A deep learning based method to automatically identify miniature EPSPs events from electrophysiology data.
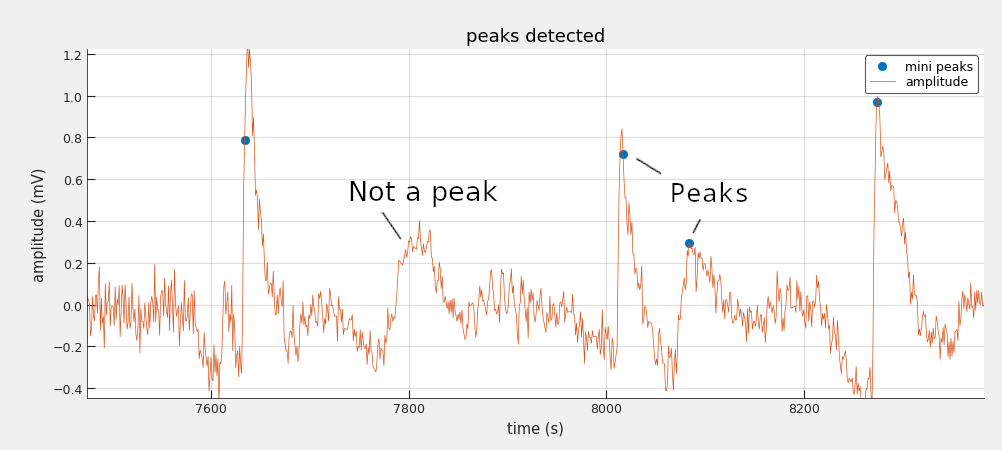
The input of the system is an electrophysiology experiment data and the output is the timestamp and amplitude of the miniature EPSPs events (aka mini peaks). As shown in the graph below it is not trivial to design a handcrafted algorithm to identify the mini peaks because the shape of the signal is the defining discriminator (the amplitude of the signal alone is not sufficient). This is why a deep learning approach has been chosen.
The main goal of this toy project is for me to apply what I learnt in the diverse deep learning courses on a real world problem from scratch.
Install pytorch following the official instructions.
From the root folder of this repository:
pip3 install .
Training is done with the script train_test_model.py which takes as input a folder containing the training data in csv files.
The csv files must contains a time serie of the electrophysiology data together with the presence or absence of mini peak for each timestep.
Here is the first four lines of a well formatted csv file:
time,amplitude,minis
0.1,-0.0909423828125,0.0
0.10200000000000001,-0.09246826171875,0.0
0.10400000000000001,-0.24658203125,0.0
- time is in second
- amplitude is in mV
- minis = 0.0 if no mini peak for this timestep; 1.0 if mini peak for this timestep
Please take a look at the minipeak.data submodule for an example of how to preprocess the raw electrophysiology data for training.
To train the network, the script train_test_model.py is used. Here is an example:
python minipeak/training/train_test_model.py example_data/ training_output/ --epochs 1000 --window-size 80
See train_test_model.py or do python minipeak/training/train_test_model.py -h to get the full list of available settings.
The output of the training is the following files structure that will be saved under the exp_folder path:
- exp_folder - root folder of the training outputs
- experiment.json - hyperparameters used to train the model and the test set validation results (loss, accuracy, precision, recall rate)
- model.pt - the saved weight of the trained model
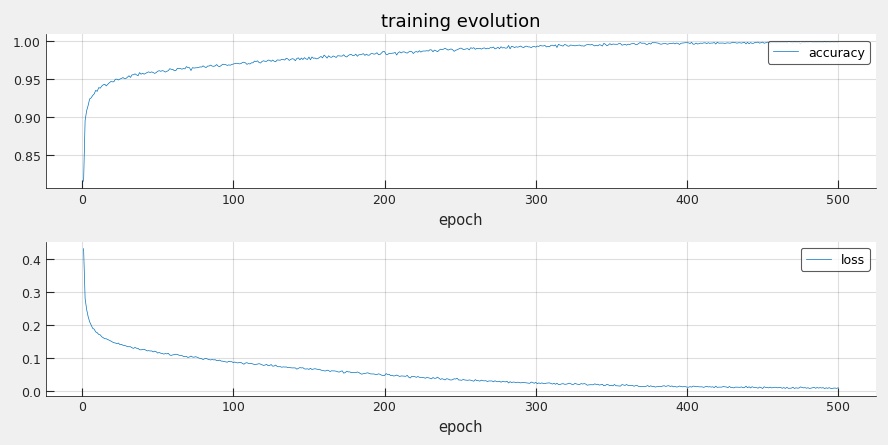
- training_results.csv - loss and accuracy evolution for each training epoch
- correct_positives - folder containing plots of all correctly detected mini peaks during test time
- false_negatives - folder containing plots of all mini peaks that don't have been detected during test time
- false_positives - folder containing plots of all wrongly detected mini peaks during test time
The inference just take the raw electrophysiology data in Axon Binary Format (ABF) file.
python minipeak/training/eval_model.py example_data/example_inference_data.abf minipeak/weights/model.pt --sampling_rate 100 --remove_trend_win_ms 100 --window-size 80
See eval_model.py or do python minipeak/training/eval_model.py -h to get the full list of available settings.
The output of the script is a csv file containing the list of detected mini peaks with timestamp (in seconds) and amplitude (in mV) for each mini peak.
minipeak- the python package of the projectdata- utility to extract relevant experiemental lab data for training.models- the neural network modelsstyles- matplotlib styles configurationtraining- the training and test loops script and the inference scriptvisualization- scripts to visualize the data and resultsweights- contains the trained weight files
TODO explain and pinpoint relevant code:
- 1D CNN choice and archi
- overlapping windows
- improve with transformer
Test set results:
- accuracy = 0.9430
- precision = 0.9102
- recall = 0.9128
- mini peak position mean error within widow <1ms