DRAX is currently under active development -- please check back soon.
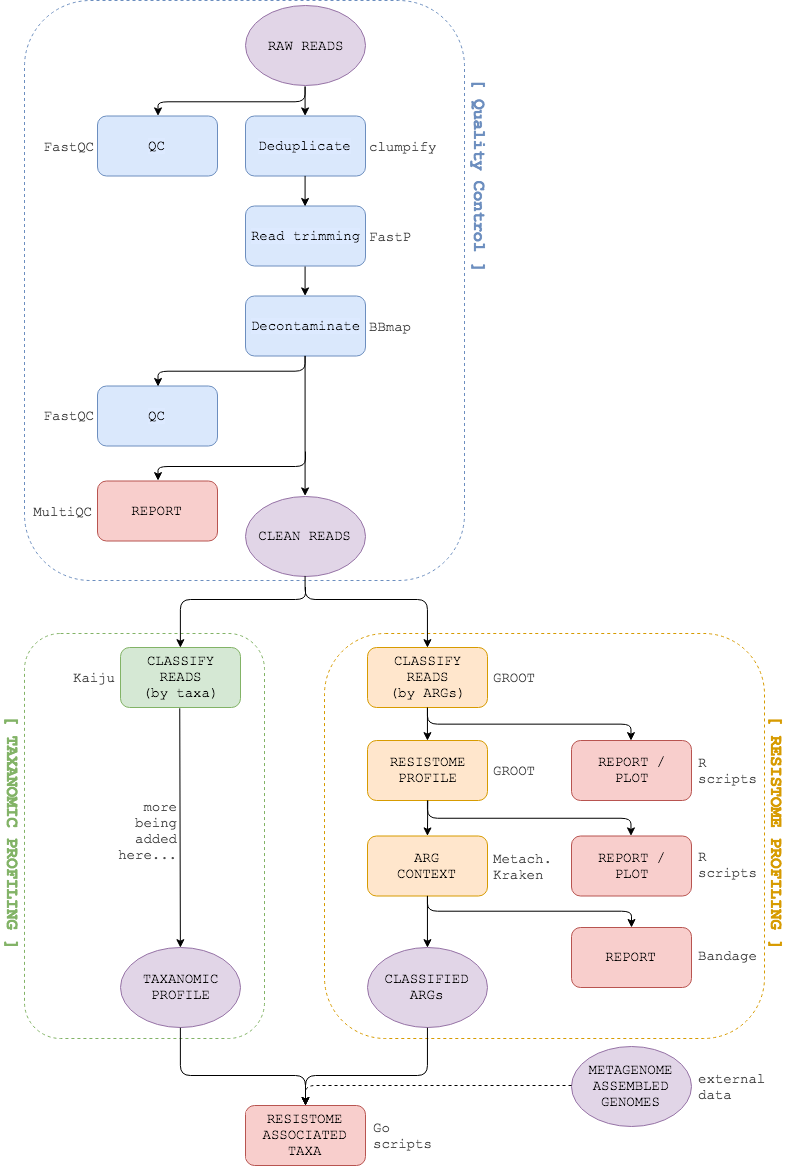
DRAX is a Nextflow pipeline to detect resistome associated taxa in metagenomes. It identifies Antimicrobial Resistance Genes (ARGs) using GROOT, extracts the genomic environment of each ARG using Metacherchant and then performs taxonomic classification of assembled environments. Further pipeline steps and full documentation are being added.
For ease of use, the DRAX pipeline is called by a wrapper script that allows you to either download the required reference data, run the DRAX Nextflow pipeline or update the pipeline. It is easiest to install DRAX using Bioconda:
conda install drax -c bioconda
You can also run DRAX using containers instead of conda. This will bypass the wrapper script so you will need to download the required reference data yourself.
# Install Nextflow
curl -s https://get.nextflow.io | bash
# Run DRAX with Docker
./nextflow run will-rowe/drax --reads 'tests/*R{1,2}.fq.gz' -profile docker
# OR run DRAX with Singularity
./nextflow run will-rowe/drax --reads 'tests/*R{1,2}.fq.gz' -with-singularity 'docker://wpmr/drax'
The first time you use DRAX, you will need to get the required data:
drax get
- this will download several databases (masked hg19 for decontamination, ResFinder for GROOT, Kaiju RefSeq DB...)
- it can take a little while (databases total ~10GB compressed)
- you only need to do this once
- the default directory is
./DRAX-filesbut you can rename/move it later - this command will also update the pipeline to the most current version
Now you can run DRAX:
drax --reads 'tests/*R{1,2}.fq.gz' --refData ./DRAX-files
For a full list of options, just run:
drax --help
Also, read the documentation for a more detailed overview of DRAX.
The DRAX pipeline comes with documentation about the pipeline, found in the docs directory. This is still being added to as the pipeline is developed.