Neural Modules with Adaptive Nonlinear Constraints and Efficient Regularizations (NeuroMANCER) is an open-source differentiable programming (DP) library for solving parametric constrained optimization problems, physics-informed system identification, and parametric model-based optimal control. NeuroMANCER is written in PyTorch and allows for systematic integration of machine learning with scientific computing for creating end-to-end differentiable models and algorithms embedded with prior knowledge and physics.
We've made some backwards-compatible changes in order to simplify integration and support multiple symbolic inputs to nn.Modules in our blocks interface.
New Colab Examples:
⭐ System identification for ordinary differential equations (ODEs)
See v1.4.1 release notes for more details.
Extensive set of tutorials can be found in the examples folder. Interactive notebook versions of examples are available on Google Colab! Test out NeuroMANCER functionality before cloning the repository and setting up an environment.
-
Part 2: NeuroMANCER syntax tutorial: variables, constraints, and objectives.
-
Part 3: NeuroMANCER syntax tutorial: modules, Node, and System class.
-
Part 1: Learning to solve a constrained optimization problem.
-
Part 2: Learning to solve a quadratically-constrained optimization problem.
-
Part 3: Learning to solve a set of 2D constrained optimization problems.
-
Part 4: Learning to solve a constrained optimization problem with projected gradient method.
-
Part 1: Neural Ordinary Differential Equations (NODEs)
-
Part 2: Parameter estimation of ODE system
-
Part 3: Universal Differential Equations (UDEs)
-
Part 4: NODEs with exogenous inputs
-
Part 5: Neural State Space Models (NSSMs) with exogenous inputs
-
Part 6: Data-driven modeling of resistance-capacitance (RC) network ODEs
Part 1: Diffusion Equation
Part 2: Burgers' Equation
Part 3: Burgers' Equation w/ Parameter Estimation (Inverse Problem)
-
Part 1: Learning to stabilize a linear dynamical system.
-
Part 2: Learning to stabilize a nonlinear differential equation.
-
Part 3: Learning to control a nonlinear differential equation.
-
Part 4: Learning neural ODE model and neural control policy for an unknown dynamical system.
-
Part 5: Learning neural Lyapunov function for a nonlinear dynamical system.
The documentation for the library can be found online. There is also an introduction video covering core features of the library.
# Neuromancer syntax example for constrained optimization
import neuromancer as nm
import torch
# define neural architecture
func = nm.modules.blocks.MLP(insize=1, outsize=2,
linear_map=nm.slim.maps['linear'],
nonlin=torch.nn.ReLU, hsizes=[80] * 4)
# wrap neural net into symbolic representation via the Node class: map(p) -> x
map = nm.system.Node(func, ['p'], ['x'], name='map')
# define decision variables
x = nm.constraint.variable("x")[:, [0]]
y = nm.constraint.variable("x")[:, [1]]
# problem parameters sampled in the dataset
p = nm.constraint.variable('p')
# define objective function
f = (1-x)**2 + (y-x**2)**2
obj = f.minimize(weight=1.0)
# define constraints
con_1 = 100.*(x >= y)
con_2 = 100.*(x**2+y**2 <= p**2)
