Spacewalk provides interactive 3D visualization of super-resolution microscopy data, with integrated genomic analysis via the genomics browser igv.js and the Hi-C map viewer juicebox.js
- Node >= v20.8.0
- NPM >= v10.1.0
Spacewalk require a modern web browser with support for Javascript ECMAScript 2015.
- Clone this repository.
git clone git@github.com:igvteam/spacewalk.git
- Install
npm install
npm run build
npm run start
- Open a browser and enter the follow url to launch the app
localhost:8080/index.html
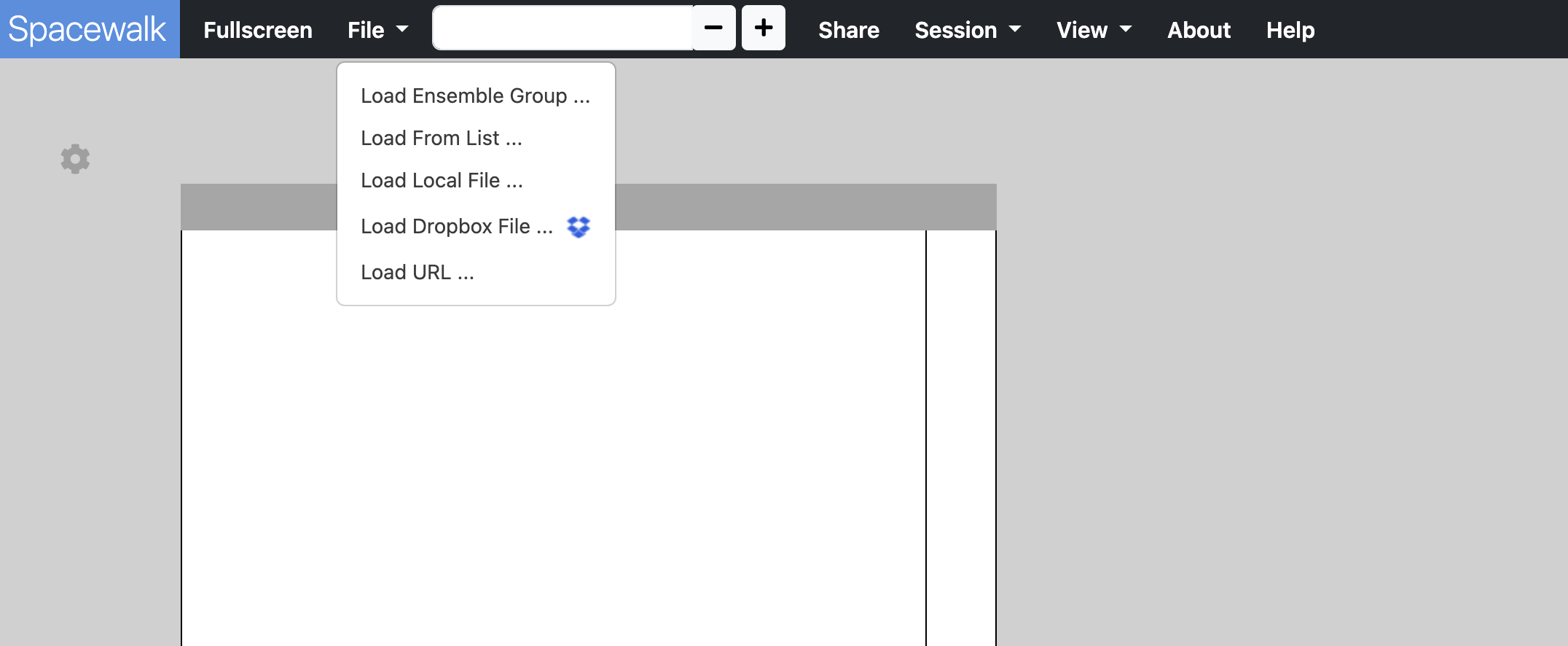
After launching the app, you will see a screen with a single empty 3D viewer. In the navbar use the File dropdown menu to load 3D structure into the 3D viewer.
Spacewalk supports the 3D visualization of
- Super-resolution microscopy (SRM) data
- Chromatin simulations
- Other forms of genome microscopy and spatial genomics
