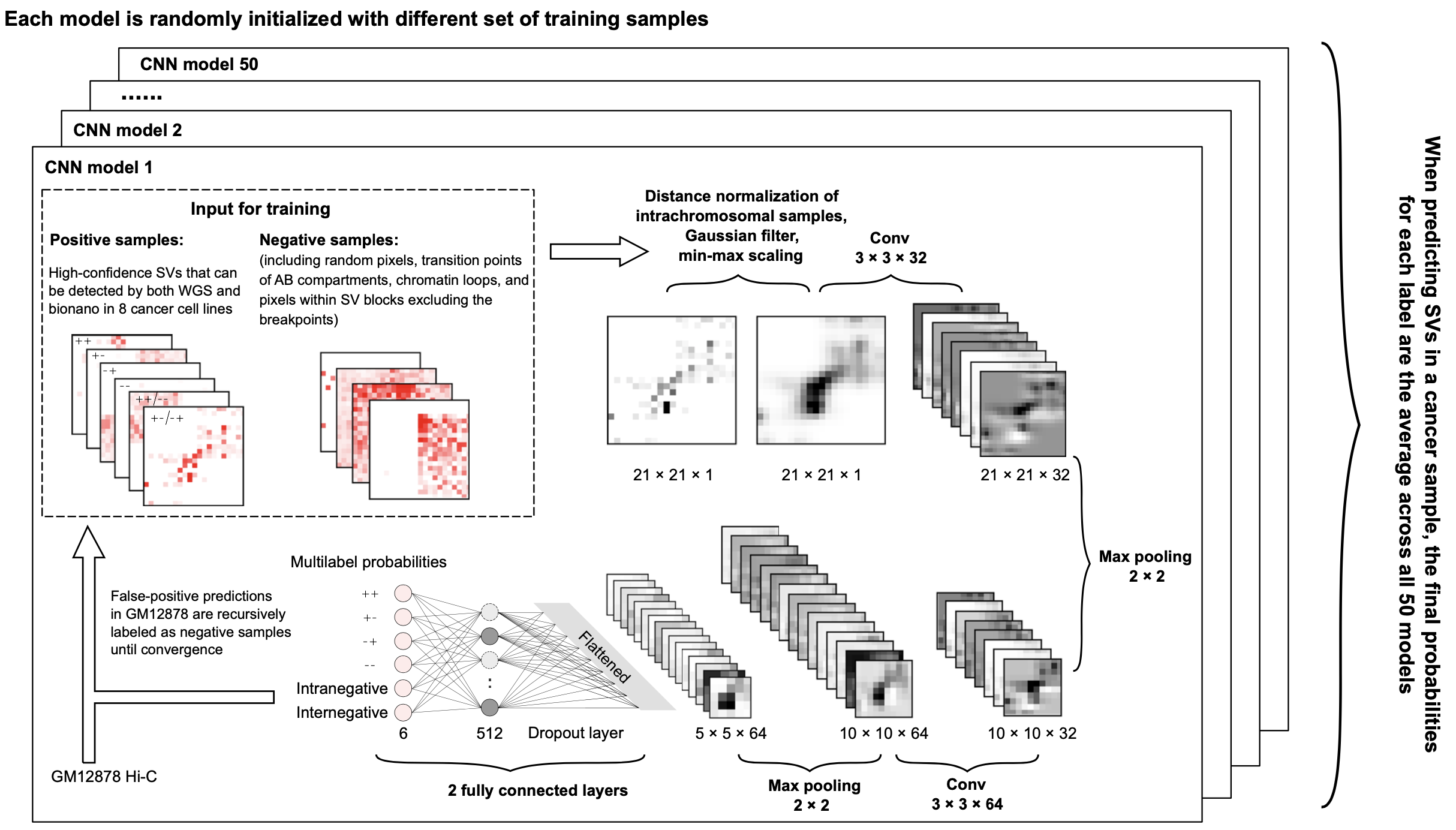
Hi-C technique has been shown to be a promising method to detect structural variations (SVs) in human genomes. However, algorithms that can use Hi-C data for a full-range SV detection have been severely lacking. Current methods can only identify inter-chromosomal translocations and long-range intra-chromosomal SVs (>1Mb) at less-than-optimal resolution. Therefore, we develop EagleC, a framework that combines deep-learning and ensemble-learning strategies to predict a full-range of SVs at high-resolution. Importantly, we show that EagleC can uniquely capture a set of fusion genes that are missed by WGS or nanopore. Furthermore, EagleC also effectively captures SVs in other chromatin interaction platforms, such as HiChIP, ChIA-PET, and capture Hi-C. We apply EagleC in over 100 cancer cell lines and primary tumors, and identify a valuable set of high-quality SVs. Finally, we demonstrate that EagleC can be applied to single-cell Hi-C and used to study the SV heterogeneity in primary tumors.
