An R implementation of the Uniform Manifold Approximation and Projection (UMAP) method for dimensionality reduction (McInnes and Healy, 2018), that also implements the supervised and metric (out-of-sample) learning extensions to the basic method. Translated from the Python implementation.
December 10 2018. Default behavior has changed so that multiple threads are no
longer used during the stochastic gradient descent phase. This (along with some
other minor changes) should ensure reproducibility (as long as you set.seed
appropriately), at the cost of a loss in speed and is in line with the behavior
of the Python UMAP implementation. If you don't care about reproducibility, the
number of threads to use during SGD can be set with a new parameter,
n_sgd_threads. Set it to n_sgd_threads = "auto" to get the old behavior.
December 9 2018. Added a pca argument that will reduce X to the specified
number of dimensions (e.g. 50, commonly used in t-SNE routines). This should
give a big speed up to the nearest neighbor search if you are using Euclidean
distance metric and you have lots of features (where lots might be as little as
100-1000), for instance
COIL-100.
December 5 2018. Some deeply experimental mixed data type features are now
available: you can now mix different metrics (e.g. euclidean for some
columns and cosine for others). The type of data that can be used with y
in supervised UMAP has also been expanded. See
Mixed Data Types
for more details.
uwot makes use of C++ code which must be compiled. You may have to carry out
a few extra steps before being able to build this package:
Windows: install
Rtools and ensure
C:\Rtools\bin is on your path.
Mac OS X: using a custom ~/.R/Makevars
may cause linking errors.
This sort of thing is a potential problem on all platforms but seems to bite
Mac owners more.
The R for Mac OS X FAQ
may be helpful here to work out what you can get away with. To be on the safe
side, I would advise building uwot without a custom Makevars.
install.packages("devtools")
devtools::install_github("jlmelville/uwot")
library(uwot)
# See function man page for help
?umapiris_umap <- umap(iris, n_neighbors = 50, learning_rate = 0.5, init = "random")
# Load mnist from somewhere, e.g.
# devtools::install_github("jlmelville/snedata")
# mnist <- snedata::download_mnist()
mnist_umap <- umap(mnist, n_neighbors = 15, min_dist = 0.001, verbose = TRUE)
# Use a specific number of threads
mnist_umap <- umap(mnist, n_neighbors = 15, min_dist = 0.001, verbose = TRUE, n_threads = 8)
# Use a different metric
mnist_umap_cosine <- umap(mnist ,n_neighbors = 15, metric = "cosine", min_dist = 0.001, verbose = TRUE, n_threads = 8)
# Supervised dimension reduction
mnist_umap_s <- umap(mnist, n_neighbors = 15, min_dist = 0.001, verbose = TRUE, n_threads = 8,
y = mnist$Label, target_weight = 0.5)
# Add new points to an existing embedding
mnist_train <- head(mnist, 60000)
mnist_test <- tail(mnist, 70000)
# You must set ret_model = TRUE to return extra data we need
# coordinates are in mnist_train_umap$embedding
mnist_train_umap <- umap(mnist_train, verbose = TRUE, ret_model = TRUE)
mnist_test_umap <- umap_transform(mnist_test, mnist_train_umap, verbose = TRUE)
# Save the nearest neighbor data
mnist_nn <- umap(mnist, ret_nn = TRUE)
# coordinates are now in mnist_nn$embedding
# Re-use the nearest neighor data and save a lot of time
mnist_nn_spca <- umap(mnist, nn_method = mnist_nn$nn, init = spca)
# No problem to have ret_nn = TRUE and ret_model = TRUE at the same time
# Calculate Petal and Sepal neighbors separately (uses intersection of the resulting sets):
iris_umap <- umap(iris, metric = list("euclidean" = c("Sepal.Length", "Sepal.Width"),
"euclidean" = c("Petal.Length", "Petal.Width")))
# Can also use individual factor columns
iris_umap <- umap(iris, metric = list("euclidean" = c("Sepal.Length", "Sepal.Width"),
"euclidean" = c("Petal.Length", "Petal.Width"),
"categorical" = "Species"))
# MNIST with PCA reduction to 50 dimensions can speed up calculation without
# affecting results much
mnist_umap <- umap(mnist, pca = 50)Apart from the man pages in R: you may be interested in:
- A description of UMAP using algorithmic terminology similar to t-SNE, rather than the more topological approach of the UMAP publication.
