| Name |
Species | Sample | Modality | Size |
Resolution |
Number | Link | Reference | Note |
|---|---|---|---|---|---|---|---|---|---|
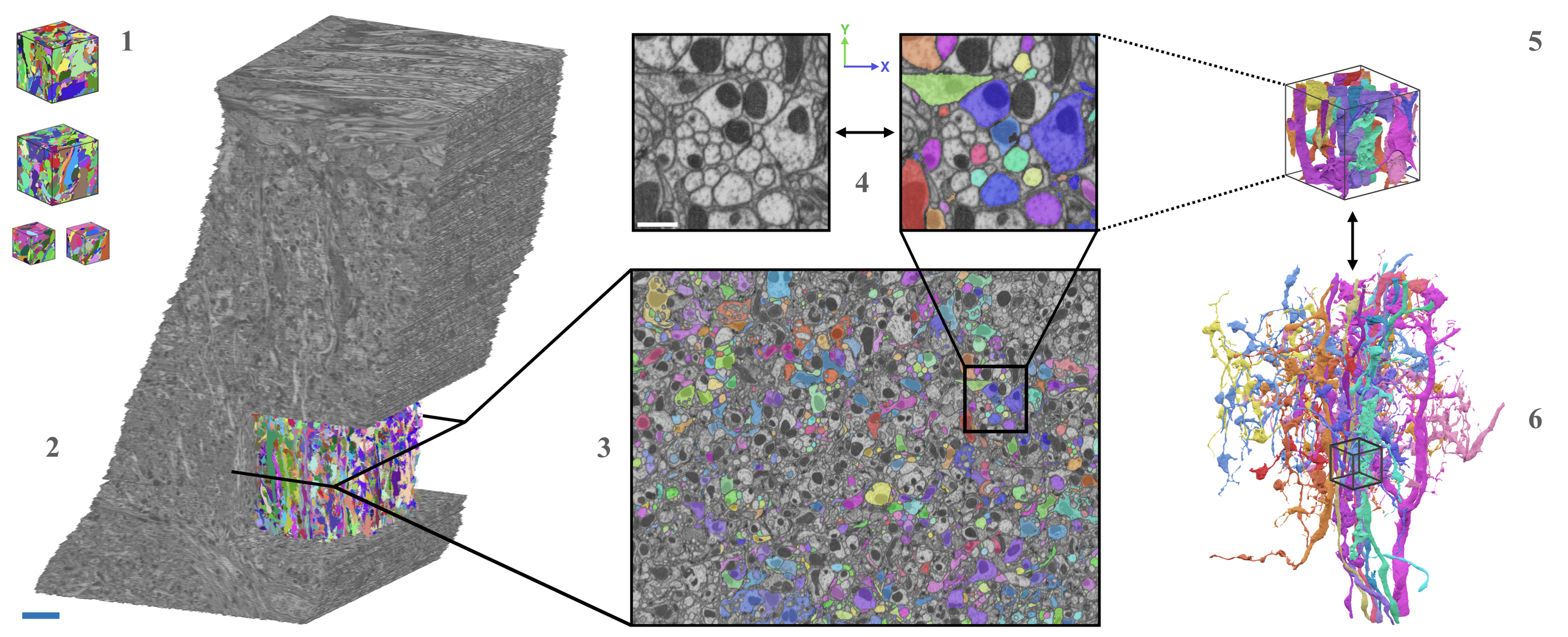
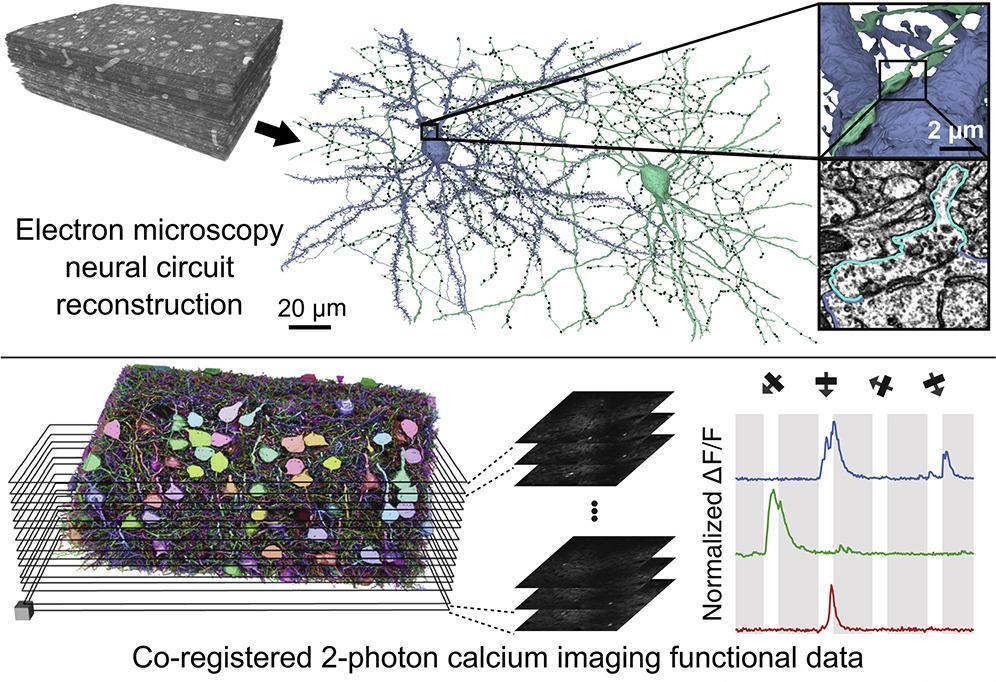
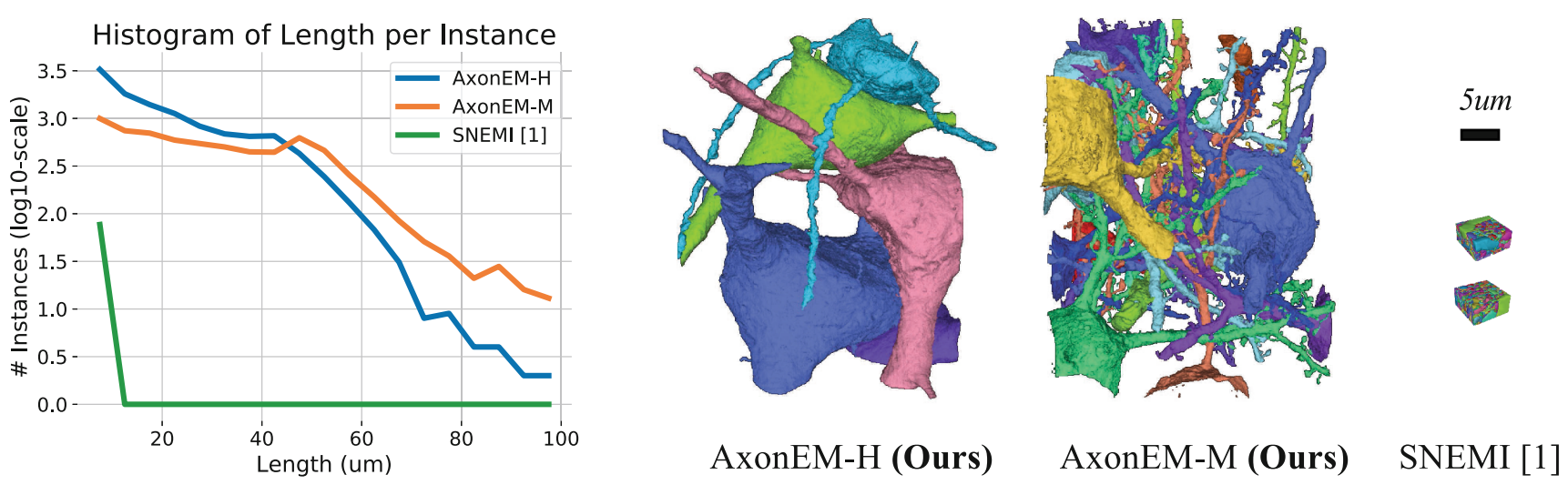
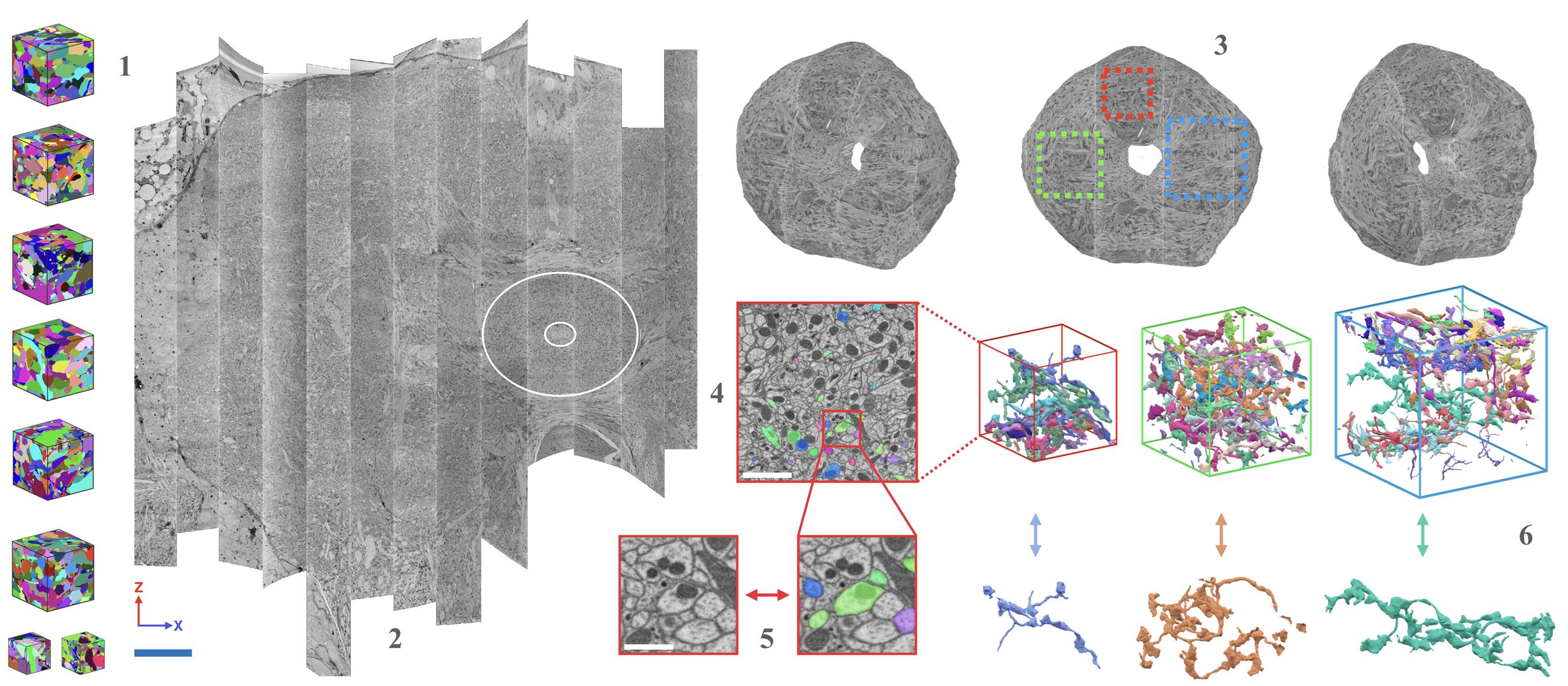