- EagleC is able to accurately detect a full range of SVs including short-range SVs with breakpoint distance less than 100kb or even 50kb
- EagleC is designed to accept any 3C-based contact maps, including Hi-C, Micro-C, HiChIP, ChIA-PET, capture Hi-C, and single-cell Hi-C
- EagleC can be used to predict SVs in any species (it has been tested in human, mouse, and zebrafish)
Wang, X., Luan, Y., Yue, F. EagleC: A deep-learning framework for detecting a full range of structural variations from bulk and single-cell contact maps. Sci Adv. 2022.
- Installation
- Download pre-trained models
- Overview of the commands
- Quick Start
- Annotate gene fusions
- Visualize predicted SVs on contact maps
- Locate high-resolution coordinates given a list of low-resolution SVs
- Predict SVs at higher resolutions
- Predict SVs in other species
First, install following python packages using mamba:
$ conda config --add channels defaults $ conda config --add channels bioconda $ conda config --add channels conda-forge $ mamba create -n EagleC scikit-learn statsmodels matplotlib cooler pyBigWig pyensembl python=3.8 joblib=1.0.1 cython=0.29.24 "tensorflow<=2.11"
Note
matplotlib and pyBigWig are only required if you want to use the visualization module to view the predicted SVs on contact maps, and pyensembl is only required if you want to annotate potential gene fusions given a list of SV breakpoints.
If you are installing EagleC in Linux, just execute the command below to install EagleC from PyPI:
$ pip install eaglec
If you are installing EagleC in MacOS, please download and install an appropriate package from here:
$ pip install eaglec-0.1.9-cp38-cp38-macosx_10_9_x86_64.whl
We have trained a series of EagleC models covering various sequencing depths for both bulk Hi-C maps and single-cell Hi-C maps. Before running EagleC, we recommend downloading these pre-trained models by simply executing the command below. In prediction, EagleC will automatically select the most appropriate models according to the number of contacts in your contact map:
$ download-pretrained-models
EagleC is distributed with 6 command-line tools. Type command [-h] in a terminal
window to learn the basic usage of each command.
predictSV
predictSV is the main command we used to predict SVs from bulk Hi-C/HiChIP/ChIA-PET contact maps in this work. It is based on predictSV-single-resolution and automatically combines predictions from 5kb, 10kb, and 50kb resolutions. For 10kb and 50kb predictions, it further searches for the most probable breakpoint coordinates within a local region on 5kb contact maps so that all the reported SVs are at the 5kb resolution.