# create penalty method-based loss function
loss = nm.loss.PenaltyLoss(objectives=[obj], constraints=[con_1, con_2])
# construct differentiable constrained optimization problem
problem = nm.problem.Problem(nodes=[map], loss=loss)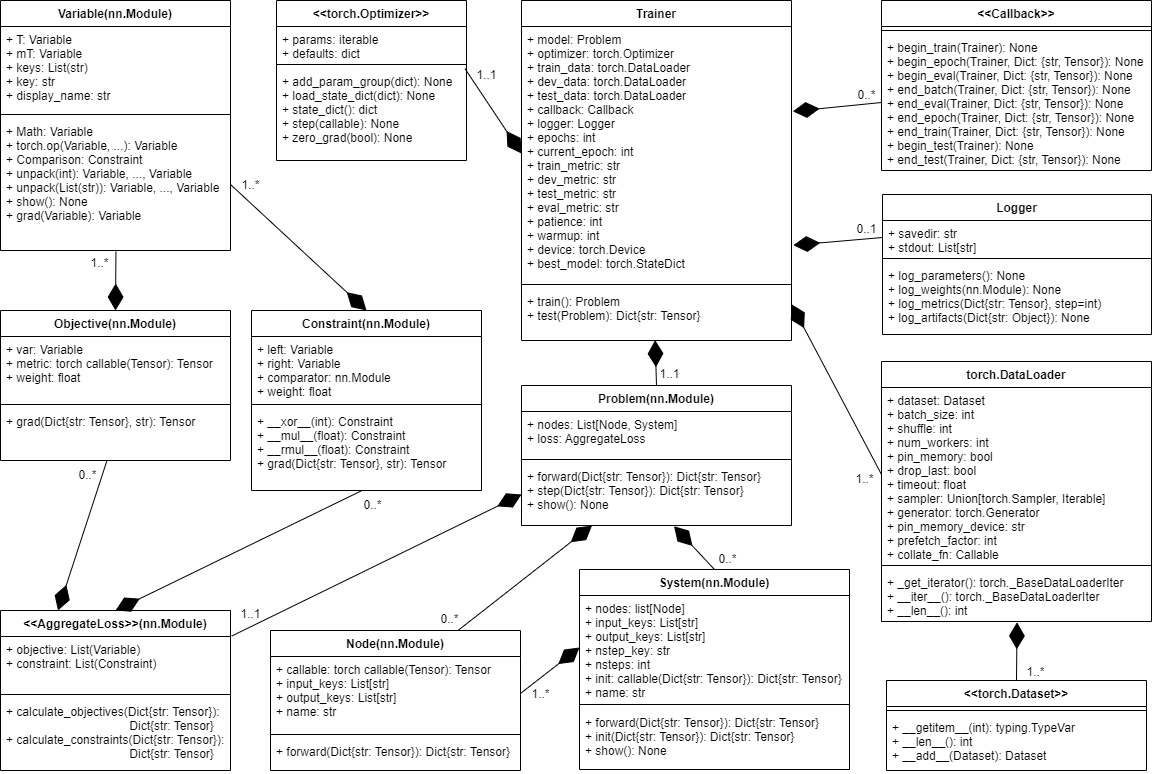 UML diagram of NeuroMANCER classes.
UML diagram of NeuroMANCER classes.
For either pip or conda installation, first clone the neuromancer package. A dedicated virtual environment (conda or otherwise) is recommended.
Note: If you have a previous neuromancer env it would be best at this point to create a new environment given the following instructions.
git clone -b master https://github.com/pnnl/neuromancer.git --single-branchRecommended installation.
Pip installation is broken up into required dependencies for core Neuromancer
and dependencies associated with the examples, tests, and generating the documentation.
Below we give instructions to install all dependencies in a conda virtual enviroment via pip.
You need at least pip version >= 21.3.
conda create -n neuromancer python=3.10.4
conda activate neuromancerFrom top level directory of cloned neuromancer run:
pip install -e.[docs,tests,examples]OR, for zsh users:
pip install -e.'[docs,tests,examples]'See the pyproject.toml file for reference.
[project.optional-dependencies]
tests = ["pytest", "hypothesis"]
examples = ["casadi", "cvxpy", "imageio"]
docs = ["sphinx", "sphinx-rtd-theme"]Before CVXPY can be installed on Apple M1, you must install cmake via Homebrew:
brew install cmakeSee CVXPY installation instructions for more details.
Conda install is recommended for GPU acceleration. In many cases the following simple install should work for the specified OS
conda env create -f linux_env.yml
conda activate neuromancerconda env create -f windows_env.yml
conda activate neuromancer
conda install -c defaults intel-openmp -fconda env create -f osxarm64_env.yml
conda activate neuromancer!!! Pay attention to comments for non-Linux OS !!!
conda create -n neuromancer python=3.10.4
conda activate neuromancer
conda install pytorch pytorch-cuda=11.6 -c pytorch -c nvidia
## OR (for Mac): conda install pytorch -c pytorch
conda config --append channels conda-forge
conda install scipy numpy"<1.24.0" matplotlib scikit-learn pandas dill mlflow pydot=1.4.2 pyts numba
conda install networkx=3.0 plum-dispatch=1.7.3
conda install -c anaconda pytest hypothesis
conda install cvxpy cvxopt casadi seaborn imageio
conda install tqdm torchdiffeq toml
## (for Windows): conda install -c defaults intel-openmp -fFrom the top level directory of cloned neuromancer (in the activated environment where the dependencies have been installed):
pip install -e . --no-depsRun pytest on the tests folder. It should take about 2 minutes to run the tests on CPU. There will be a lot of warnings that you can safely ignore. These warnings will be cleaned up in a future release.
We welcome contributions and feedback from the open-source community!
Discussions should be the first line of contact for new users to provide direct feedback on the library. Post your Ideas for new features or examples, showcase your work using neuromancer in Show and tell, or get support for usage in Q&As, please post them in one of our categories.