Spacewalk supports 3D visualization of data that comes in two general forms:
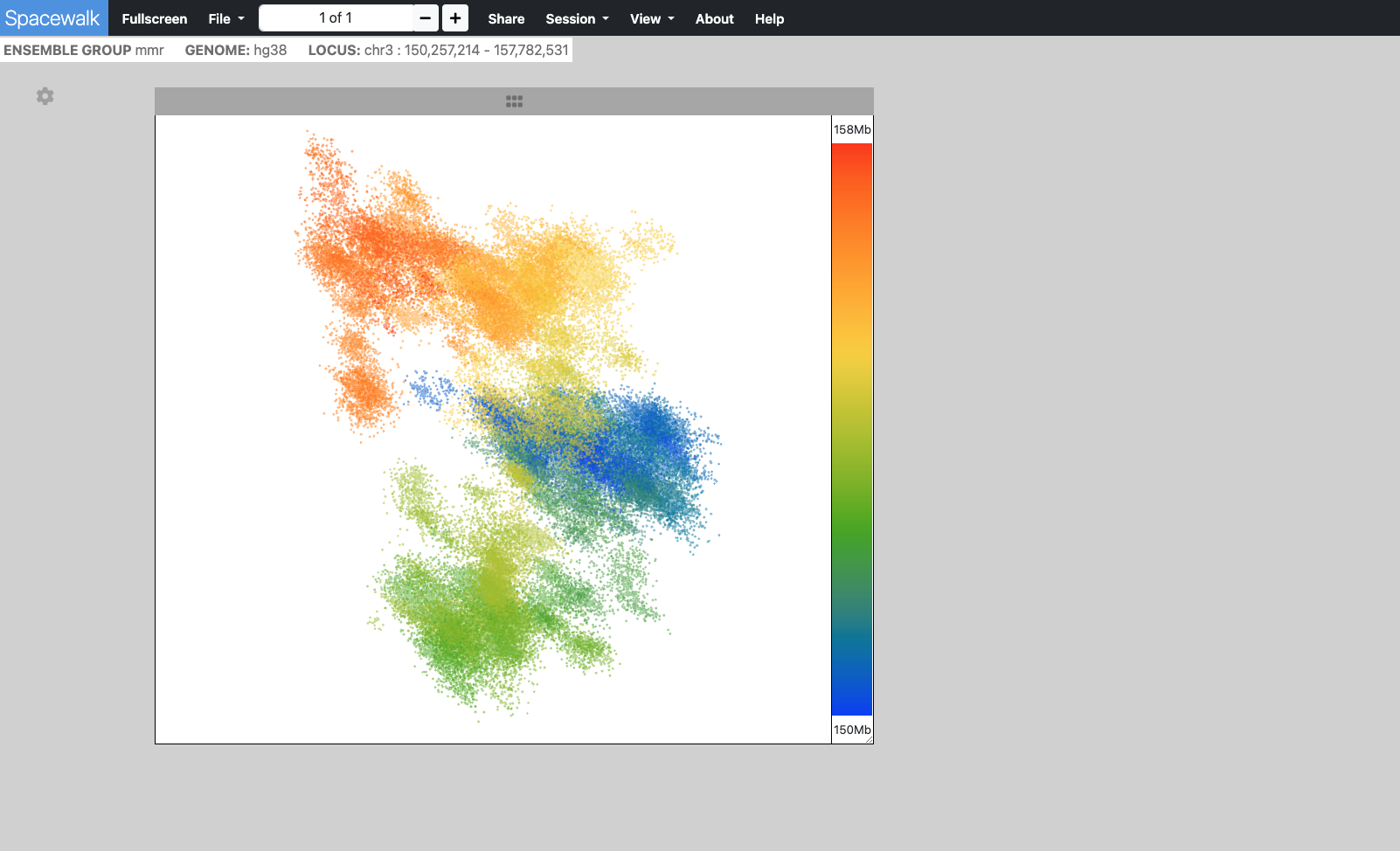
- Point Cloud - Typically derived from OlioSTORM data
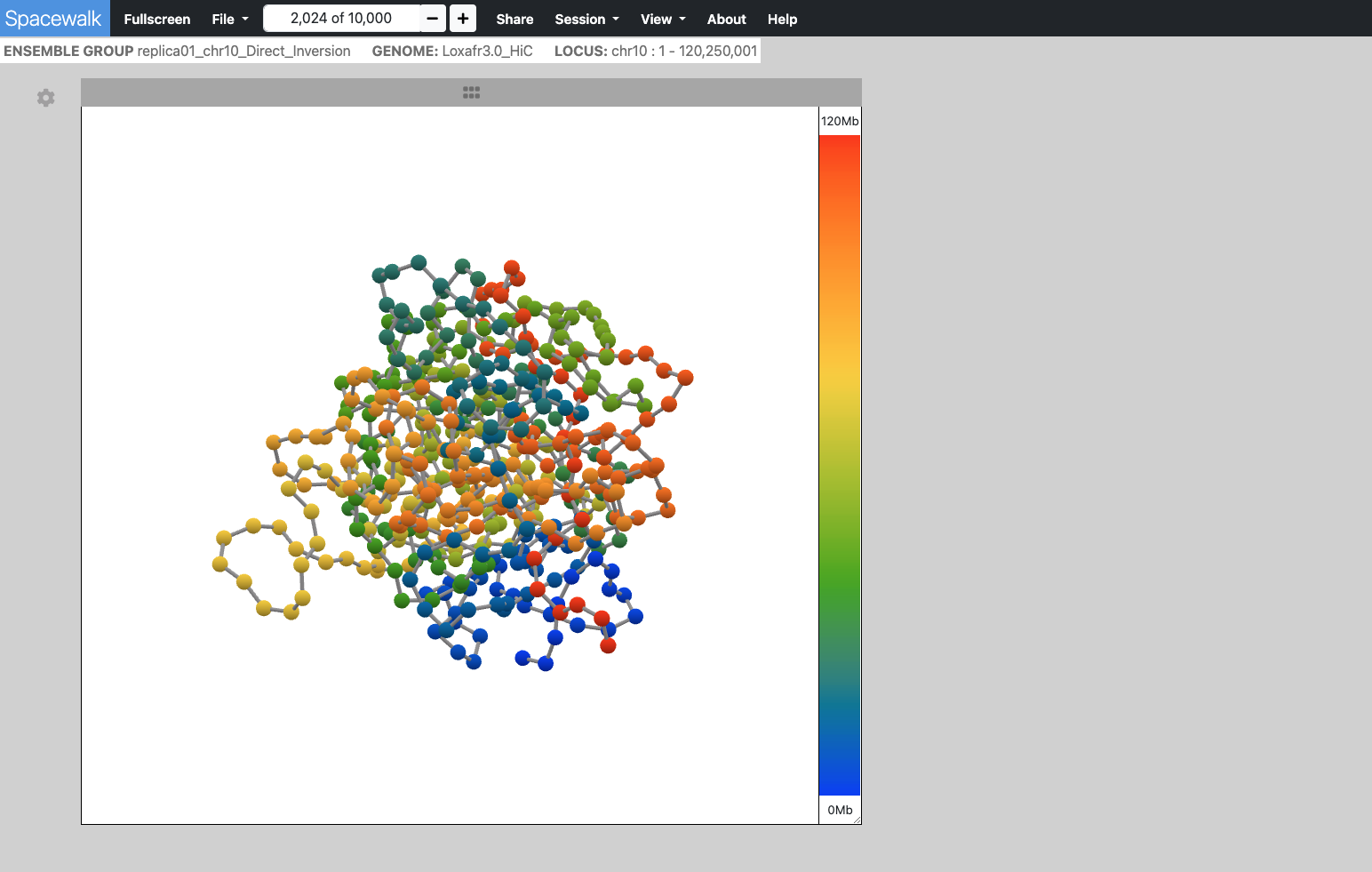
- Ball & Stick - Typically derived from Chromatin simulations
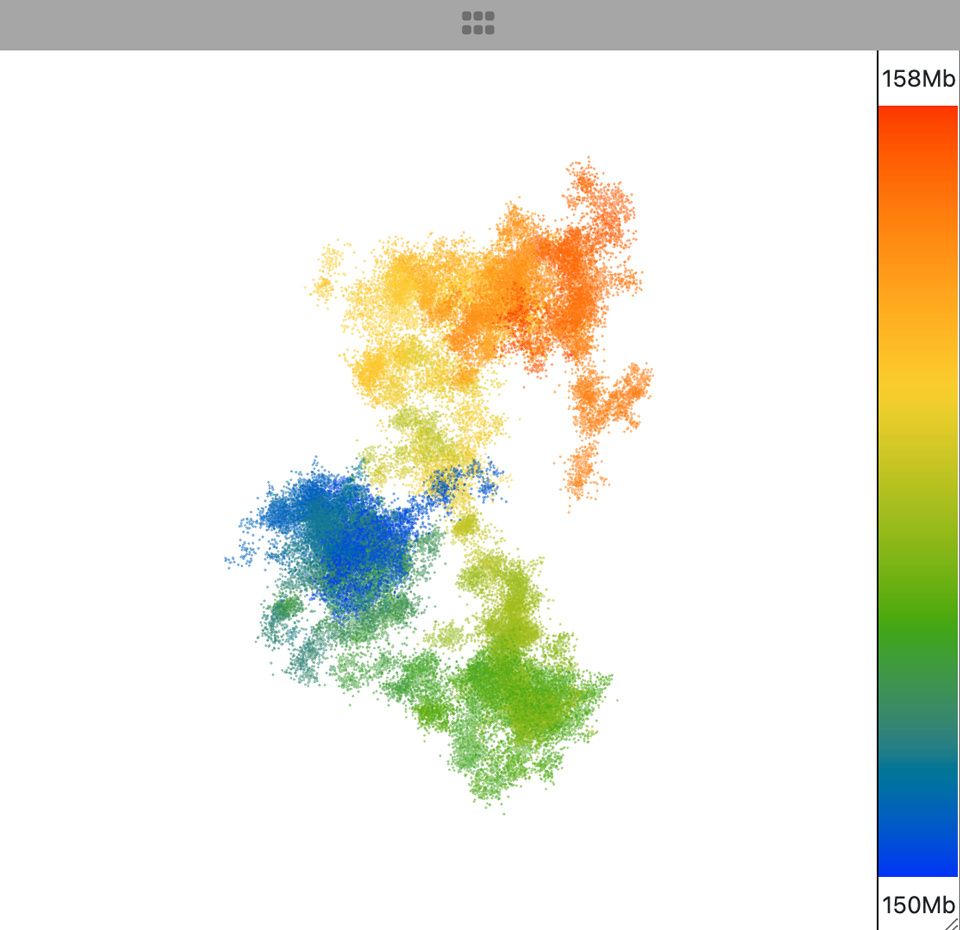
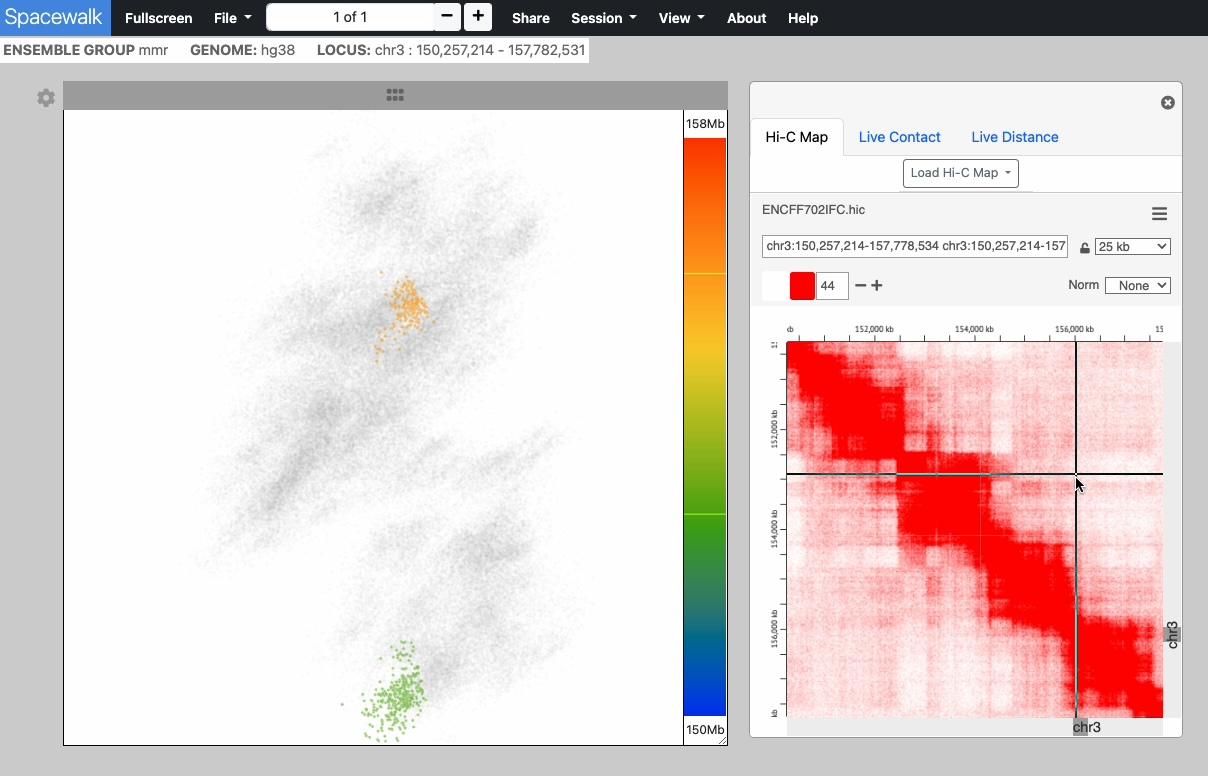
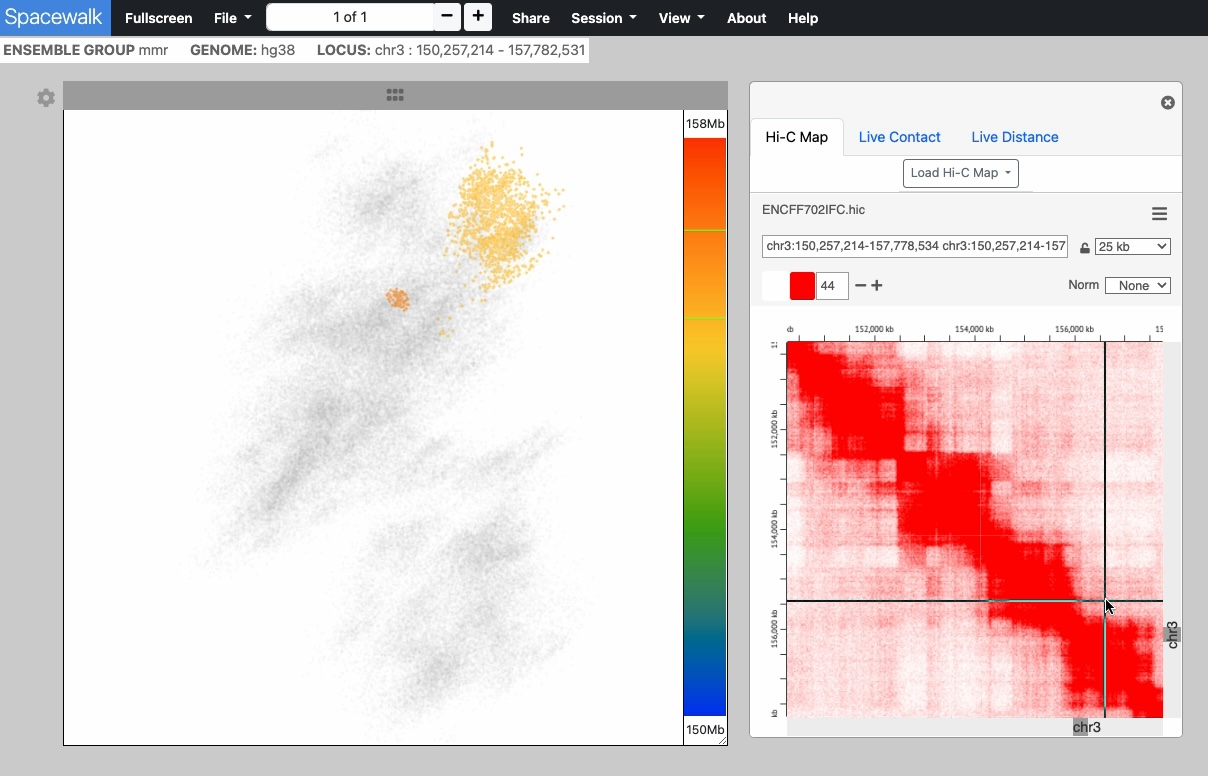
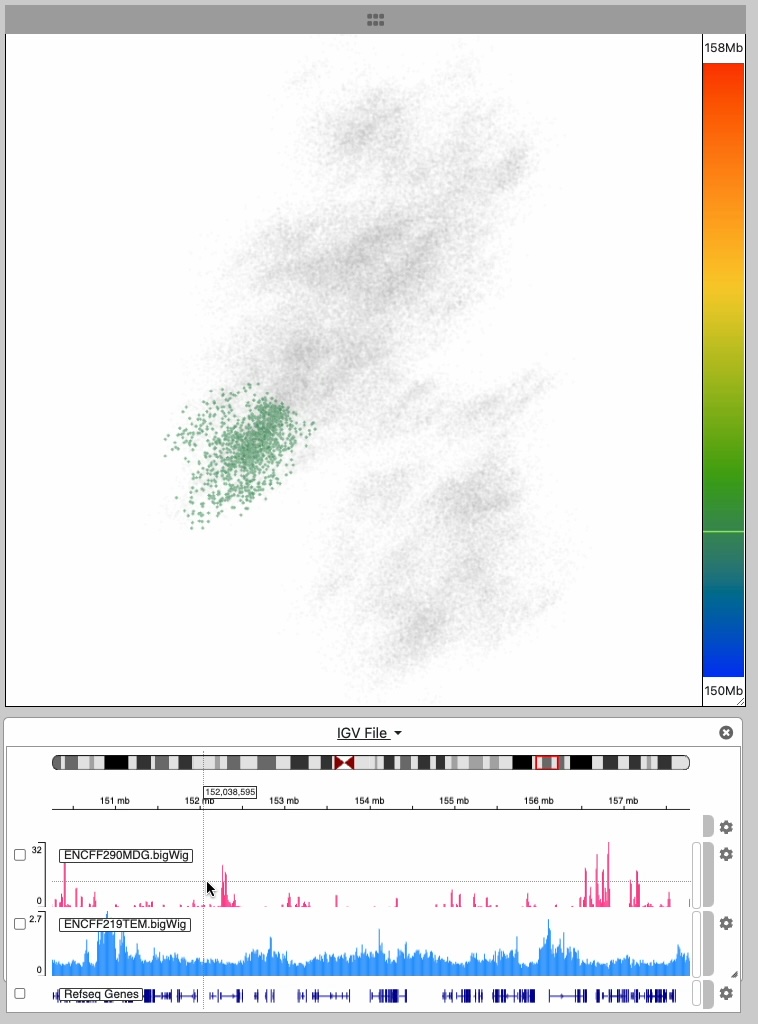
