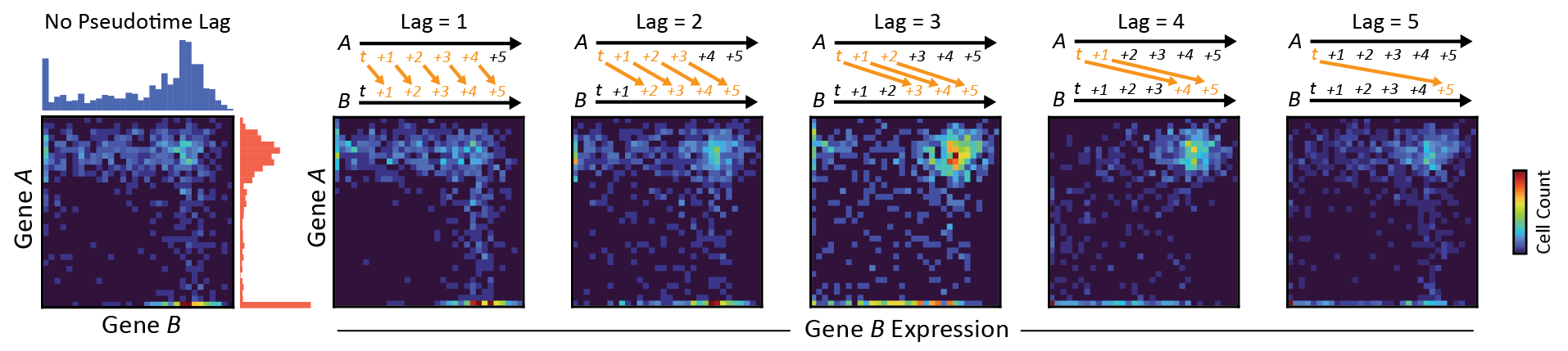
DELAY: DEpicting LAgged causalitY across single-cell trajectories for accurate gene-regulatory inference
-
Follow these instructions to install the latest version of PyTorch with CUDA support: https://pytorch.org
- Please note, DELAY currently requires CUDA-capable GPUs for training and prediction
-
Confirm that two additional dependencies have been satisfied:
pytorch-lightningandpandas -
Navigate to the location where you want to clone the repository and run:
git clone https://github.com/calebclayreagor/DELAY.git
1. Fine-tune DELAY on datasets with partially-known ground-truth interactions, e.g. from ChIP-seq experiments:
python RunDELAY.py [datadir] [outdir] -k [val_fold] [--atac] -p -ft
-kis the validation fold and--ataccan optionally specify scATAC-seq input data (default is scRNA-seq)- Use TensorBoard to monitor training by runnning
tensorboard --logdir RESULTSfrom the main directory - By default, DELAY will save the best model weights to a checkpoint file in
RESULTS/outdir
python RunDELAY.py [datadir] [outdir] -m [RESULTS/outdir/BEST_WEIGHTS.ckpt] -p -g 1 -bs 1024
- DELAY will save the predicted gene-regulation probabilities as a
tfs x genesmatrix inoutdirnamedregPredictions.csv - By default, DELAY will load batches from existing directories, so make sure to delete created folders for all
training,validationandpredictionbatches when finished
For additional help, run python RunDELAY.py --help
DELAY will expect unique sub-directories for each dataset in datadir containing the following files:
-
NormalizedData.csv— A labeledgenes x cellsmatrix of gene-expression or accessibility values -
PseudoTime.csv— A single-column table (cells x "PseudoTime") of inferred pseudotime values -
refNetwork.csv— A two-column table of ground-truth interactions between TFs ("Gene1") and target genes ("Gene2") -
TranscriptionFactors.csv(REQUIRED FOR INFERENCE) — A list of known transcription factors and co-factors in the dataset -
splitLabels.csv(REQUIRED FOR VALIDATION) — A single-column table (tfs x "Split") of training and validation folds for TFs in therefNetwork
For more help, see the example-data directory1
python RunDELAY.py [datadir] [outdir] --train -k [val_fold] \
--model_type vgg -cfg 32 32 M 64 64 M 128 128 M
Read the peer-reviewed paper: https://doi.org/10.1093/pnasnexus/pgad113
Footnotes
-
Example data taken from Hayashi et al., Nature Communications (2018) ↩