The inputs to this command are three genome-wide contact maps at 5kb, 10kb, and 50kb resolutions in .cool format (cool URIs, refer to cooler if you are not familiar with this format). If you only have .hic files, consider converting your files to the ".cool" format using hic2cool or pairLiftOver. The predicted SVs can be selected to be reported in two formats: 1) "--output-format full" will report 8 columns for each SV, including breakpoint coordinates (chrom1, pos1, chrom2, pos2) and probability values of each fusion type (++, +-, -+, and --) (refer to Figures S1-S2 for the definition of each fusion type); 2) "--output-format NeoLoopFinder" will output a file (6 columns) that can be directly used as the NeoLoopFinder input.
predictSV-single-resolution
This command predicts SVs at single resolution. By default, it searches for SVs throughout the whole genome; however, it can also perform a local search on high-resolution matrices if SVs at lower resolutions are provided through the parameter "--low-resolution-breaks".
merge-redundant-SVs
This command merges multiple SV calls from the same sample. The inputs are one or multiple SV files from predictSV or predictSV-single-resolution in "full" format (8 columns). Again, the output format has two options ("full" and "NeoLoopFinder").
annotate-gene-fusion
This command can be used to annotate gene fusion events for a list of SV breakpoints. The input to this command is an SV file with breakpoint coordinate information (chrom1, pos1, chrom2, pos2) in the first four columns and a release number of ensembl genes.
plot-interSVs
This command can be used to plot a chromosome-wide contact map with predicted SVs marked on it.
plot-intraSVs
This command can be used to plot a local intra-chromosomal contact map with predicted SVs marked on it.
First, let's download a processed Hi-C dataset (~163M contact pairs) in SK-N-AS (a neuroblastoma cell line):
$ wget -O SKNAS-MboI-allReps-filtered.mcool -L https://www.dropbox.com/s/f80bgn11d7wfgq8/SKNAS-MboI-allReps-filtered.mcool?dl=0
The downloaded ".mcool" file contains contact matrices at multiple resolutions. To list all
individual cool URIs within it, execute the cooler ls command below:
$ cooler ls SKNAS-MboI-allReps-filtered.mcool SKNAS-MboI-allReps-filtered.mcool::/resolutions/5000 SKNAS-MboI-allReps-filtered.mcool::/resolutions/10000 SKNAS-MboI-allReps-filtered.mcool::/resolutions/25000 SKNAS-MboI-allReps-filtered.mcool::/resolutions/50000 SKNAS-MboI-allReps-filtered.mcool::/resolutions/100000 SKNAS-MboI-allReps-filtered.mcool::/resolutions/250000 SKNAS-MboI-allReps-filtered.mcool::/resolutions/500000 SKNAS-MboI-allReps-filtered.mcool::/resolutions/1000000 SKNAS-MboI-allReps-filtered.mcool::/resolutions/2500000 SKNAS-MboI-allReps-filtered.mcool::/resolutions/5000000
Next, let's use the predictSV command to predict SVs on this dataset:
$ predictSV --hic-5k SKNAS-MboI-allReps-filtered.mcool::/resolutions/5000 \
--hic-10k SKNAS-MboI-allReps-filtered.mcool::/resolutions/10000 \
--hic-50k SKNAS-MboI-allReps-filtered.mcool::/resolutions/50000 \
-O SK-N-AS -g hg38 --balance-type CNV --output-format full \
--prob-cutoff-5k 0.8 --prob-cutoff-10k 0.8 --prob-cutoff-50k 0.99999
As we mentioned in Overview of the commands, contact matrices at three resolutions 5kb, 10kb, and 50kb will be used. Here are some suggestions for individual parameters:
--balance-type, here by specifying "--balance-type CNV", predictSV will perform predictions on CNV-normalized matrices. You can also select to use ICE-normalized matrices by specifying "--balance-type ICE" or Raw matrices by specifying "--balance-type Raw". According to our test, for the same sample, running on the Raw matrix tends to detect more SVs with lower accuracy, while running on the CNV/ICE normalized matrices usually achieves higher accuracy but detects fewer SVs.
Note
If you choose CNV, make sure you have run "correct-cnv" of the NeoLoopFinder toolkit before you run this command; if you choose ICE, make sure you have run "cooler balance" on your Hi-C matrices before you run this command.