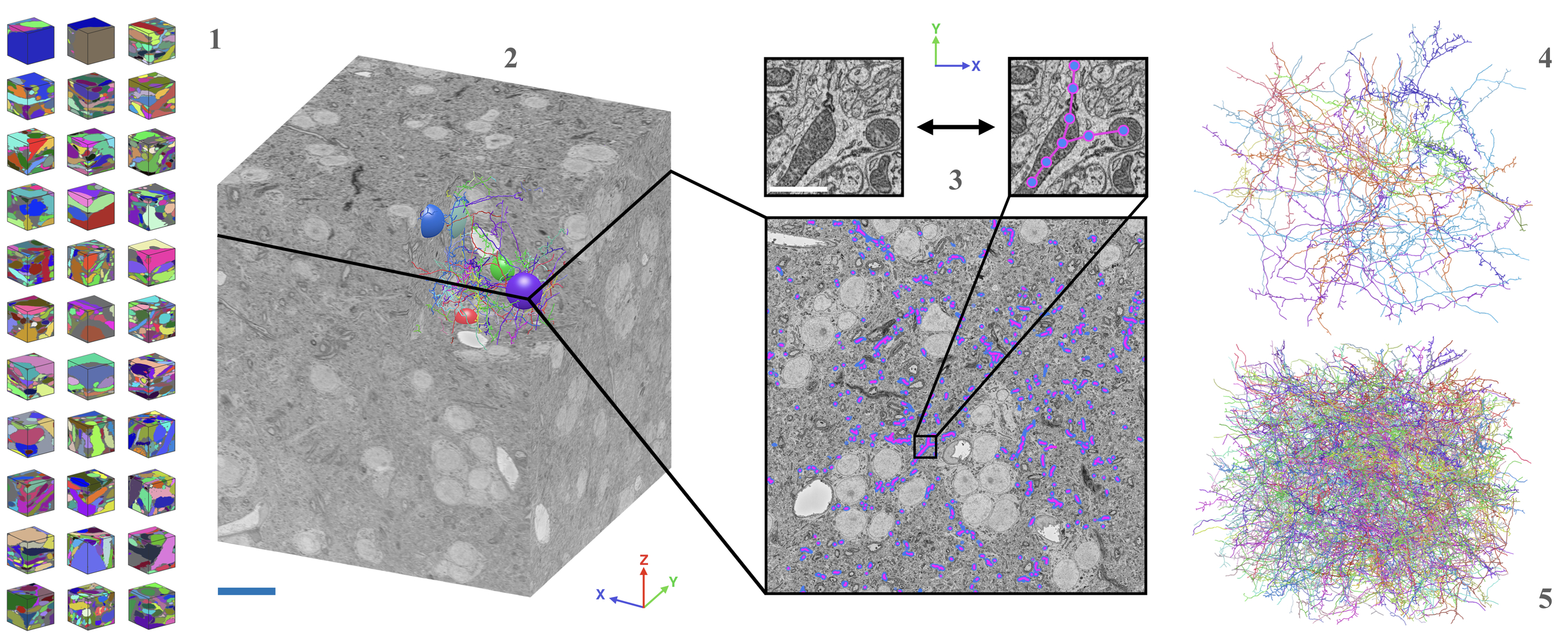
| AxonEM | Mouse Human |
layer 2/3 in the primary visual cortex layer 2 in the temporal lobe |
TEM SEM |
|
18,000 axons | n/a | Wei |
two subset of MICrONS and H01 |
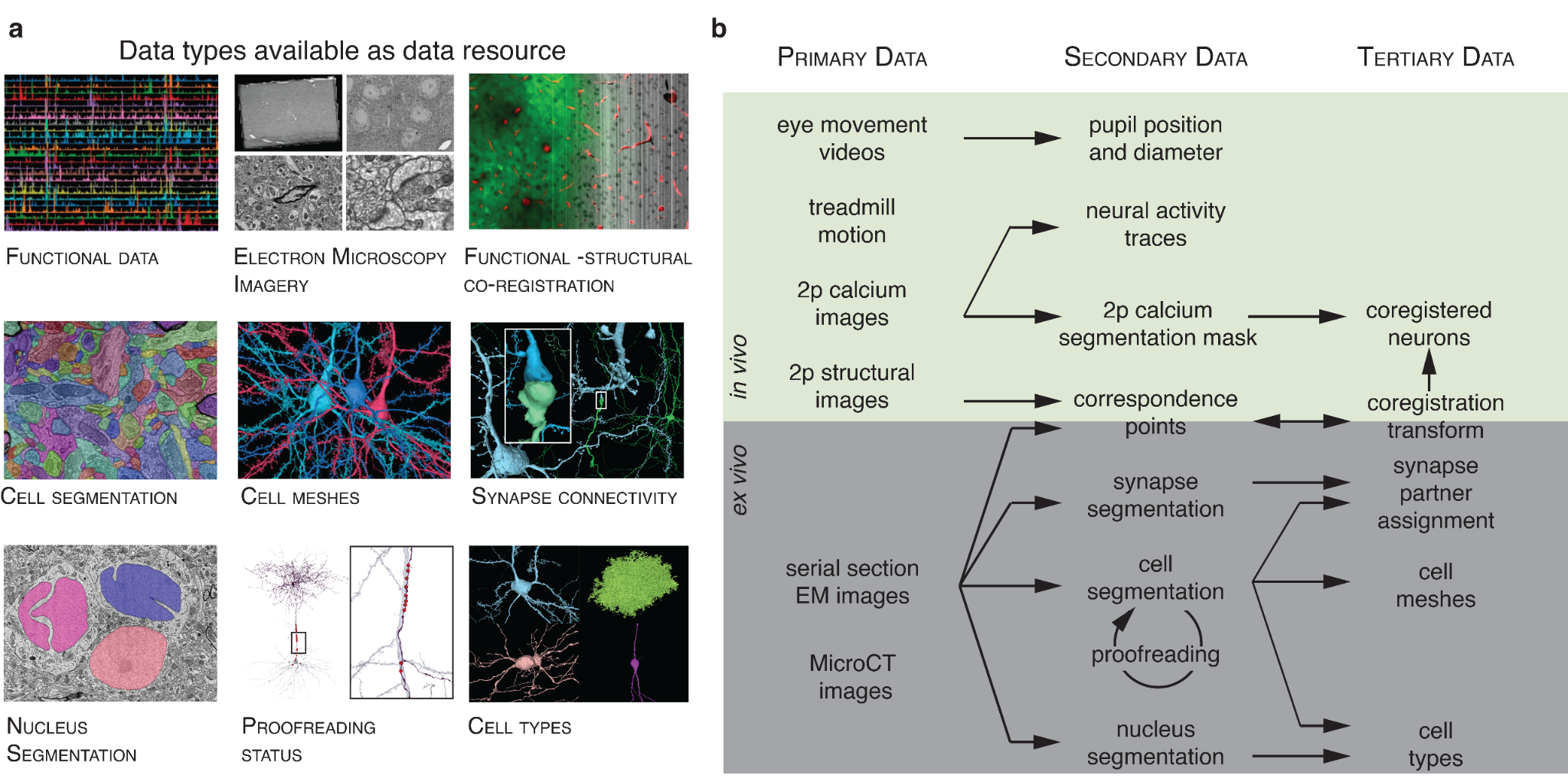
- Turner