If you have an example of using NeuroMANCER to solve an interesting problem, or of using NeuroMANCER in a unique way please share them in Show and tell discussions. The best examples might be incorporated into our current library of examples. To submit an example, create a folder for your example/s in the example folder if there isn't currently an applicable folder and place either your executable python file or notebook file there. Push your code back to Github and then submit a pull request. Please make sure to note in a comment at the top of your code if there are additional dependencies to run your example and how to install those dependencies.
We welcome contributions to NeuroMANCER. Please accompany contributions with some lightweight unit tests via pytest (see test/ folder for some examples of easy to compose unit tests using pytest). In addition to unit tests a script utilizing introduced new classes or modules should be placed in the examples folder. To contribute a new well-developed feature please submit a pull request (PR). Before creating a PR, we encourage developers to discuss and document the intended feature in Ideas discussion category.
If you find a bug in the code or want to request a new well-developed feature, please open an issue.
Here are some upcoming features we plan to develop. Please let us know if you would like to get involved and contribute so we may be able to coordinate on development. If there is a feature that you think would be highly valuable but not included below, please open an issue and let us know your thoughts.
- Control and modelling for networked systems
- Support for stochastic differential equations (SDEs)
- Easy to implement modeling and control with uncertainty quantification
- Proximal operators for dealing with equality and inequality constraints
- Interface with CVXPYlayers
- Online learning examples
- Benchmark examples of DPC compared to deep RL
- Conda and pip package distribution
- More versatile and simplified time series dataloading
- Discovery of governing equations from learned RHS via NODEs and SINDy
- More domain science examples
- To simplify integration, interpolation of control input is no longer supported in
integrators.py- The
interp_uparameter ofIntegratorand subclasses has been deprecated
- The
- Additional inputs (e.g.,
u,t) can now be passed as*args(instead of as a single tensor input stacked withx) in:Integratorand subclasses inintegrators.pyBlock- new base class for all other classes inblocks.pyODESysteminode.py
- New Physics-Informed Neural Network (PINN) examples for solving PDEs in
/examples/PDEs/ - New system identification examples for ODEs in
/examples/ODEs/ - Fixed a bug in the
show(...)method of theProblemclass - Hotfix:
*argsforGeneralNetworkedODE
- Refactored PSL
- Better PSL unit testing coverage
- Consistent interfaces across system types
- Consistent perturbation signal interface in signals.py
- Refactored Control and System ID learning using Node and System class (system.py)
- Classes used for system ID can now be easily interchanged to accommodate downstream control policy learning
- Merged Structured Linear Maps and Pyton Systems Library into Neuromancer
- The code in neuromancer was closely tied to psl and slim. A decision was made to integrate the packages as submodules of neuromancer. This also solves the issue of the package names "psl" and "slim" already being taken on PyPI.
Import changes for psl and slim
# before
import psl
import slim
# now
from neuromancer import psl
from neuromancer import slim- New example scripts and notebooks
- Interactive Colab notebooks for testing Neuromancer functionality without setting up an environment
- See Examples for links to Colab
- RC-Network modeling using Graph Neural Time-steppers example:
- See neuromancer/examples/graph_timesteppers/
- Baseline NODE dynamics modeling results for all nonautonomous systems in Python Systems Library
- See neuromancer/examples/benchmarks/node/
- Updated install instructions for Linux, Windows, and MAC operating systems
- New linux_env.yml, windows_env.yml, osxarm64_env.yml files for installation of dependencies across OS
- Interactive Colab notebooks for testing Neuromancer functionality without setting up an environment
- Corresponding releases of SLiM and PSL packages
- Make sure to update these packages if updating Neuromancer
- Release 1.4 will roll SLiM and PSL into core Neuromancer for ease of installation and development
- Tutorial YouTube videos to accompany tutorial scripts in examples folder:
- Closed loop control policy learning examples with Neural Ordinary Differential Equations
- examples/control/
- vdpo_DPC_cl_fixed_ref.py
- two_tank_sysID_DPC_cl_var_ref.py
- two_tank_DPC_cl_var_ref.py
- two_tank_DPC_cl_fixed_ref.py
- examples/control/
- Closed loop control policy learning example with Linear State Space Models.