- Examples of the output of UMAP on some datasets, compared to t-SNE.
- How to use UMAP for Supervised and Metric Learning.
For small (N < 4096) and Euclidean distance, exact nearest neighbors are found
using the FNN package. Otherwise,
approximate nearest neighbors are found using
RcppAnnoy. The supported
distance metrics (set by the metric parameter) are:
- Euclidean
- Cosine
- Manhattan
- Hamming
If you need other metrics, and can generate the nearest neighbor info
externally, you can pass the data directly to uwot via the nn_method
parameter. See the
Nearest Neighbor Data Format section
for more details.
Coordinate initialization uses RSpectra to do the eigendecomposition of the normalized Laplacian.
The optional PCA initialization and initial dimensionality reduction uses irlba.
The smooth k-nearest neighbor distance and stochastic gradient descent optimization routines are written in C++ (using Rcpp and RcppArmadillo), aping the Python code as closely as possible. It is my first time using Rcpp, so let's assume I did a horrible job.
For the datasets I've tried it with, the results look at least
reminiscent of those obtained using the
official Python implementation.
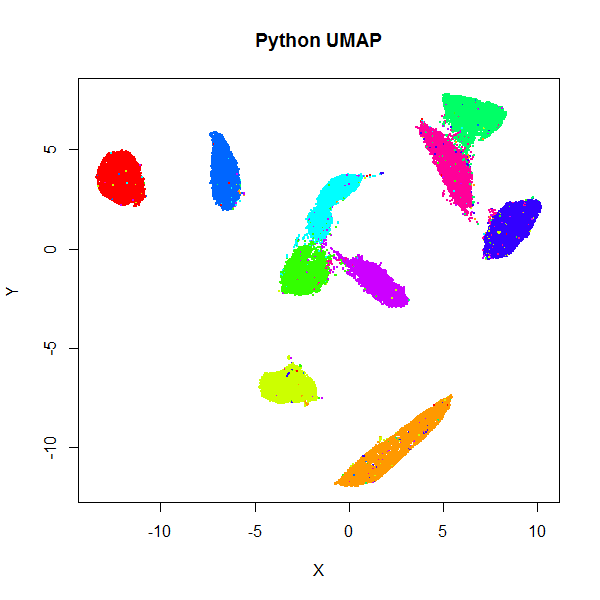
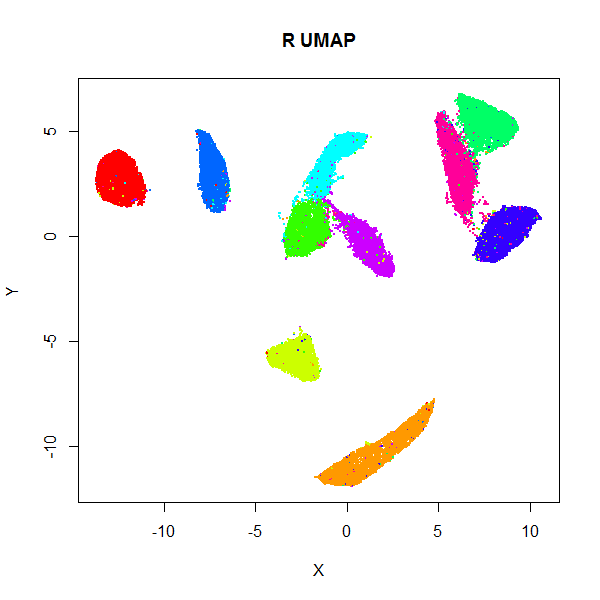
Below are results for the 70,000 MNIST digits (downloaded using the
snedata package). On the left
is the result of using the official Python UMAP implementation
(via the reticulate package).
The right hand image is the result of using uwot.
 |
 |
The project documentation contains some more examples.
