pyGEMMA: A Fast, User-Friendly Python/R Implementation of Linear Mixed Models for Genome-Wide Association Studies
The current implementation of pyGEMMA was tested on the following configuration:
- Ubuntu 18.04.6 LST (Bionic Beaver)
- gcc/g++ 7.5.0 (Ubuntu 7.5.0-3ubuntu1~18.04)
- Python 3.7.15
- Numpy 1.21.6
- Cython 0.29.33
- Pandas 1.3.5
- Scipy 1.5.0
- Scikit-learn 0.24.2
- Matplotlib 3.3.0
- Seaborn 0.12.2
- rich 13.3.2
- qnorm 0.8.1
The installation of pyGEMMA is straightforward and can be performed using Python's pip package manager. Here, we detail the installation process using a virtualenv Python enviroment. This has been tested with the configuration listed in the Software Requirements section. While installation may be possible with other configurations, we it has only been tested with the configuration we list.
- Create your Python environment and activate it. Using
virtualenv, this can be done by running
pip3 install virtualenv
python3 -m virtualenv pygemma_env
source pygemma_env/bin/activateNote: If pip3 install virtualenv fails because it can't find pip3, you can try running python3 -m pip install virtualenv instead. This looks for the pip module directly if pip3 isn't in your PATH.
- Ensure the
NumpyandCythonpackages are both installed prior to installingpyGEMMA(they will not be installed automatically). This can be done by running
pip install numpy Cython- Clone this repository.
- Install
pyGEMMA's dependencies. From thepygemmadirectory, this can be done by running
pip install -r requirements.txt-
Ensure that you have a valid
C/C++compiler loaded.pyGEMMAhas been tested usinggcc/g++. -
Install
pyGEMMA. From thepygemmadirectory, this can be done by running
python setup.py installThe pyGEMMA package contains both high-level and low-level functions for fitting the linear mixed model outlined in the original GEMMA paper by Zhou et al. (Nat Gen 2012).