- By default, we apply probability cutoffs of 0.8, 0.8, and 0.99999 at 5kb, 10kb, and 50kb resolutions, respectively. We found this set of cutoffs achieved a good tradeoff between sensitivity and specificity in most of our tests. If you care more about sensitivity, just tune down these cutoffs.
Running predictSV on a single CPU core is expected to be slow, as it iterates submatrices of all candidate pixels on these contact matrices. To speed up the calculation, predictSV supports parallel computation for different intra-chromosomal and inter-chromosomal matrices, by creating hidden lock files to avoid conflicts between jobs. This strategy is especially efficient when you are performing the calculation in a computational cluster. Depending on your cluster environment, you need to create a job submission script. Here is an example slurm script named as "slurm-predictSV.sh":
#!/bin/bash
#SBATCH -A b1042
#SBATCH -p genomicsguestA
#SBATCH -t 48:00:00
#SBATCH -N 1
#SBATCH --mem=16G
#SBATCH --cpus-per-task=1
#SBATCH --job-name=predictSV
#SBATCH --output=predictSV.%j.%N.txt
#SBATCH --error=predictSV.%j.%N.err
source /home/xwl2576/.bashrc
mamba activate EagleC
predictSV --hic-5k SKNAS-MboI-allReps-filtered.mcool::/resolutions/5000 \
--hic-10k SKNAS-MboI-allReps-filtered.mcool::/resolutions/10000 \
--hic-50k SKNAS-MboI-allReps-filtered.mcool::/resolutions/50000 \
-O SK-N-AS -g hg38 --balance-type CNV --output-format full \
--prob-cutoff-5k 0.8 --prob-cutoff-10k 0.8 --prob-cutoff-50k 0.99999
Then all you need to do is to submit this script for a certain number of times:
$ for i in {1..16}; do sbatch slurm-predictSV.sh; sleep 40s; done
The above command will launch 16 parallelized jobs and should be able to finish within 2 hours.
Note
EagleC will cache all the intermediate results within hidden folders in
your current working directory. In this example, these folders will be prefixed with
".SKNAS-MboI-allReps-filtered.mcool". If you want to start a fresh job without using
previous cached results, or if your previous jobs were killed or terminated
by the system, you may need to first remove those intermediate files by executing