To get a feel for the performance of uwot, here are some timings for
processing the MNIST dataset on my not-particularly-beefy laptop, compared with
some other packages with their default settings:
| Method | Time |
|---|---|
uwot(n_threads = 1) |
3.5 minutes |
| Barnes-Hut t-SNE (Rtsne 0.15) | 14 minutes |
| largeVis | 56 minutes |
| official LargeVis implementation | 10 minutes |
| UMAP (Python) | 2 minutes |
uwot(n_threads = 4) |
2 minutes |
uwot(n_threads = 1, approx_pow = TRUE) |
3 minutes |
uwot(n_threads = 4, approx_pow = TRUE) |
1.5 minutes |
The difference in performance between the Python UMAP (powered by the JIT-magic of
Numba) and uwot with one thread is due to:
- nearest neighbor search: takes 40 seconds in Python which also has the
experimental parallel support in Numba turned on, versus just over 2 minutes in
single-threaded
uwot. Using 4 threads for the index search part reduces this to 1 minute. This part is the performance bottleneck at the moment. The Python version of UMAP uses pynndescent, a nearest neighbor descent approach, rather than Annoy. Alternative nearest neighbors libraries e.g. kgraph (which is based on the same paper as pynndescent), or HNSW would be interesting to try, but all of the ones I've looked at either don't currently build on Windows or have non-portable compilation flags, so will require some fiddling with. - the optimization stage: takes 60 seconds in Python (no parallel option
here), versus about 66 seconds with
uwotwith one thread . I think the difference here is due to thepowoperations in the gradient. If you like living dangerously, you can try using thefastPrecisePowapproximation to thepowfunction suggested by Martin Ankerl:
# Set approx_pow = TRUE to use the approximation
mnist_umap <- umap(mnist, n_neighbors = 15, min_dist = 0.001, approx_pow = TRUE, verbose = TRUE)For what I think seem like typical values of b (between 0.7 and 0.9)
and the squared distance (0-1000), I found the maximum relative error was
about 0.06. However, I haven't done much testing, beyond looking to see that
the MNIST results are not obviously worsened. Results in the table above with
approx_pow = TRUE do show a worthwhile improvement.
I would welcome any further suggestions on improvements (particularly speeding up the optimization loop). However, it's certainly fast enough for my needs.
By the deeply unscientific method of me looking at how much memory the R session
was taking up according to the Task Manager, processing MNIST with four threads
saw the memory usage increase by nearly 1 GB at some points. There are some manual
calls to gc() after some stages to avoid holding onto unused memory for longer
than usual. The larger the value of n_neighbors, the more memory you can expect
to take up (see, for example, the discussion of the lvish function below).
RcppParallel is used for the nearest neighbor index search, the smooth knn/perplexity calibration, and the optimization, which is the same approach that LargeVis takes.
You can (and should) adjust the number of threads via the n_threads parameter;
for now, the default is half of whatever RcppParallel thinks should be the
default. I have also exposed the grain_size parameter. If a thread would
process less than grain_size number of items, then no multithreading is
carried out. Set n_threads = 0 to use the previous non-threaded search; with
n_threads = 1, you get the new multi-threaded code but with only one thread.
I've not experienced any problems with using multiple threads for a little while, but if you have any problems with crashing sessions, please file an issue.
- As noted in the
Implementation Details,
only Euclidean, Cosine, Hamming, and Manhattan distances are supported for finding
nearest neighbors from data frame and dense matrix input. For other metrics,
you can pass nearest neighbor data directly: see the
Nearest Neighbor Data Format
section. Or if you can calculate a distance matrix for your data, you can pass
it in as
distobject. For larger distance matrices, you can pass in asparseMatrix(from the Matrix package). Neither approach is supremely efficient at the moment. Proper sparse matrix support is limited by the nearest neighbor search routine: Annoy is intended for dense vectors. Adding a library for sparse nearest neighbor search would be a good extension. - For supervised dimensionality reduction using a numeric vector, only the Euclidean distance is supported for building the target graph. Again, see the Nearest Neighbor Data Format for a possible alternative.
