Implementation for EMNLP 2022 paper:
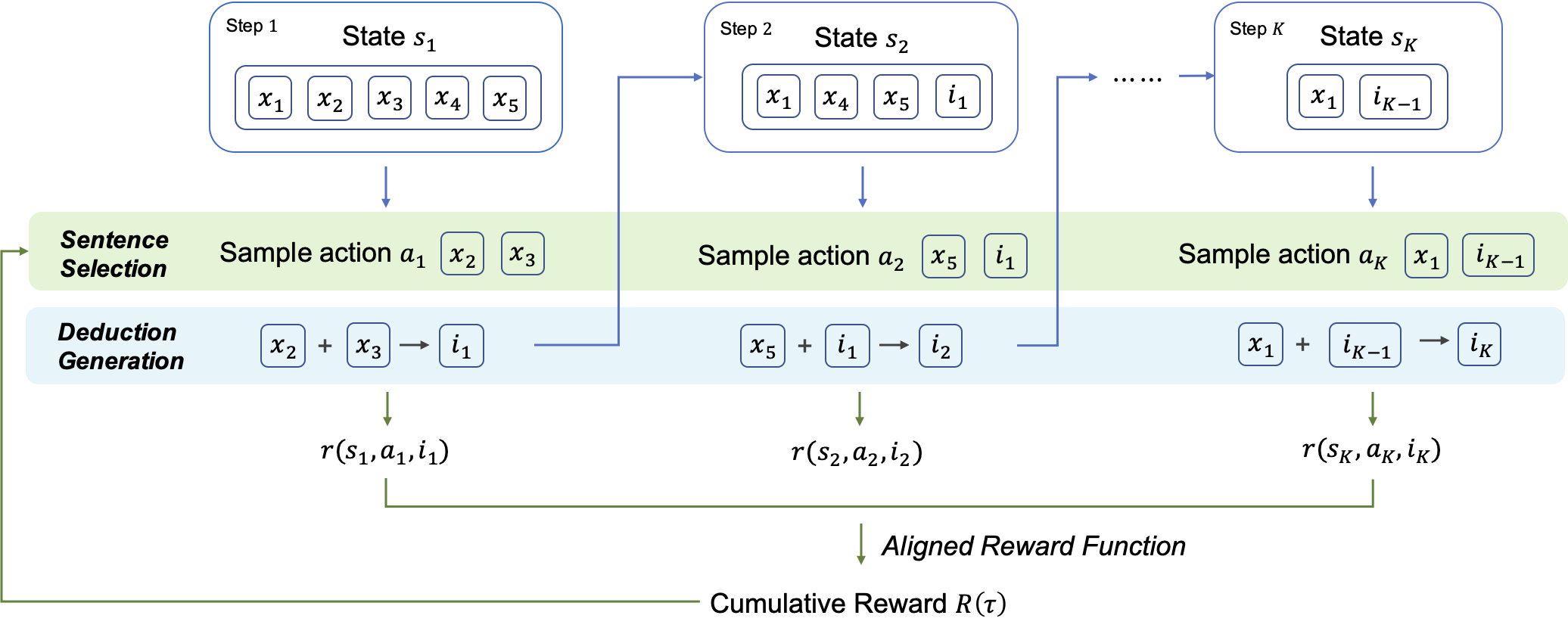
RLET: A Reinforcement Learning Based Approach for Explainable QA with Entailment Trees
Install all the dependencies in the requirements.txt file:
pip -r requirements.txt
Install BLEURT
BLEURT
pip install --upgrade pip # ensures that pip is current
git clone https://github.com/google-research/bleurt.git
cd bleurt
pip install .
Install spacy
python3 -m spacy download en_core_web_sm
Download the dataset from EntailmentBank and put it in the data folder. The folder should look like this:
├── data
│ ├── entailment_trees_emnlp2021_data_v3
│ │ ├── dataset
│ │ │ ├── ...
│ │ └── supporting_data
│ │ └── ...
│ ├── prepare_warmup_data.py
└── └── prepare_warmup_data_task23.py
Run the following command to preprocess the data for warmup training stage:
bash scripts/prepare_data.sh
That should generate the data at data/warmup/task1 and data/warmup/task23.
Before training, please initiate fitlog by running the following command:
fitlog init
Run the following command to start training:
# Task 1
bash scripts/run_warmup.sh
# Task 2/3
bash scripts/run_warmup_task23.sh
Run the following command to start training. Remember to change the --policy_model_path to the path of the checkpoint from the warmup training stage.
# Task 1
bash scripts/run_rl_task1.sh
# Task 2/3
bash scripts/run_rl_task23.sh
To evaluate the model, change the --policy_model_path to the path of the checkpoint from the RL training stage.
You can also download the checkpoint from here and put it in the outputs/examples folder. Then run the following command:
# Task 1
bash scripts/test_rl_task1.sh
# Task 2
bash scripts/test_rl_task2.sh
# Task 3
bash scripts/test_rl_task3.sh
You should be able to get the following output:
[Metric] Saved EB format predictions as PATH_TO_PREDICTION/pred_para.tsv
Use the generated file and follow the instructions in this repo to evaluate the results.
If you have any questions, please feel free to open an issue or contact me at txliu21@m.fudan.edu.cn.
Our code is based on ParaPattern, rlhr. Great thanks to the authors for their work!