A R package for Grade of Membership (GoM) model fit and Visualization of counts data-
Kushal K Dey, Chiaowen Joyce Hsiao, Matthew Stephens
CountClust requires the following CRAN-R packages: maptpx, slam, ggplot2, cowplot, parallel along with the Bioconductor package: limma.
source("http://bioconductor.org/biocLite.R")
biocLite("CountClust")
For installing the working version of this package and loading the data required for this vignette, we use CRAN-R package devtools.
library(devtools)
install_github('kkdey/CountClust')
Then load the package with:
library(CountClust)
We load the single cell RNA-seq data due to Deng et al 2014. The data contains RNA-seq read counts for single cells at different stages of mouse embryo development (from zygote to blastocyst).
library(singleCellRNASeqMouseDeng2014)
deng.counts <- exprs(Deng2014MouseESC)
deng.meta_data <- pData(Deng2014MouseESC)
deng.gene_names <- rownames(deng.counts)
We apply the StructureObj function (which is a wrapper of the topics function in the maptpx package) for a vector of number of clusters, ranging from 2 to 7.
FitGoM(t(deng.counts),
K=c(3,6), tol=0.1,
path_rda="data/MouseDeng2014.FitGoM.rda")
This function will output a list, each element representing a GoM model fit output for a particular cluster number.
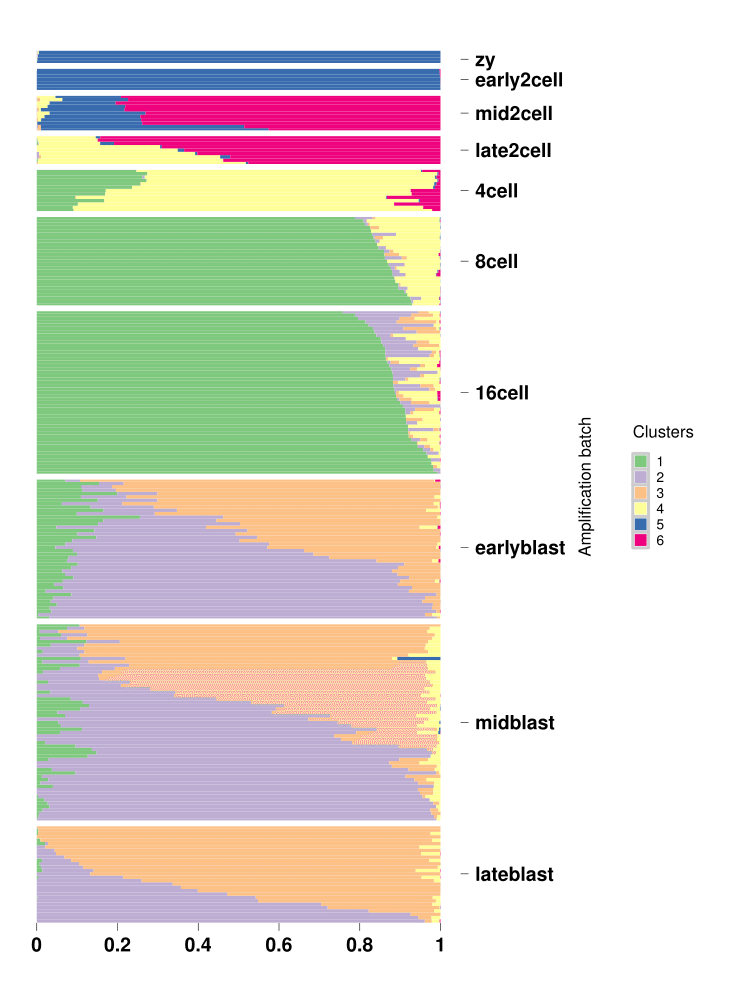
One can plot the omega from the StructureObj fit using a Structure plot. Here we provide an example of the Structure plot for K=6 for the above GoM model fit.
data("MouseDeng2014.FitGoM")
names(MouseDeng2014.FitGoM)
omega <- MouseDeng2014.FitGoM$clust_6$omega
annotation <- data.frame(
sample_id = paste0("X", c(1:NROW(omega))),
tissue_label = factor(rownames(omega),
levels = rev( c("zy", "early2cell",
"mid2cell", "late2cell",
"4cell", "8cell", "16cell",
"earlyblast","midblast",
"lateblast") ) ) )
rownames(omega) <- annotation$sample_id;
StructureGGplot(omega = omega,
annotation = annotation,
palette = RColorBrewer::brewer.pal(8, "Accent"),
yaxis_label = "Amplification batch",
order_sample = TRUE,
axis_tick = list(axis_ticks_length = .1,
axis_ticks_lwd_y = .1,
axis_ticks_lwd_x = .1,
axis_label_size = 7,
axis_label_face = "bold"))
We can extract the features that drive the clusters for K=6 as follows
theta_mat <- MouseDeng2014.topicFit$clust_6$theta;
top_features <- ExtractTopFeatures(theta_mat, top_features=100,
method="poisson", options="min");
gene_list <- do.call(rbind, lapply(1:dim(top_features)[1],
function(x) deng.gene_names[top_features[x,]]))
It will provide you with a list of top 100 variables/features per cluster that are relatively most highly expressed in that cluster compared to the other clusters, or in other words, plays the most important role in driving or separating out that cluster from the rest.
The CountClust package is distributed under [GPL - General Public License (>= 2)]
For any questions or comments, please contact kkdey@uchicago.edu
- Raman Shah