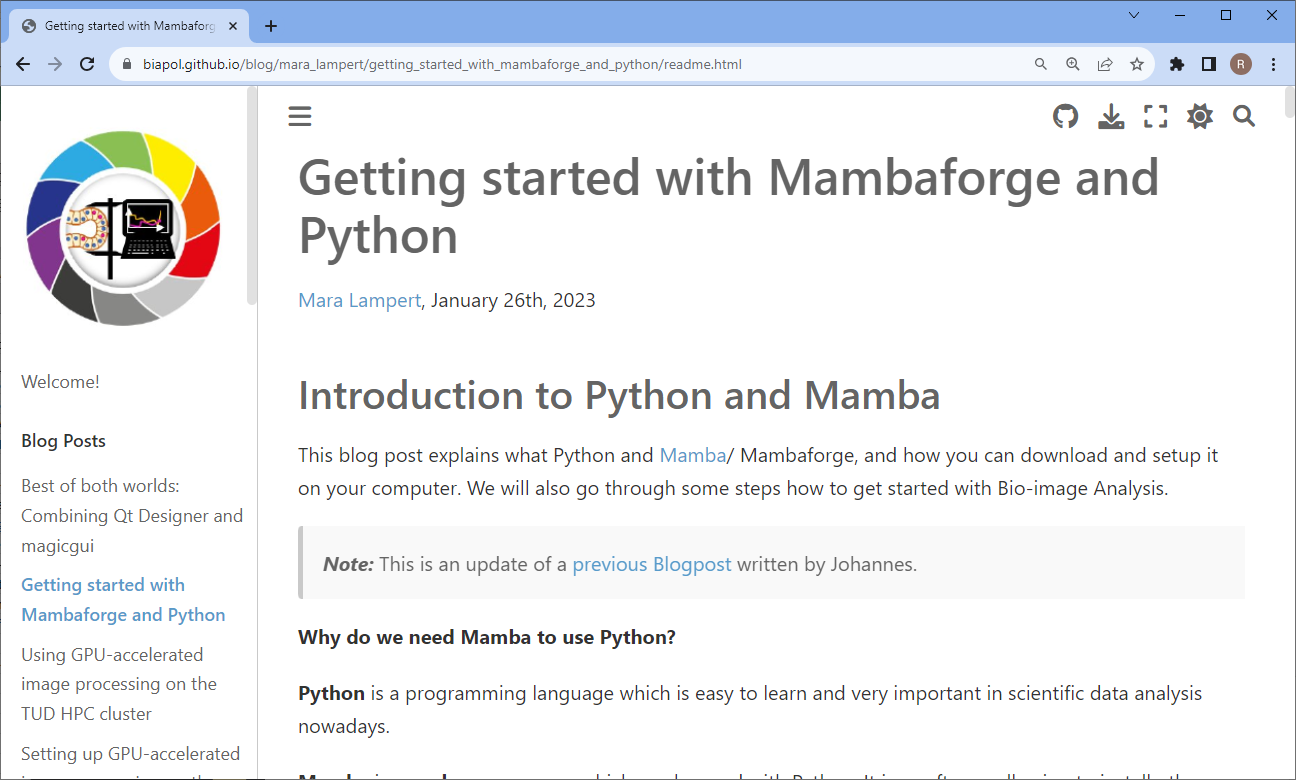
As life scientists in our local environment commonly struggle with installing napari, we also wrote a blog-post outlining how to do this:
As guidance for napari-plugin developers and data scientists more broadly, we wrote blog posts introducing them to good practices for user documentation, licensing and sharing.
- Scientific Data Analysis: User Documentation 101
- If you license it, it'll be harder to steal it
- Sharing research data with zenodo
We also shared slides of our presentations online:
- Bio-image analysis using Napari and Python
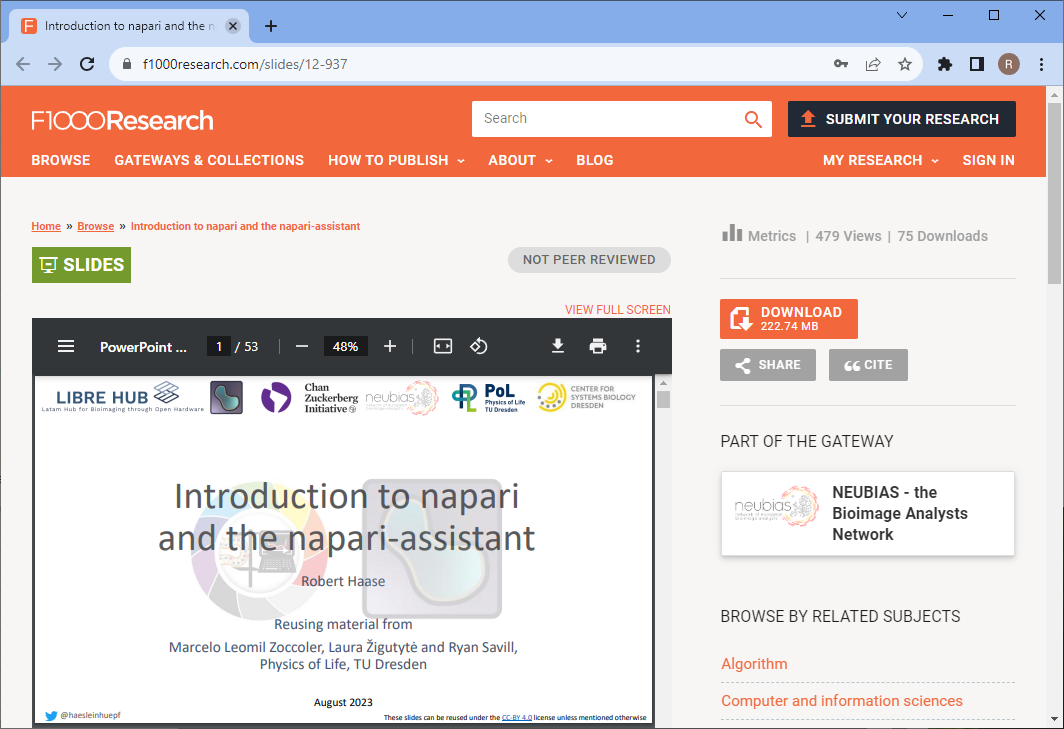
- Introduction to napari and the napari-assistant
- GPU-acclerated Bio-Image Analysis using clesperanto
- Introduction to Napari
- [From assistant to notebooks](https://f1000research.com/slides/12-573 https://f1000research.com/slides/11-1351)
We attended to teach others how to use Napari
- Napari LaTam Workshop Image data science with Python and Napari
- PoL Bio-Image Analysis Training School on GPU-Accelerated Image Analysis
- PoL Bio-Image Analysis Training School - Early Career Track
- Quantitative the Bio-Image Analysis with Python
- DIGS-BB - Light Microscopy Course - Bio-Image Analysis 2022
- Image data science with Python and Napari @EPFL
- NEUBIAS Pasteur course on Bioimage Analysis
Some of these workshops were organized as part of the CZI-EOSS cylce 4 project GPU-accelerating Fiji and Friends Using Distributed CLIJ, NEUBIAS-style.
You find a complete list of links to our actions in this csv file.
This project has been made possible by grant number 2022-252520 from the Chan Zuckerberg Initiative DAF, an advised fund of the Silicon Valley Community Foundation. This project was supported by the Deutsche Forschungsgemeinschaft under Germany’s Excellence Strategy – EXC2068 – Cluster of Excellence “Physics of Life” of TU Dresden.