- examples/control/
- double_integrator_dpc_ol_fixed_ref.py
- vtol_dpc_ol_fixed_ref.py
- examples/control/
- New class for Linear State Space Models (LSSM)
- LinearSSM in dynamics.py
- Refactored closed-loop control policy simulations
- simulator.py
- Interfaces for open and closed loop simulation (evaluation after training) for several classes
- Dynamics
- Estimator
- Policy
- Constraint
- PSL Emulator classes
- New class for closed-loop policy learning of non-autonomous ODE systems
- ControlODE class in ode.py
- Added support for NODE systems
- Torchdiffeq integration with fast adjoint method for NODE optimization
Lead developers: Aaron Tuor, Jan Drgona
Active PNNL developers: James Koch, Madelyn Shapiro, Draguna Vrabie
Community contributors: Seth Briney, Bo Tang
Past contributors: Shrirang Abhyankar, Mia Skomski, Stefan Dernbach, Zhao Chen, Christian Møldrup Legaard
- James Koch, Zhao Chen, Aaron Tuor, Jan Drgona, Draguna Vrabie, Structural Inference of Networked Dynamical Systems with Universal Differential Equations, arXiv:2207.04962, (2022)
- Ján Drgoňa, Sayak Mukherjee, Aaron Tuor, Mahantesh Halappanavar, Draguna Vrabie, Learning Stochastic Parametric Differentiable Predictive Control Policies, IFAC ROCOND conference (2022)
- Sayak Mukherjee, Ján Drgoňa, Aaron Tuor, Mahantesh Halappanavar, Draguna Vrabie, Neural Lyapunov Differentiable Predictive Control, IEEE Conference on Decision and Control Conference 2022
- Wenceslao Shaw Cortez, Jan Drgona, Aaron Tuor, Mahantesh Halappanavar, Draguna Vrabie, Differentiable Predictive Control with Safety Guarantees: A Control Barrier Function Approach, IEEE Conference on Decision and Control Conference 2022
- Ethan King, Jan Drgona, Aaron Tuor, Shrirang Abhyankar, Craig Bakker, Arnab Bhattacharya, Draguna Vrabie, Koopman-based Differentiable Predictive Control for the Dynamics-Aware Economic Dispatch Problem, 2022 American Control Conference (ACC)
- Drgoňa, J., Tuor, A. R., Chandan, V., & Vrabie, D. L., Physics-constrained deep learning of multi-zone building thermal dynamics. Energy and Buildings, 243, 110992, (2021)
- E. Skomski, S. Vasisht, C. Wight, A. Tuor, J. Drgoňa and D. Vrabie, "Constrained Block Nonlinear Neural Dynamical Models," 2021 American Control Conference (ACC), 2021, pp. 3993-4000, doi: 10.23919/ACC50511.2021.9482930.
- Skomski, E., Drgoňa, J., & Tuor, A. (2021, May). Automating Discovery of Physics-Informed Neural State Space Models via Learning and Evolution. In Learning for Dynamics and Control (pp. 980-991). PMLR.
- Drgoňa, J., Tuor, A., Skomski, E., Vasisht, S., & Vrabie, D. (2021). Deep Learning Explicit Differentiable Predictive Control Laws for Buildings. IFAC-PapersOnLine, 54(6), 14-19.
- Tuor, A., Drgona, J., & Vrabie, D. (2020). Constrained neural ordinary differential equations with stability guarantees. arXiv preprint arXiv:2004.10883.
- Drgona, Jan, et al. "Differentiable Predictive Control: An MPC Alternative for Unknown Nonlinear Systems using Constrained Deep Learning." Journal of Process Control Volume 116, August 2022, Pages 80-92
- Drgona, J., Skomski, E., Vasisht, S., Tuor, A., & Vrabie, D. (2020). Dissipative Deep Neural Dynamical Systems, in IEEE Open Journal of Control Systems, vol. 1, pp. 100-112, 2022
- Drgona, J., Tuor, A., & Vrabie, D., Learning Constrained Adaptive Differentiable Predictive Control Policies With Guarantees, arXiv preprint arXiv:2004.11184, (2020)
@article{Neuromancer2023,
title={{NeuroMANCER: Neural Modules with Adaptive Nonlinear Constraints and Efficient Regularizations}},
author={Tuor, Aaron and Drgona, Jan and Koch, James and Shapiro, Madelyn and Vrabie, Draguna and Briney, Seth},
Url= {https://github.com/pnnl/neuromancer},
year={2023}
}This research was partially supported by the Mathematics for Artificial Reasoning in Science (MARS) and Data Model Convergence (DMC) initiatives via the Laboratory Directed Research and Development (LDRD) investments at Pacific Northwest National Laboratory (PNNL), by the U.S. Department of Energy, through the Office of Advanced Scientific Computing Research's “Data-Driven Decision Control for Complex Systems (DnC2S)” project, and through the Energy Efficiency and Renewable Energy, Building Technologies Office under the “Dynamic decarbonization through autonomous physics-centric deep learning and optimization of building operations” and the “Advancing Market-Ready Building Energy Management by Cost-Effective Differentiable Predictive Control” projects. PNNL is a multi-program national laboratory operated for the U.S. Department of Energy (DOE) by Battelle Memorial Institute under Contract No. DE-AC05-76RL0-1830.