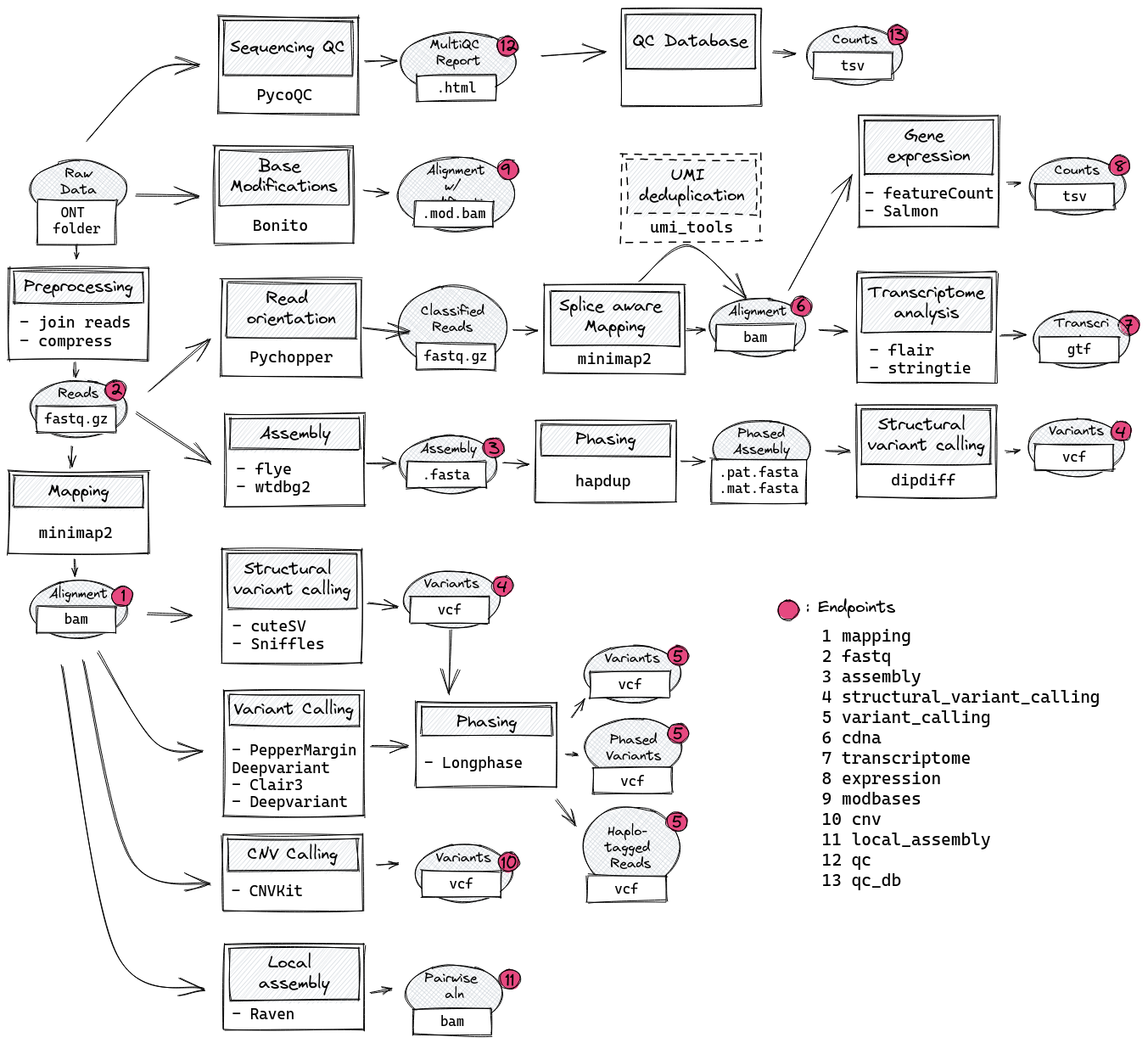
This is a Snakemake pipeline for comprehensive analysis of longread sequencing data. It is mainly intented for research and method establishment at the Institute of medicinal genetics and applied genomics (IMGAG) of UKT Tübingen but can be adapted easily to fit your requirements.
Detailed information for the transcriptome analysis can be found in the document at doc/transcriptome_analysis.md.
Dependencies:
- Snakemake (
pip install snakemake) - Conda
- Docker/Singularity for Variant Calling with Pepper-Margin-Deepvariant
For cDNA (Transcriptome) analysis you need to manually install the following tool and adjust the application paths in the config file:
All other required tools will be installed into generated Conda environments.
All configuration takes place in a config file (config.yml) located in the work directory. Look into the default settings (config/config_defaults) for a list of options. Most important is to to change the path to the genome reference and select required analysis steps.
The pipeline should be run in an analysis folder that will contain final and intermediate results. The location of the raw data is defined by the sample_run_table.tsv placed in the working dir. In the simplest form this is a two-column tab separated textfile containing sample names and data folder:
sample1
sample1
sample2
For more config options look into the input configuration.
Assuming you are in the working directory which contains a config.yml and a sample_run_table.tsv run the pipeline with:
snakemake --use-conda -s /path_to_repo/megLR/workflow/Snakefile