rm -rf .SKNAS-MboI-allReps-filtered.mcool*.
If no errors occurred, 6 files ("SK-N-AS.CNN_SVs.10K_highres.txt", "SK-N-AS.CNN_SVs.10K.txt", "SK-N-AS.CNN_SVs.50K_highres.txt", "SK-N-AS.CNN_SVs.50K.txt", "SK-N-AS.CNN_SVs.5K_combined.txt", and "SK-N-AS.CNN_SVs.5K.txt") will be outputed in current working directory. Among them, the file "SK-N-AS.CNN_SVs.5K_combined.txt" contains the final non-redundant SVs combined from 5kb, 10kb, and 50kb resolutions:
$ head SK-N-AS.CNN_SVs.5K_combined.txt chrom1 pos1 chrom2 pos2 ++ +- -+ -- chr10 100540000 chr10 101175000 1.885e-15 4.558e-22 1 1.827e-16 chr11 100080000 chr11 100160000 1.319e-26 1 1.47e-23 1.292e-15 chr11 40120000 chr11 40300000 2.869e-13 7.797e-17 0.964 1.603e-17 chr11 71720000 chr17 32285000 3.397e-23 1 8.086e-15 1.674e-18 chr12 111605000 chr16 83395000 6.232e-29 1.972e-28 1 8.747e-27 chr13 63030000 chr17 22155000 1.812e-10 1.975e-16 0.9197 2.687e-12 chr16 21580000 chr16 22695000 1 4.339e-28 6.561e-27 1.242e-17 chr17 73790000 chr19 780000 1.392e-21 2.4e-29 2.071e-24 1 chr18 47755000 chr18 48025000 1.861e-13 3.204e-14 0.9863 1.928e-16
To annotate potential gene fusion events from the predicted SVs above, just execute the command below:
$ annotate-gene-fusion --sv-file SK-N-AS.CNN_SVs.5K_combined.txt \
--output-file SK-N-AS.gene-fusions.txt \
--buff-size 10000 --skip-rows 1 --ensembl-release 93 --species human
Here by specifying "--ensembl-release 93 --species human", annotate-gene-fusion uses the Ensembl gene release 93 of the human genome as a database to search for genes at any genomic loci. The "--buff-size" parameter determines the genomic span (in base pair) of the breakpoints for each SV. Here, a gene will be considered at a breakpoint if its interval is overlapped with the +/-10kb region centered at the breakpoint:
$ head -5 SK-N-AS.gene-fusions.txt chr12 111605000 chr16 83395000 6.232e-29 1.972e-28 1 8.747e-27 ATXN2-CDH13 chr1 1930000 chr1 10975000 2.572e-25 1 1.017e-17 1.627e-20 CFAP74-C1orf127 chr1 25255000 chr1 25330000 8.584e-19 0.8123 1.172e-19 4.559e-14 RSRP1-TMEM50A,RSRP1-RHD chr1 1765000 chr1 1905000 2.688e-11 1.744e-18 0.8671 6.763e-09 NADK-CALML6 chr3 60625000 chr17 42830000 6.303e-24 8.763e-27 7.642e-27 1 FHIT-BECN1,FHIT-PSME3