- Even if you use
set.seed, results of the embeddings are not repeatable, unless you only use one thread (i.e. use the argumentn_threads = 1). This is because there is no locking carried out on the underlying coordinate matrix, and work is partitioned by edge not vertex and a given vertex may be processed by different threads. The order in which reads and writes occur is of course at the whim of the thread scheduler. - I haven't applied
uwoton anything much larger than MNIST and Fashion MNIST (so at least around 100,000 rows with 500-1,000 columns works fine). Bear in mind that Annoy itself says it works best with dimensions < 100, but still works "surprisingly well" up to 1000. - The spectral initialization default for
umap(and the Laplacian eigenmap initialization,init = "laplacian") can sometimes run into problems. If it fails to converge it will fall back to random initialization, but on occasion I've seen it take an extremely long time (a couple of hours) to converge. If initialization is taking more than a few minutes, I suggest stopping the calculation and using the scaled PCA (init = "spca") instead. R CMD checkcurrently reports the following note:GNU make is a SystemRequirements., which is expected and due to using RcppParallel. On Linux, it sometimes notes that thelibssub-directory is over 1 MB. I am unsure if this is anything to worry about.
Some other dimensionality reduction methods are also available in uwot:
If you choose the UMAP curve parameters to be a = 1 and b = 1, you get
back the Cauchy distribution used in
t-Distributed Stochastic Neighbor Embedding
and LargeVis. This also happens to
significantly simplify the gradient leading to a noticeable speed-up: for MNIST,
I saw the optimization time drop from 66 seconds to 18 seconds. The trade off is
that you will see larger, more spread-out clusters than with the typical UMAP
settings (they're still more compact than you see in t-SNE, however). To try
t-UMAP, use the tumap function:
mnist_tumap <- tumap(mnist, n_neighbors = 15, verbose = TRUE)Note that using umap(a = 1, b = 1) doesn't use the simplified gradient, so
you won't see any speed-up that way.
As UMAP's implementation is similar to LargeVis in some respects, this package
also offers a LargeVis-like method, lvish:
# perplexity, init and n_epoch values shown are the defaults
# use perplexity instead of n_neighbors to control local neighborhood size
mnist_lv <- lvish(mnist, perplexity = 50, init = "lvrand", n_epochs = 5000,
verbose = TRUE)
# Make hilarious Lembas bread jokeAlthough lvish is like the real LargeVis in terms of the input weights, output
weight function and gradient, and so should give results that resemble the real
thing, note that:
- Like the real LargeVis, matrix input data is normalized by centering each
column and then the entire matrix is scaled by dividing by the maximum absolute
value. This differs from
umap, where no scaling is carried out. Scaling can be controlled by thescaleparameter. - Nearest neighbor results are not refined via the neighbor expansion method.
The
search_kparameter is twice as large than Annoy's default to compensate. - The other nearest neighbor index parameter,
n_trees, is not dynamically chosen based on data set size. In LargeVis, it ranges between 10 (for N < 100,000) and 100 (for N > 5,000,000). Thelvishdefault of 50 would cover datasets up to N = 5,000,000, and combined with the defaultsearch_k, seems suitable for the datasets I've looked at. - Negative edges are generated by uniform sampling of vertexes rather than their degree ^ 0.75.
- The default number of epochs is dataset-dependent, to generate the same number
of edge samples that would be used by the default settings of the reference
LargeVis implementation. This normally results in a substantially longer run
time than for
umap. You may be able to get away with fewer epochs, and using the UMAP initialization ofinit = "spectral", rather than the default Gaussian random initialization (init = "lvrand") can help.