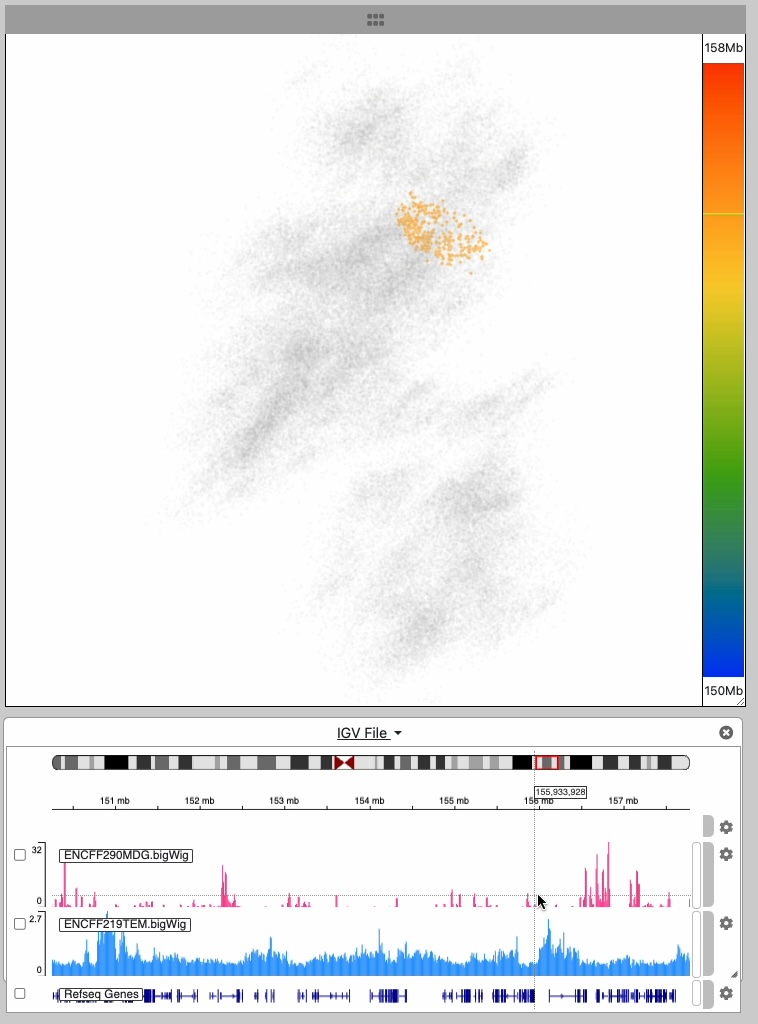
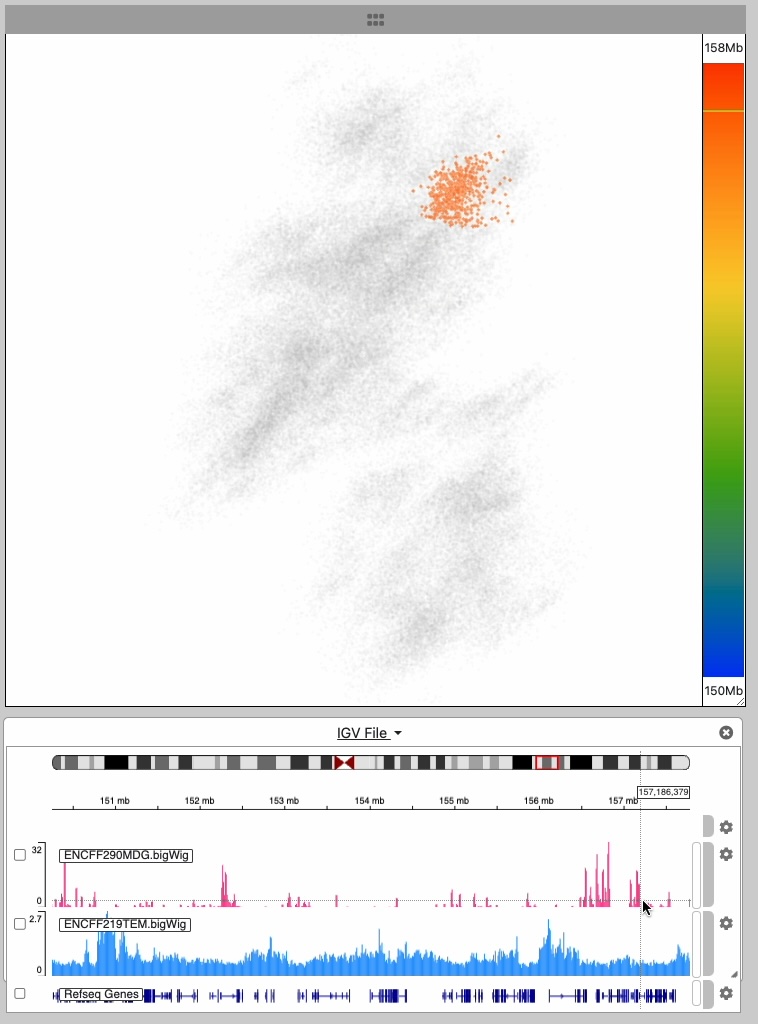
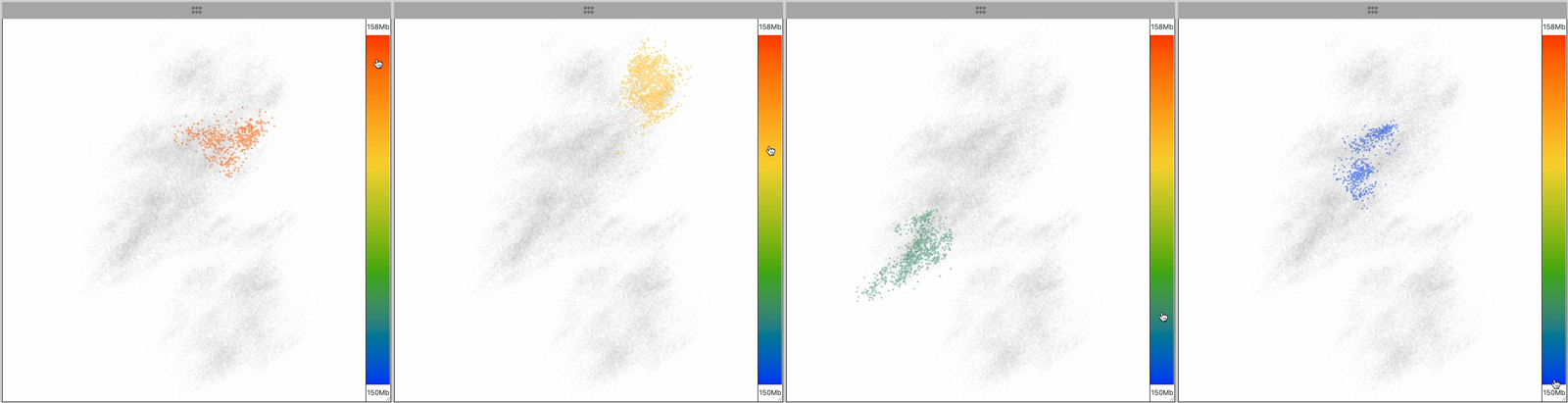
The point cloud is rendered as a collection of 3D point clusters, each corresponding to a specific genomic extent. The color of each cluster is determined by the genomic navigator's color ramp bar, located on the right side of the 3D viewer.
When you mouse over the genomic navigator the corresponding 3D point cluster is highlighted.
Chromatin centroids are rendered as balls, each colored according to its genomic location. Sticks (cylinders) connect the balls in the order they appear along the genomic range. As the user moves the cursor along the genomic navigator on the right side of the 3D viewer, the corresponding ball is highlighted based on its genomic location.
Alternatively, hovering over a ball will display its genomic location in the genomic navigator's color ramp.