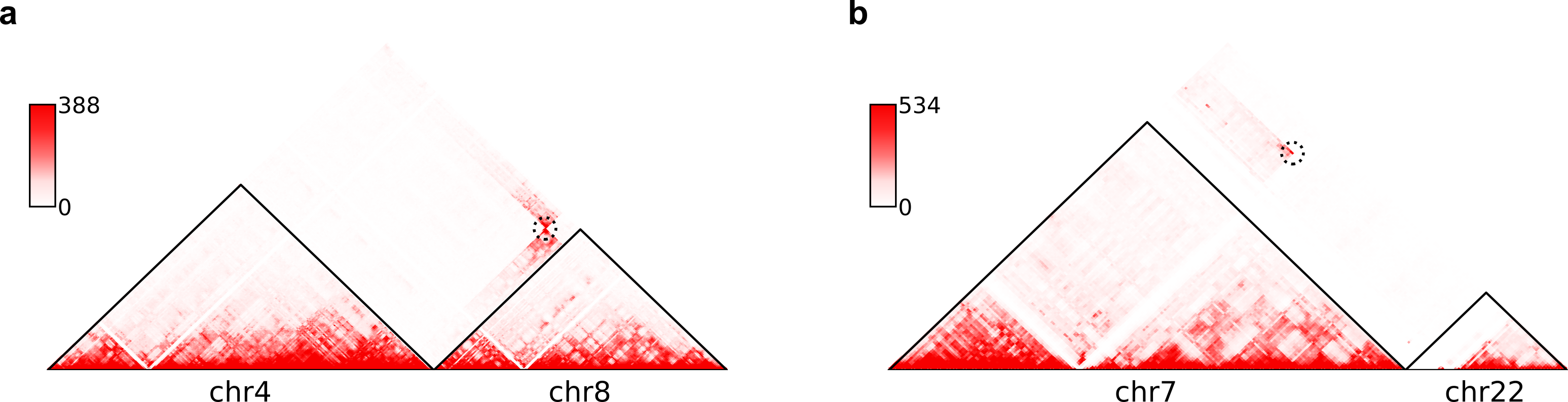
Here are example commands to visualize SVs on chromosome-wide contact maps:
$ plot-interSVs --cool-uri SKNAS-MboI-allReps-filtered.mcool::resolutions/1000000 \
--full-sv-file SK-N-AS.CNN_SVs.5K_combined.txt --output-figure-name chr4-chr8.png \
-C chr4 chr8 --balance-type Raw --dpi 800 # panel A
$ plot-interSVs --cool-uri SKNAS-MboI-allReps-filtered.mcool::resolutions/1000000 \
--full-sv-file SK-N-AS.CNN_SVs.5K_combined.txt --output-figure-name chr7-chr22.png \
-C chr7 chr22 --balance-type Raw --dpi 800 # panel B
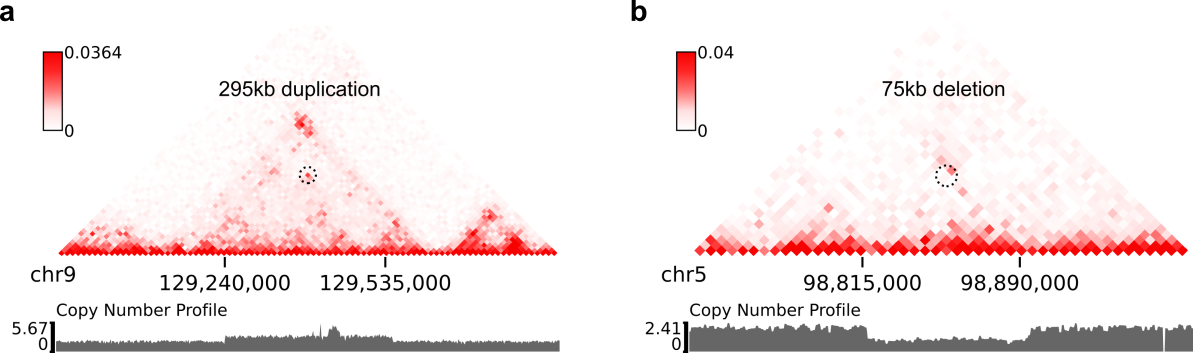
Here are example commands to visualize SVs on local intra-chromosomal contact maps:
$ wget -O SKNAS_merged.dedup.bam_ratio.bw -L https://www.dropbox.com/s/usjjc6neqs5fk3a/SKNAS_merged.dedup.bam_ratio.bw?dl=0
$ plot-intraSVs --cool-uri SKNAS-MboI-allReps-filtered.mcool::resolutions/10000 \
--full-sv-file SK-N-AS.CNN_SVs.5K_combined.txt \
--cnv-file SKNAS_merged.dedup.bam_ratio.bw \
--region chr9:128940000-129835000 --output-figure-name intraSV-example1.png \
--coordinates-to-display 129240000 129535000 \
--balance-type CNV --dpi 800 # panel A
$ plot-intraSVs --cool-uri SKNAS-MboI-allReps-filtered.mcool::resolutions/5000 \
--full-sv-file SK-N-AS.CNN_SVs.5K_combined.txt \
--cnv-file SKNAS_merged.dedup.bam_ratio.bw \
--region chr5:98735000-98970000 --output-figure-name intraSV-example2.png \
--coordinates-to-display 98815000 98890000 \
--contact-max-value 0.04 \
--balance-type CNV --dpi 800 # panel B
In above figures, the predicted SVs are marked by black dashed circles.
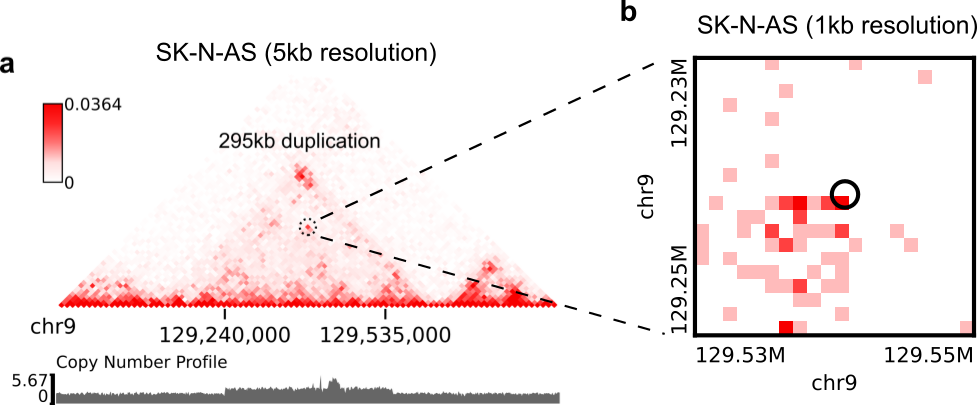
Suppose you have a high-resolution contact map at the 1kb or 2kb resolution, and a list of SVs detected from relatively low-resolution (50kb/10kb) contact maps of the same sample, you want to use the high-resolution map to find more precise breakpoint coordinates for these SVs, rather than perform a genome-wide SV prediction on the high-resolution map. With the predictSV-single-resolution command, you can easily get this job done by specifying the "--low-resolution-breaks" parameter.
