Structure template for computational research project
- Write README template in cookiecutter folder
- look into rclone/rsync scripts/automation
- create Rmarkdown template
- Update this document according to my changes
Fantastic Work by Albert Henry
- Prepare the pre-requisites softwares and accounts above
- Generate a new GitHub personal access token, fill in the Note field and tick repo box under Select scopes, copy the generated token
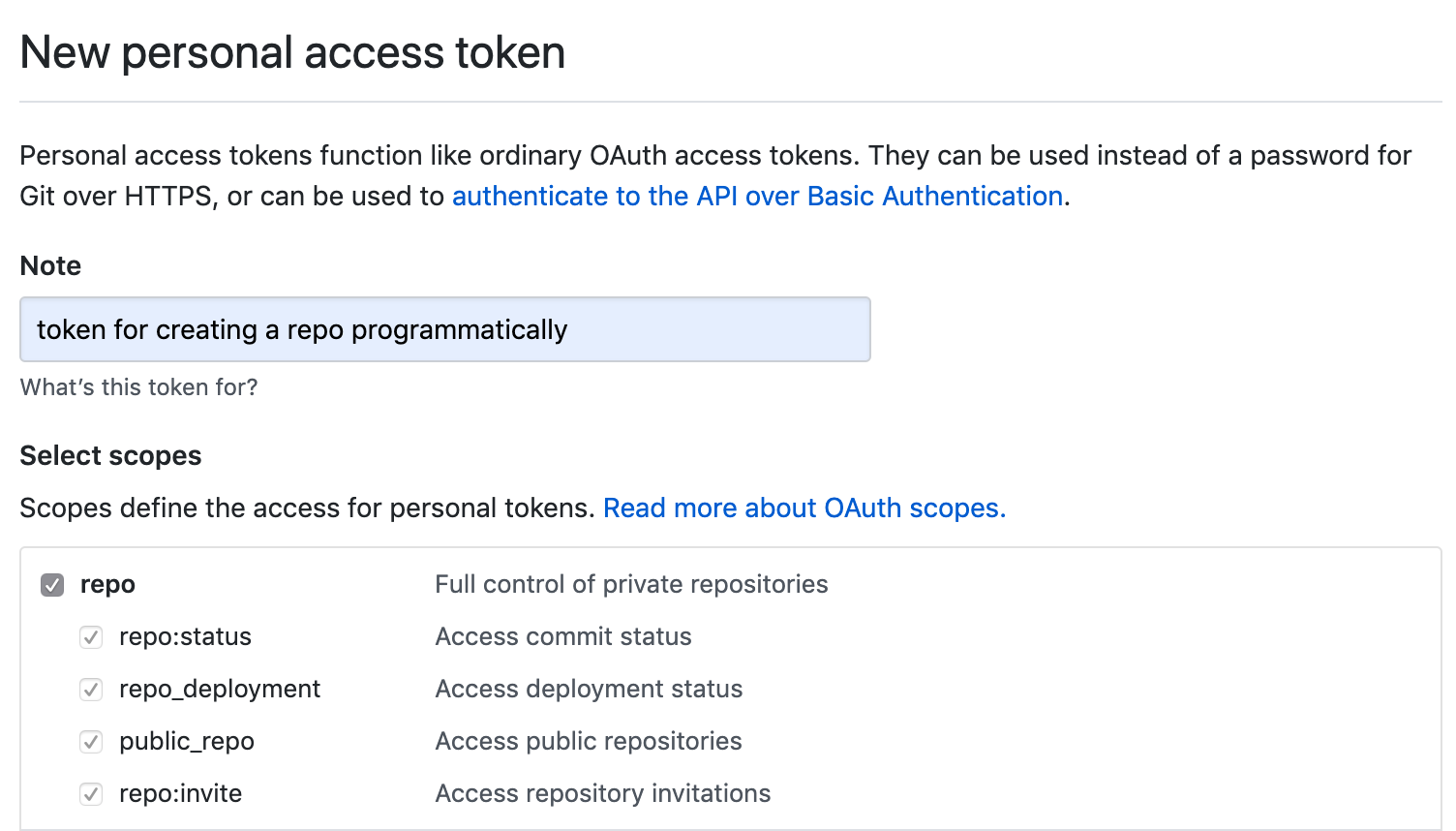
-
Open Command Line Interface (e.g. Terminal in Mac)
-
Change directory to parent project directory
cd my_directory -
Run
cookiecutterpointing to project template git repocookiecutter https://github.com/stef-mueller/project_template -
When prompted, enter the project title, project directory name, GitHub username, GitHub repository name (make sure name is available), and GitHub personal access token. Leave blank to use the default value (shown in square bracket).
The default project structure is outlined below:
./
├── .git/
|
├── admin/
| └── meeting_notes.csv
|
├── code/
| ├── s01_data_generation.sh
| |── s02.1_calculate_desc_stats_table1.R
| └── s02.2_figure1.ipynb
|
├── data/
| ├── input/
| └── output/
| |
├── exploratory/
|
├── results/
| ├── table1.tsv
| └── figure1.png
|
├── writing/
| ├── main_text.doc
| ├── supplementary_table.xls
| └── analysis_plan.doc
|
└── ...
- .git - hidden folder for git version control
- admin - Admin documents, e.g. meeting notes, applications, ethical approvals, MTA
- code - Analysis scripts
- data - data files used as input for analysis and results
- exploratory - Scratch space for temporary output files / exploratory data analysis generated as part of the analysis, can be deleted before archiving
- results - output of data analysis e.g. tables, figures, jupyter notebook
- writing - Analysis write ups, subfolders can be created specifically for early analysis drafts and later on manuscript drafts and final editions ready for submission to specific journals (this can also include reviewer comments and reply)