Small pipeline to wavelet transform 3D-Stacks (time,Y,X) based on the analysis tools provided by pyBOAT. The supplied input movie gets analyzed pixel by pixel along the time axis, yielding up to four output movies:
- phase movie
- period movie
- amplitude movie
- power movie
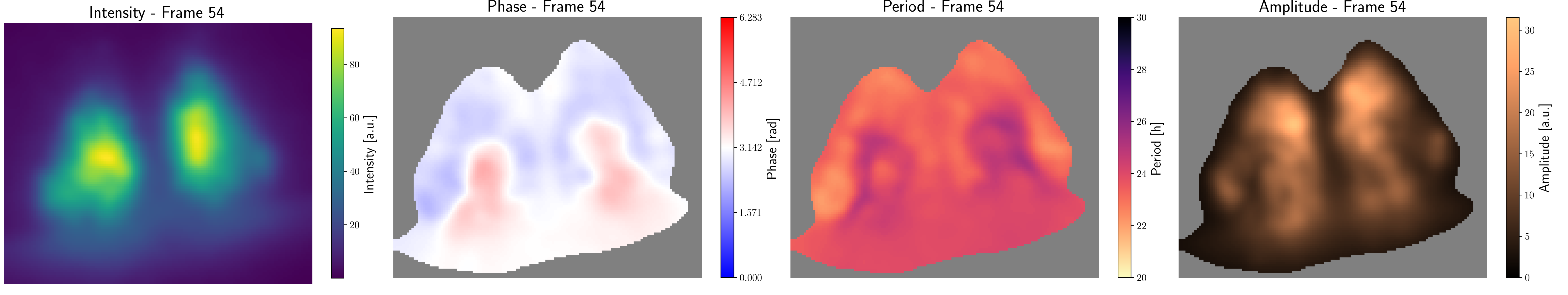
A snapshot of a typical output of the pipeline might look like this:
From left to right: Intensity of the blurred and down-sampled input; Phasefield; Periods and Amplitude
This is the complete 2D-analogue to the results for univariate time-series provided by pyBOAT.
For new users and/or exploratory data analysis it is advisable to first analyse at least a few 1D time-series (e.g. ROI tracks) with pyBOAT, which gets automatically installed along SpyBOAT. This allows to sanity check the key parameters of the time-frequency analysis:
- period range
- sinc filter cut-off period
- amplitude normalization window size (optional)
- sampling interval
before starting the computationally costly 2D stack transforms with SpyBOAT.
Typically bio-image data aquired by tissue microscopy has much higher spatial resolution than temporal resolution. To save computing time and ressources, it is advisable to spatially downscale the input movie. SpyBOAT offers a simple rescaling based on a scaling factor in %.
Often not everywhere in space an oscillatory signal can be found, hence masking the results after the transformation is an important step. SpyBOAT provides some simple masking capabilities, but the user is welcome to work with the raw output and devise a masking strategy most suitable for her needs.
This repository also provides a Fiji plugin in /local-scripts/mask_wmovie_ij.py, which allows for interactive
masking.
Soon..!
See the /galaxy folder.
Install SpyBOAT via pip:
bash pip3 install spyboat
or conda (soon..):
conda config --add channels conda-forge
conda install spyboatThe scripting-template.py in the root of this repository
showcases the analysis with the example spyboat.datasets.