$\textit{et al.}$ Reconstruction of Neocortex: Organelles, Compartments, Cells, Circuits, and Activity
- Wei
$\textit{et al.}$ AxonEM Dataset: 3D Axon Instance Segmentation of Brain Cortical Regions
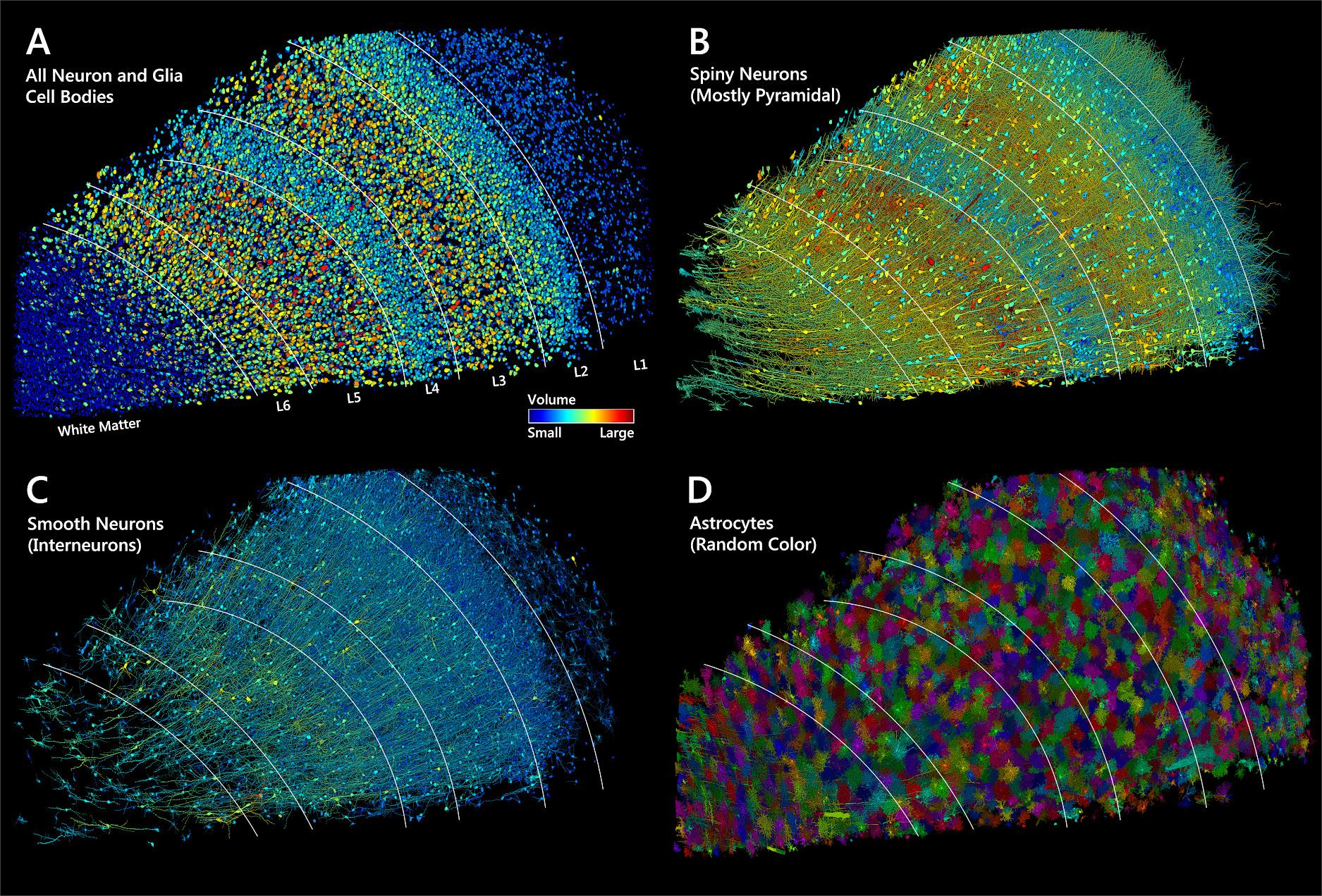
- Shapson-Coe