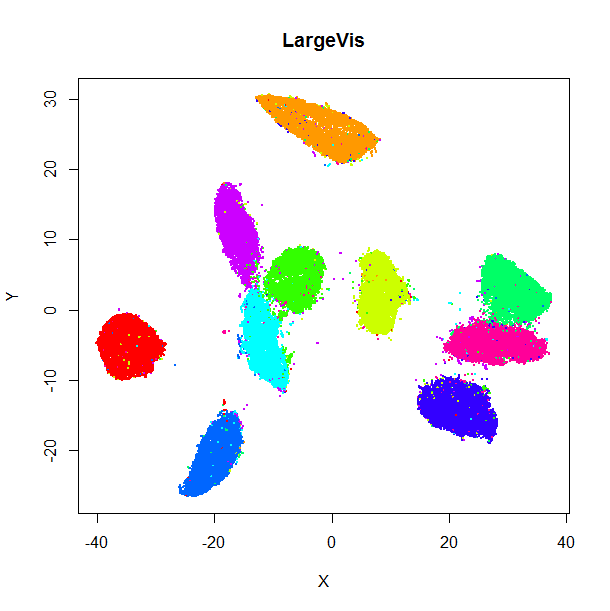
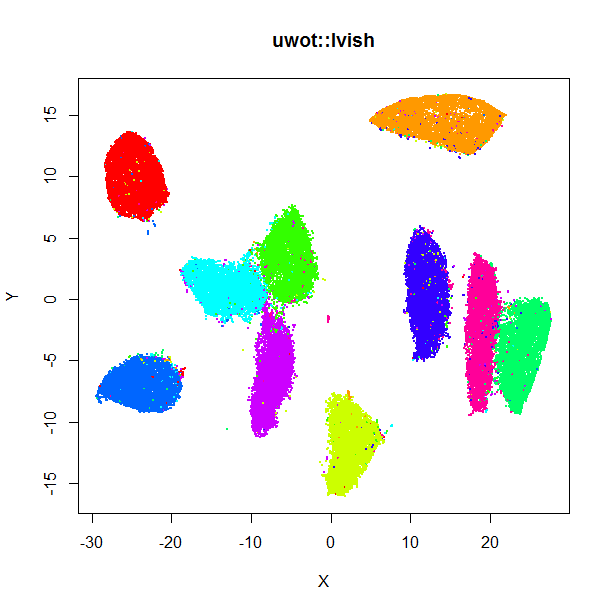
The left-hand image below is the result of running the official LargeVis
implementation on MNIST. The image on the right is that from running lvish
with its default settings (apart from setting n_threads = 8). Given they were
both initialized from different random configurations, there's no reason to
believe they would be identical, but they look pretty similar:
 |
 |
Because the default number of neighbors is 3 times the perplexity, and the
default perplexity = 50, the nearest neighbor search needs to find 150 nearest
neighbors per data point, an order of magnitude larger than the UMAP defaults.
This leads to a less sparse input graph and hence more edges to sample. Combined
with the increased number of epochs, expect lvish to be slower than umap:
with default single-threaded settings, it took about 20 minutes to embed the
MNIST data under the same circumstances as described in the "Performance"
section. With n_threads = 4, it took 7 minutes. In addition, storing those
extra edges requires a lot more memory than the umap defaults: my R session
increased by around 3.2 GB, versus 1 GB for umap.
As an alternative to the usual Gaussian input weight function, you can use the
k-nearest neighbor graph itself, by setting kernel = "knn". This will give
each edge between neighbors a uniform weight equal to 1/perplexity, which
leads to each row's probability distribution having the target perplexity.
This matrix will then be symmetrized in the usual way. The advantage of this is
that the number of neighbors is reduced to the same as the perplexity (indeed,
the n_neighbors parameter is ignored with this setting), and leads to less
memory usage and a faster runtime. You can also get away with setting the
perplexity to a much lower value than usual with this kernel (e.g. perplexity = 15) and get closer to UMAP's performance. If you use the default LargeVis
random initialization, you will still need more epochs than UMAP, but you can
still expect to see a big improvement. Something like the following works for
MNIST:
mnist_lv <- lvish(mnist, kernel = "knn", perplexity = 15, n_epochs = 1500,
init = "lvrand", verbose = TRUE)The default approach of UMAP is that all your data is numeric and will be
treated as one block using the Euclidean distance metric. To use a different
metric, set the metric parameter, e.g. metric = "cosine".
Treating the data as one block may not always be appropriate. uwot now
supports a highly experimental approach to mixed data types. It is not based on
any deep understanding of topology and sets, so consider it subject
to change, breakage or completely disappearing.