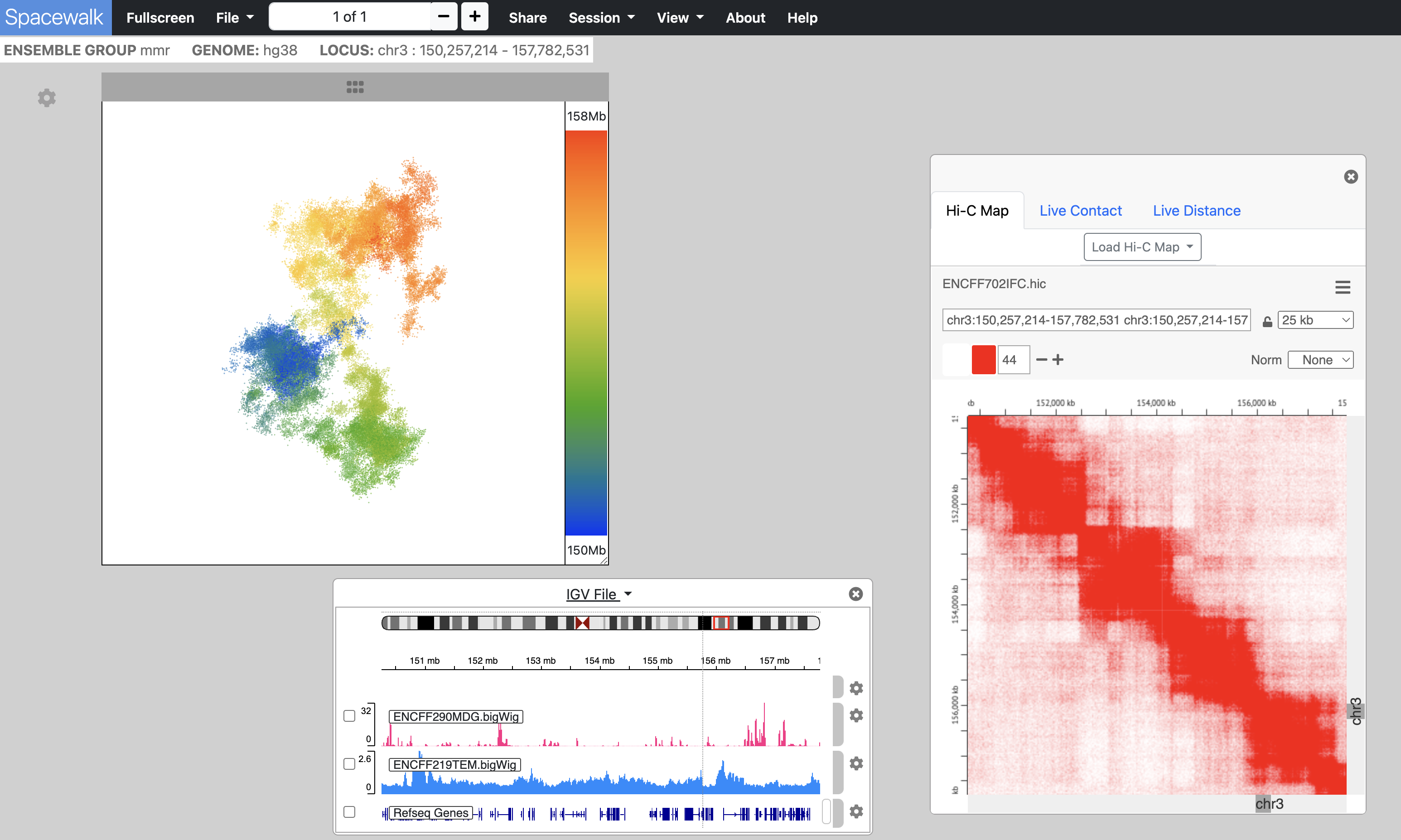
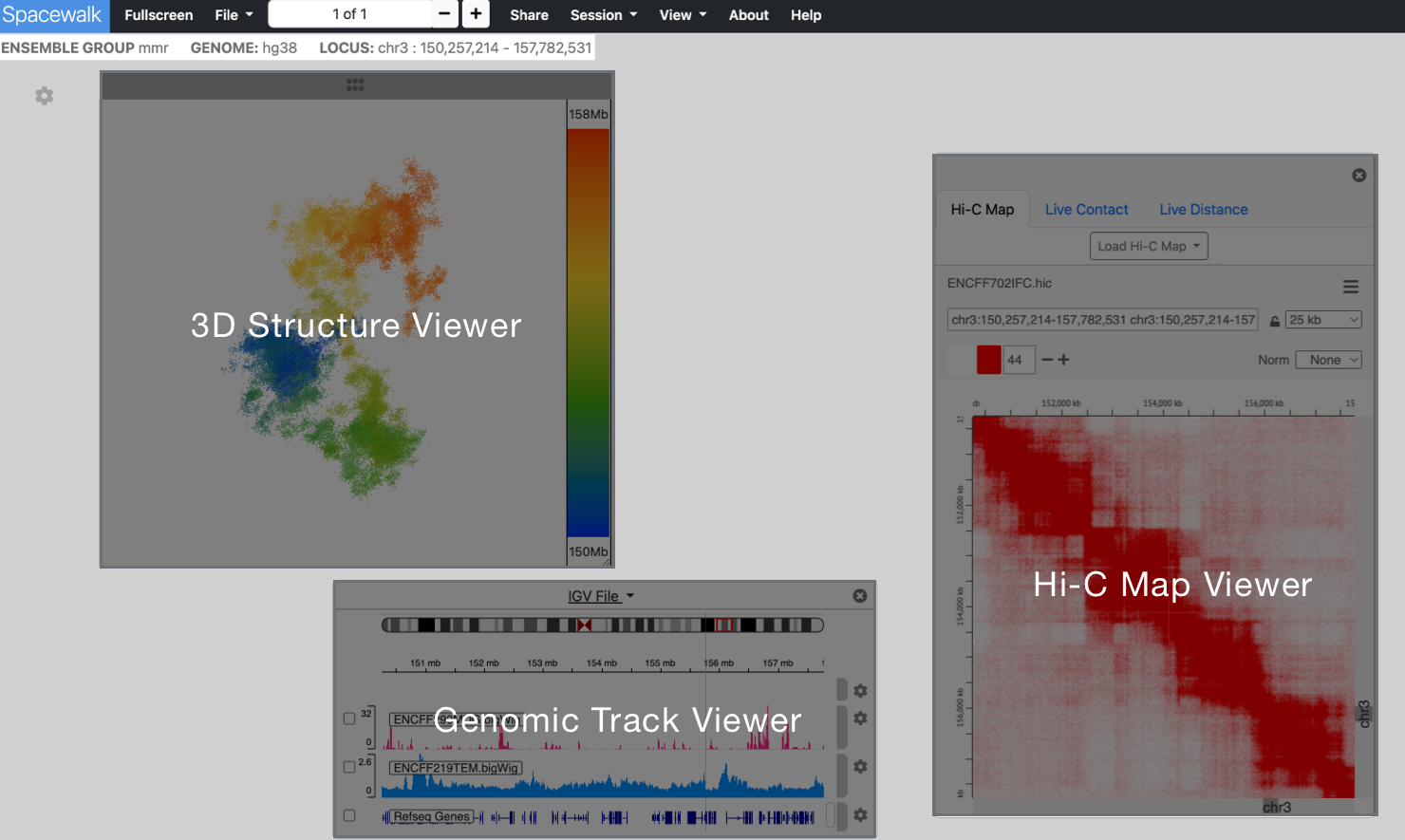
Spacewalk is organized around three visualization panels, each responsible for one aspect of genomic visualization:
A 3D structure - SRM, simulation, etc. - is