$\textit{et al.}$ A Connectomic Study of a Petascale Fragment of Human Cerebral Cortex
- MICrONS Consortium
$\textit{et al.}$ Functional Connectomics Spanning Multiple Areas of Mouse Visual Cortex
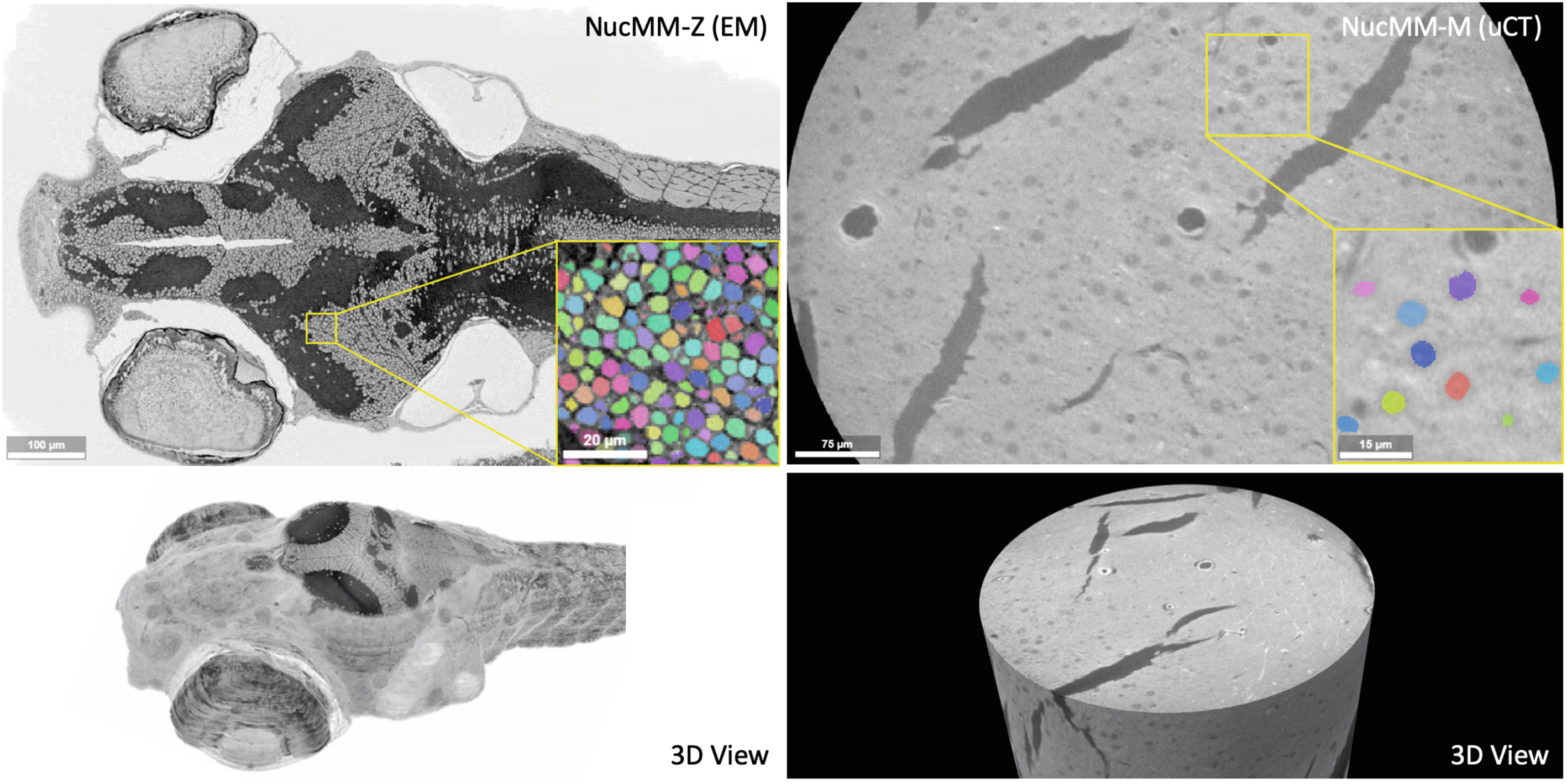
- Lin
$\textit{et al.}$ NucMM Dataset: 3D Neuronal Nuclei Instance Segmentation at Sub-Cubic Millimeter Scale