To use different metrics for different parts of a data frame, pass a list to
the metric parameter. The name of each item is the metric to use and the
value is a vector containing the names of the columns (or their integer id, but
I strongly recommend names) to apply that metric to, e.g.:
metric = list("euclidean" = c("A1", "A2"), "cosine" = c("B1", "B2", "B3"))this will treat columns A1 and A2 as one block of data, and generate
neighbor data using the Euclidean distance, while a different set of neighbors
will be generated with columns B1, B2 and B3, using the cosine distance.
This will create two different simplicial sets. The final set used for
optimization is the intersection of these two sets. This is exactly the same
process that is used when carrying out supervised UMAP (except the contribution
is always equal between the two sets and can't be controlled by the user).
You can repeat the same metric multiple times. For example, to treat the
petal and sepal data separately in the iris dataset, but to use Euclidean
distances for both, use:
metric = list("euclidean" = c("Petal.Width", "Petal.Length"),
"euclidean" = c("Sepal.Width", "Sepal.Length"))As the iris example shows, using column names can be very verbose. Integer
indexing is supported, so the equivalent of the above using integer indexing
into the columns of iris is:
metric = list("euclidean" = 3:4, "euclidean" = 1:2)but internally, uwot strips out the non-numeric columns from the data, and if
you use Z-scaling (i.e. specify scale = "Z"), zero variance columns will also
be removed. This is very likely to change the index of the columns. If you
really want to use numeric column indexes, I strongly advise not using the
scale argument and re-arranging your data frame if necessary so that all
non-numeric columns come after the numeric columns.
Supervized UMAP allows for a factor column to be used. You may now also specify
factor columns in the X data. Use the special metric name "categorical".
For example, to use the Species factor in standard UMAP for iris along
with the usual four numeric columns, use:
metric = list("euclidean" = 1:4, "categorical" = "Species")Factor columns are treated differently from numeric columns:
- They are always treated separately, one column at a time. If you have two
factor columns,
cat1, andcat2, and you would like them included in UMAP, you should write:
metric = list("categorical" = "cat1", "categorical" = "cat2", ...)As a convenience, you can also write:
metric = list("categorical" = c("cat1", "cat2"), ...)but that doesn't combine cat1 and cat2 into one block, just saves some
typing.
- Because of the way categorical data is intersected into a simplicial set, you
cannot have an X
metricthat specifies onlycategoricalentries. You must specify at least one of the standard Annoy metrics for numeric data. Foriris, the following is an error:
# wrong and bad
metric = list("categorical" = "Species")Specifying some numeric columns is required:
# OK
metric = list("categorical" = "Species", "euclidean" = 1:4)- Factor columns not explicitly included in the
metricare still removed as usual. - Categorical data does not appear in the model returned when
ret_model = TRUEand so does not affect the project of data used inumap_transform. You can still use the UMAP model to project new data, but factor columns in the new data are ignored (effectively working like supervized UMAP).
The handling of y data has been extended to allow for data frames, and
target_metric works like metric: multiple numeric blocks with different
metrics can be specified, and categorical data can be specified with
categorical. However, unlike X, the default behavior for y is to include
all factor columns. Any numeric data found will be treated as one block, so if
you have multiple numeric columns that you want treated separately, you should
specify each column separately:
target_metric = list("euclidean" = 1, "euclidean" = 2, ...)I suspect that the vast majority of y data is one column, so the default
behavior will be fine most of the time.
The Python implementation of UMAP supports lots of distance metrics; uwot does
not, because it depends on the distance metrics supported by RcppAnnoy, which
in turn depends on those supported by Annoy. For more flexibility, at the
cost of convenience, you can generate nearest neighbor data for X by some
other means and pass that to umap (or tumap or lvish) directly via the
nn_method parameter.
The format expected by nn_method is a list containing the following two
entries:
idx: a matrix of dimensionn_vertices x n_neighbors, where each row contains the indexes (starting at1) of the nearest neighbors of each item (vertex) in the dataset. Each item is always the nearest neighbor of itself, so the first element in rowishould always bei. If it isn't then either you are using a really weird non-metric distance or your approximate nearest neighbor method is returning way too approximate results. In either case, you should expect bad results.dist: a matrix of dimensionn_vertices x n_neighbors, where each row contains the distances of the nearest neighbors of each item (vertex) in the dataset, in Each item is always the nearest neighbor of itself, so the first element in rowishould always be0.0.