For example, the coordinate of the duplication showed in the previous figure (panel A) were determined as ("chr9", 129,240,000, "chr9", 129,535,000) at the 5kb resolution. Now let's try to use the predictSV-single-resolution command to find more precise coordinates at the 1kb resolution.
First, let's extract the line containing this duplication from "SK-N-AS.CNN_SVs.5K_combined.txt" and put it into a new TXT file:
$ head -1 SK-N-AS.CNN_SVs.5K_combined.txt > test.txt $ grep '129240000\|129535000' SK-N-AS.CNN_SVs.5K_combined.txt >> test.txt $ cat test.txt chrom1 pos1 chrom2 pos2 ++ +- -+ -- chr9 129240000 chr9 129535000 1.35e-13 5.094e-14 0.8477 6.37e-18
Then download the SK-N-AS Hi-C map at the 1kb resolution:
$ wget -O SKNAS-MboI-allReps-filtered.1kb.cool -L https://www.dropbox.com/s/m8tqsr7ics9juas/SKNAS-MboI-allReps-filtered.1kb.cool?dl=0
And execute the command below:
$ predictSV-single-resolution -H SKNAS-MboI-allReps-filtered.1kb.cool -O test.1k.txt \
-g hg38 --balance-type Raw --low-resolution-breaks test.txt \
--region-size 10000
Here by specifying --region-size 10000, we limit the program to perform a local search within
+/-10kb of the input coordinates. Wait ~1 minutes, then you can find more precise breakpoint
coordinates in "test.1k.txt":
$ cat test.1k.txt chrom1 pos1 chrom2 pos2 ++ +- -+ -- chr9 129239000 chr9 129536000 1.35e-13 5.094e-14 0.8477 6.37e-18
Note that when you run predictSV-single-resolution with the parameter "--low-resolution-breaks", the program will keep the probability scores the same and only change the coordinates in the 2nd and 4th columns.
If you want to predict smaller SVs, try "predictSV-single-resolution" on high-resolution maps (1kb or 2kb) without specifying the "--low-resolution-breaks" parameter.
In this example, we will use a CTCF ChIA-PET dataset (containing ~266M usable reads) to predict SVs in MCF7 at the 2kb resolution:
$ wget -O ChIA-PET_hg38_MCF7_CTCF_pairs.2K.cool -L https://www.dropbox.com/s/bqz71zn9pg5si6a/ChIA-PET_hg38_MCF7_CTCF_pairs.2K.cool?dl=0
Again, let's create a job submission script "slurm-predictSV-2k.sh":
#!/bin/bash
#SBATCH -A b1042
#SBATCH -p genomicsguestA
#SBATCH -t 48:00:00
#SBATCH -N 1
#SBATCH --mem=20G
#SBATCH --cpus-per-task=1
#SBATCH --job-name=eaglec
#SBATCH --output=eaglec.%j.%N.txt
#SBATCH --error=eaglec.%j.%N.err
source /home/xwl2576/.bashrc
mamba activate EagleC
predictSV-single-resolution --hic ChIA-PET_hg38_MCF7_CTCF_pairs.2K.cool \
-O MCF7_CTCF-ICE.SVs.2k.txt -g hg38 \
-C 1 18 --maximum-size 100000 --balance-type ICE \
--add-log-header --logFile eaglec-ice-2k.log
And submit it for a number of times:
for i in {1..2}; do sbatch slurm-predictSV-2k.sh; sleep 40s; done
Note that identifying SVs on 1kb/2kb contact maps is really time consuming. Here by specifying
-C 1 18 --maximum-size 100000, we limit our search space to chromosomes 1 and 18, and only
consider SV candidates with breakpoint distance less than 100kb.