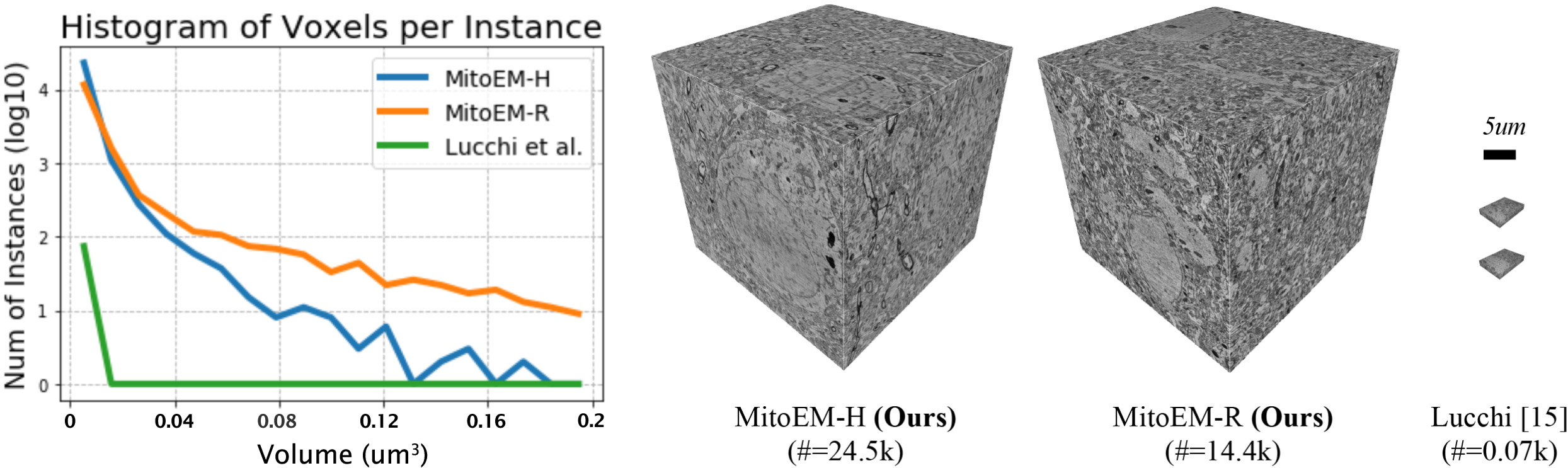
- Wei
$\textit{et al.}$ MitoEM Dataset: Large-Scale 3D Mitochondria Instance Segmentation from EM Images
- Scheffer
$\textit{et al.}$ A connectome and analysis of the adult Drosophila central brain
- Januszewski
$\textit{et al.}$ High-precision automated reconstruction of neurons with flood-filling networks
- Takemura
$\textit{et al.}$ Synaptic circuits and their variations within different columns in the visual system of Drosophila
- Kasthuri
$\textit{et al.}$ Saturated Reconstruction of a Volume of Neocortex
- Lichtman Lab, Department of Molecular and Cellular Biology, Center for Brain Science, Harvard University
- Seung Lab, Computer Science Department, Princeton Neuroscience Institute
- Helmstaedter Lab, Department of Connectomics, Max Planck Institute for Brain Research
- Pfister Lab, Visual Computing Group, School of Engineering and Applied Sciences, Harvard University
- Funke Lab, Janelia Research Campus, Howard Hughes Medical Institute
Please contribute if you think a new dataset or a relevant paper is missing.
Let's enjoy the beauty of EM data!