The pyGEMMA package is designed to fit the same model as GEMMA. That is, it fits
where
This model can be fit using the function pygemma.lmm.pygemma.
from pygemma import lmm
lmm.pygemma(Y, X, W, K, snps=snps, verbose=1)Note that snps is a list of SNP names that will be used to label the pandas DataFrame returned by the function. verbose controls whether to output run progress.
We have also developed an R interface for pyGEMMA, enabling its use within the R programming environment. A comprehensive tutorial for this integration can be found here
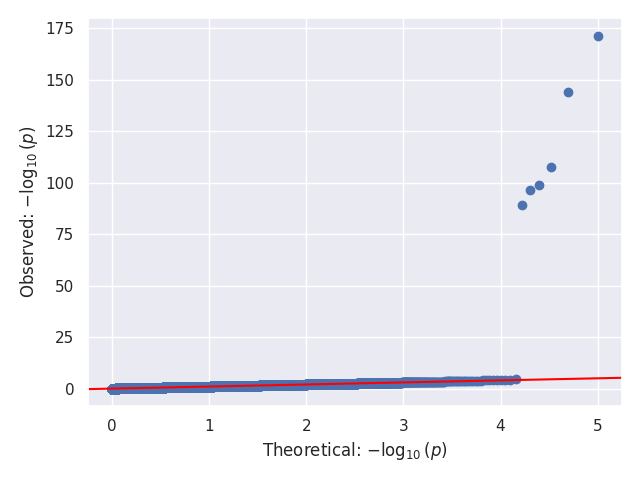
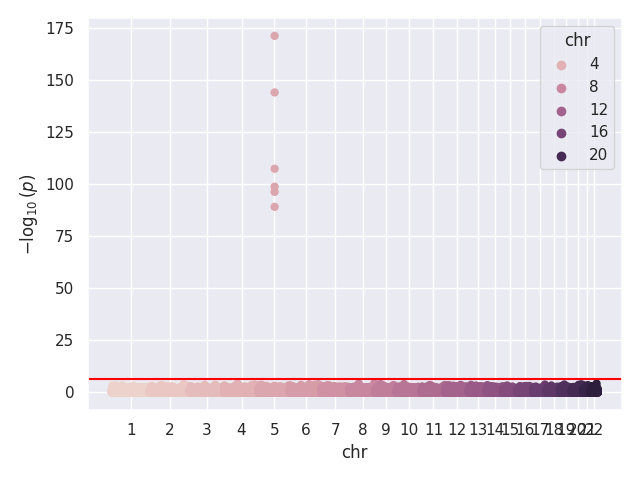
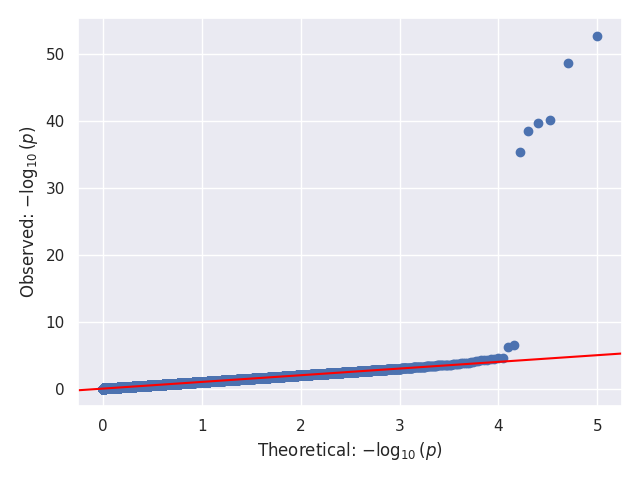
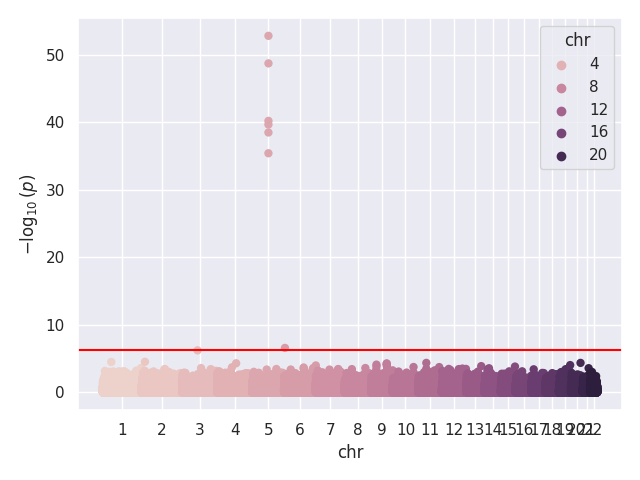
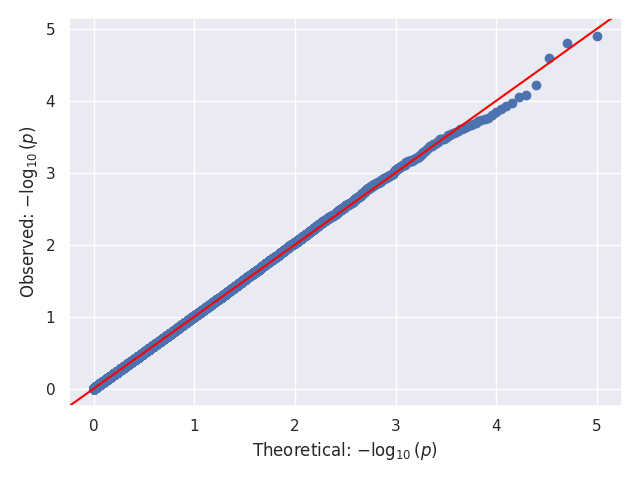
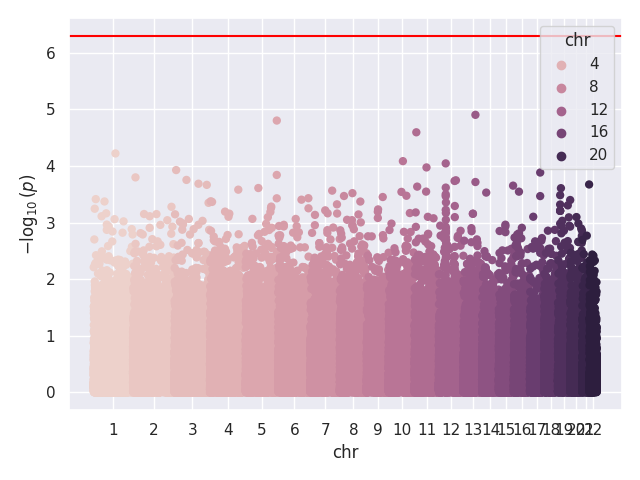
We provide a test script designed to test almost all basic functions and to run on three GWAS test cases (10,000 SNPs each). Before using the script, the paths for the data should be updated, as they are currently hardcoded (to be changed later). In our tests, this script took around 20 to 25 minutes to run on 32 cores. Below, we show the Q-Q and Manhattan plots for the three test cases.
 |
 |
 |
|
 |
|
We provide a number of scripts to run eQTL analyses for over 7100 genes using data from 1000 Genomes. These scripts are provided here.
The jobs may be launched by batching the run_pyGEMMA.sh script. Note that all files in the directory should be examined and paths modified before running. The scripts were designed to run using the SLURM scheduler.
@rlangefe - Robert Langefeld (Department of Biostatistics - University of Michigan)
If you have any questions or comments, please feel free to contact me.