This job will finish within 10 minutes. Now let's plot the predicted SVs:
$ cat MCF7_CTCF-ICE.SVs.2k.txt
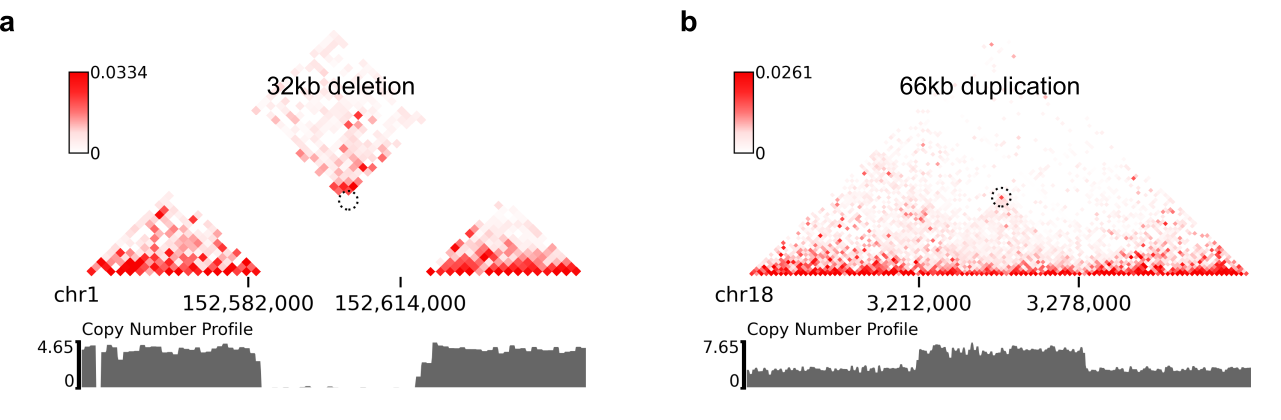
chr18 3212000 chr18 3278000 8.129e-18 7.563e-19 0.9997 2.274e-16
chr1 152584000 chr1 152616000 1.324e-09 0.9528 6.386e-10 6.64e-08
$ wget -O MCF7_merged.dedup.bam_ratio.bw -L https://www.dropbox.com/s/rstx3lzvpin8d0m/MCF7_merged.dedup.bam_ratio.bw?dl=0
$ plot-intraSVs --cool-uri ChIA-PET_hg38_MCF7_CTCF_pairs.2K.cool \
--full-sv-file MCF7_CTCF-ICE.SVs.2k.txt \
--cnv-file MCF7_merged.dedup.bam_ratio.bw \
--region chr1:152547000-152649000 --output-figure-name intraSV-example3.png \
--coordinates-to-display 152582000 152614000 \
--balance-type ICE --dpi 800 # panel A
$ plot-intraSVs --cool-uri ChIA-PET_hg38_MCF7_CTCF_pairs.2K.cool \
--full-sv-file MCF7_CTCF-ICE.SVs.2k.txt \
--cnv-file MCF7_merged.dedup.bam_ratio.bw \
--region chr18:3142000-3348000 --output-figure-name intraSV-example4.png \
--coordinates-to-display 3212000 3278000 \
--balance-type ICE --dpi 800 # panel B
To predict SVs in other species, just specify "--genome other" when you run predictSV or predictSV-single-resolution.