loaded by clicking on the File dropdown menu in the navbar
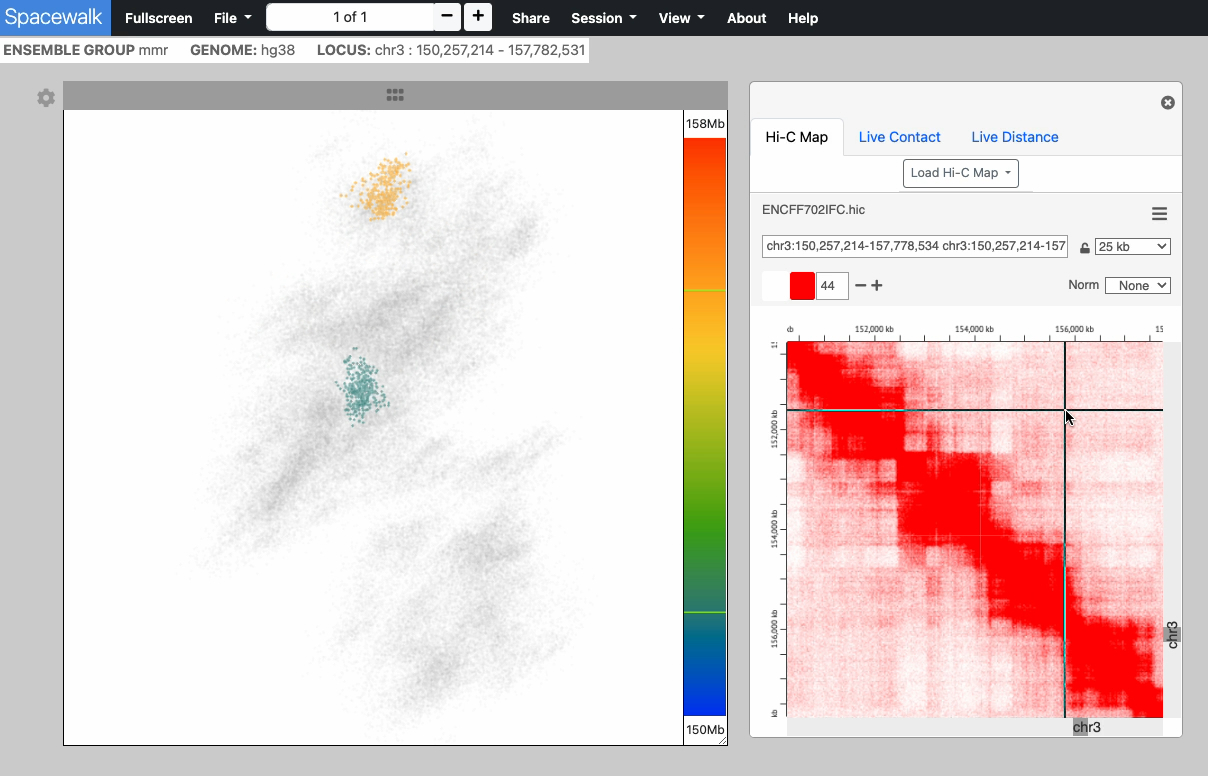
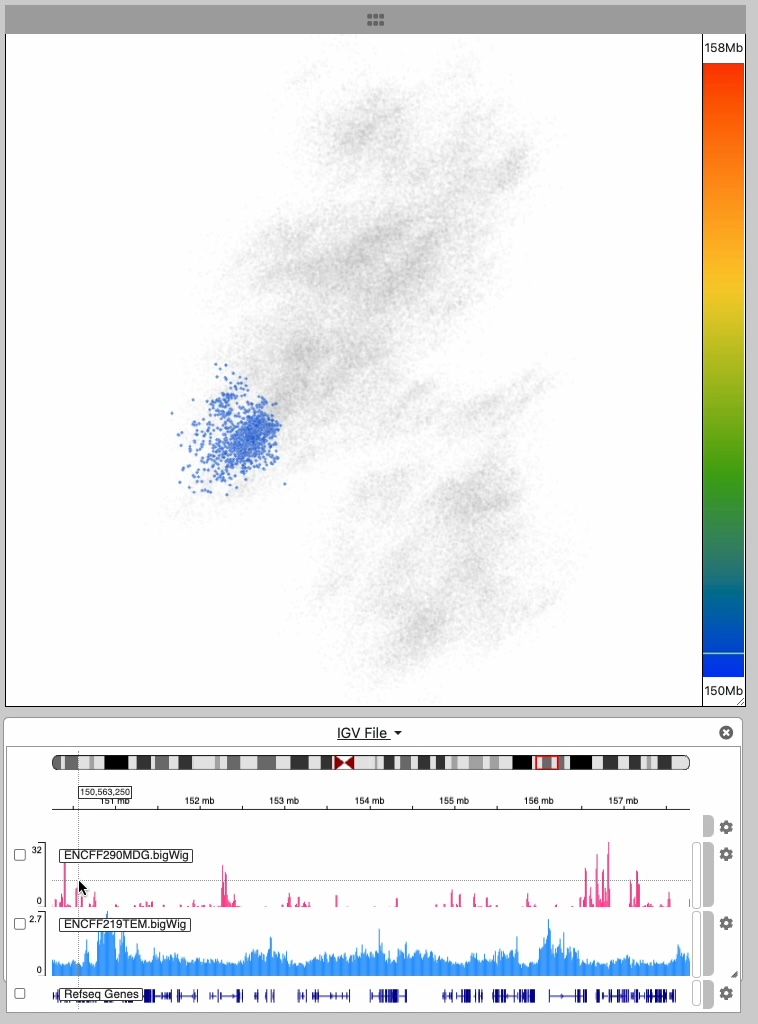
This image series shows the cursor moving along the genomic extent of the 3D structure.