If you use pre-computed nearest neighbor data, be aware that:
- You can't use pre-computed nearest neighbor data and also use
metric. - You can explicitly set
Xto NULL, as long as you don't try and use an initialization method that makes use ofX(init = "pca"orinit = "spca"). - Setting
ret_model = TRUEdoes not produce a valid model.umap_transformwill not work with this setting.
If you set ret_nn = TRUE, the return value of umap will be a list, and the
nn item contains the nearest neighbor data in a format that can be used
with nn_method. This is handy if you are going to be running UMAP multiple
times with the same data and n_neighbors and scale settings, because the
nearest neighbor calculation can be the most time-consuming part of the
calculation.
Normally the contents of nn is itself a list, the value of which is the
nearest neighbor data. The name is the type of metric that generated the data.
As an example, here's what the first few items of the iris 5-NN data should
look like:
lapply(umap(iris, ret_nn = TRUE, n_neighbors = 5)$nn$euclidean, head)
$`idx`
[,1] [,2] [,3] [,4] [,5]
[1,] 1 18 5 40 29
[2,] 2 35 46 13 10
[3,] 3 48 4 7 13
[4,] 4 48 30 31 3
[5,] 5 38 1 18 41
[6,] 6 19 11 49 45
$dist
[,1] [,2] [,3] [,4] [,5]
[1,] 0 0.1000000 0.1414214 0.1414214 0.1414214
[2,] 0 0.1414214 0.1414214 0.1414214 0.1732051
[3,] 0 0.1414214 0.2449490 0.2645751 0.2645751
[4,] 0 0.1414214 0.1732051 0.2236068 0.2449490
[5,] 0 0.1414214 0.1414214 0.1732051 0.1732051
[6,] 0 0.3316625 0.3464102 0.3605551 0.3741657If for some reason you specify ret_nn while supplying precomputed nearest
neighbor data to nn_method, the returned data should be identical to what
you passed in, and the list item names will be precomputed.
As discussed under the
Mixed Data Types section,
you can apply multiple distance metrics to different parts of matrix or
data frame input data. if you do this, then ret_nn will return all the
neighbor data. The list under nn will now contain as many items as metrics,
in the order they were specified. For instance, if the metric argument is:
metric = list("euclidean" = c("Petal.Width", "Petal.Length"),
"cosine" = c("Sepal.Width", "Sepal.Length"))The nn list will contain two list entries. The first will be called
euclidean and the second cosine.
If you have access to multiple distance metrics, you may also provide multiple
precomputed neighbor data to nn_method in the same format: a list of lists,
where each sublist has the same format as described above (i.e. the two
matrices, idx and dist). The names of the list items are ignored, so you
don't need to set them. Roughly, do something like this:
nn_metric1 <- list(idx = matrix(...), dist = matrix(...))
nn_metric2 <- list(idx = matrix(...), dist = matrix(...))
umap_res <- umap(nn_method = list(nn_metric1, nn_metric2), ...)The different neighbor data must all have the same number of neighbors, i.e. the number of columns in all the matrices must be the same.
If you are using supervised UMAP with a numeric y, then you can also pass
nearest neighbor data to y, using the same format as above. In this case the
nearest neighbors should be with respect to the data in y.
Note that you cannot pass categorical y as nearest neighbor data. This is
because the processing of the data goes through a different code path that
doesn't directly calculate nearest neighbors: if y is a factor, when there
are only a small number of levels, the number of neighbors of an item can be
vastly larger than n_neighbors.
Nearest neighbor data for y is not returned from umap for re-use.
- The UMAP reference implementation and publication.
- There is now a UMAP package on CRAN (see also its github repo). Another R package is umapr.
uwotuses the RcppProgress package to show a text-based progress bar whenverbose = TRUE.- My somewhat convoluted method to ensure the C++ random numbers are repeatable
makes use of a (not convoluted)
get_seedfunction suggested in a blog post by Rory Nolan.
