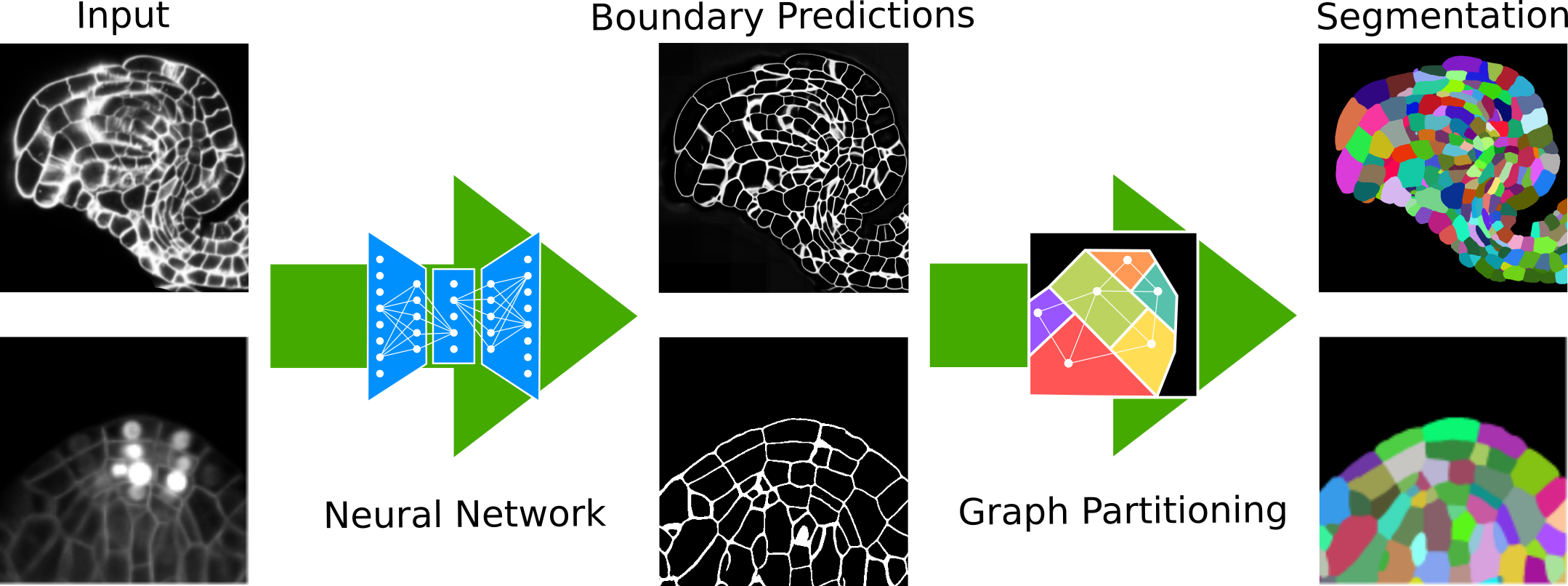
PlantSeg is a tool for cell instance aware segmentation in densely packed 3D volumetric images. The pipeline uses a two stages segmentation strategy (Neural Network + Segmentation). The pipeline is tuned for plant cell tissue acquired with confocal and light sheet microscopy. Pre-trained models are provided.
For detailed usage checkout our documentation 📖.
Please go to the documentation for more detailed instructions. In short, we recommend using mamba to install PlantSeg, which is currently supported on Linux and Windows.
-
GPU version, CUDA=12.x (replace
12.1with11.8for CUDA=11.x)mamba create -n plant-seg -c pytorch -c nvidia -c conda-forge pytorch pytorch-cuda=12.1 plant-seg --no-channel-priority
-
CPU version
mamba create -n plant-seg -c pytorch -c nvidia -c conda-forge pytorch cpuonly plant-seg --no-channel-priority
The above command will create new conda environment plant-seg together with all required dependencies.
The PlantSeg repository is organised as follows:
- plantseg: Contains the source code of PlantSeg.
- conda-reicpe: Contains all necessary code and configuration to create the anaconda package.
- docs: Contains the documentation of PlantSeg.
- evaluation: Contains all script required to reproduce the quantitative evaluation in Wolny et al..
- examples: Contains the files required to test PlantSeg.
- tests: Contains automated tests that ensures the PlantSeg functionality are not compromised during an update.
@article{wolny2020accurate,
title={Accurate and versatile 3D segmentation of plant tissues at cellular resolution},
author={Wolny, Adrian and Cerrone, Lorenzo and Vijayan, Athul and Tofanelli, Rachele and Barro, Amaya Vilches and Louveaux, Marion and Wenzl, Christian and Strauss, S{\"o}ren and Wilson-S{\'a}nchez, David and Lymbouridou, Rena and others},
journal={Elife},
volume={9},
pages={e57613},
year={2020},
publisher={eLife Sciences Publications Limited}
}