Notice the highlighting of the 3D structure during the interaction
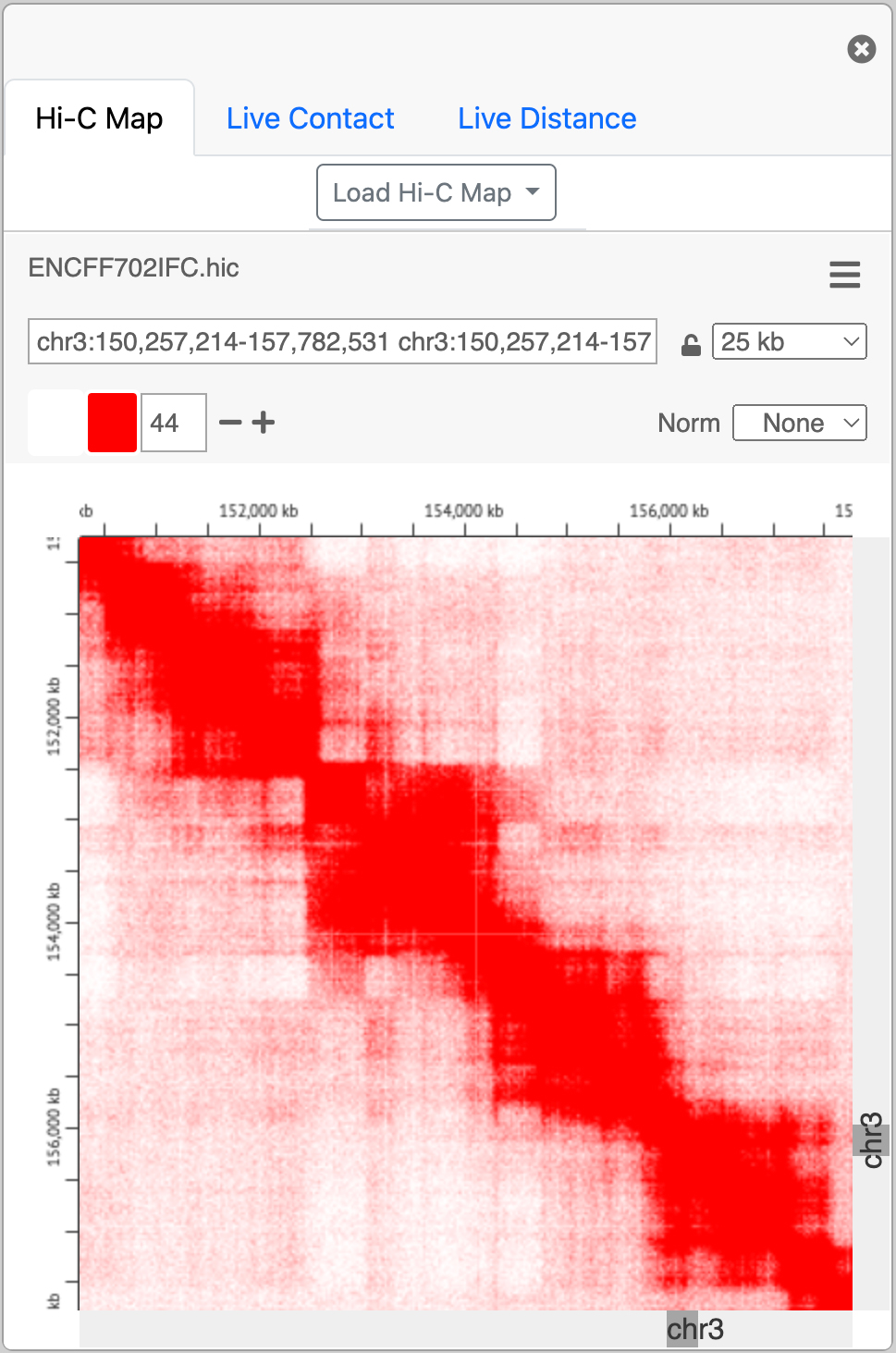
An embedded instance juicebox.js for viewing Hi-C maps
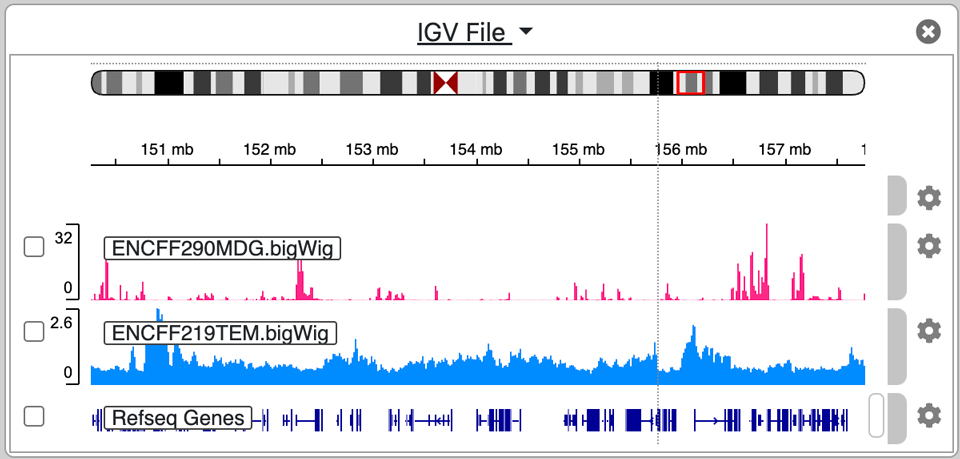
An embedded instance of the igv.js genome browser