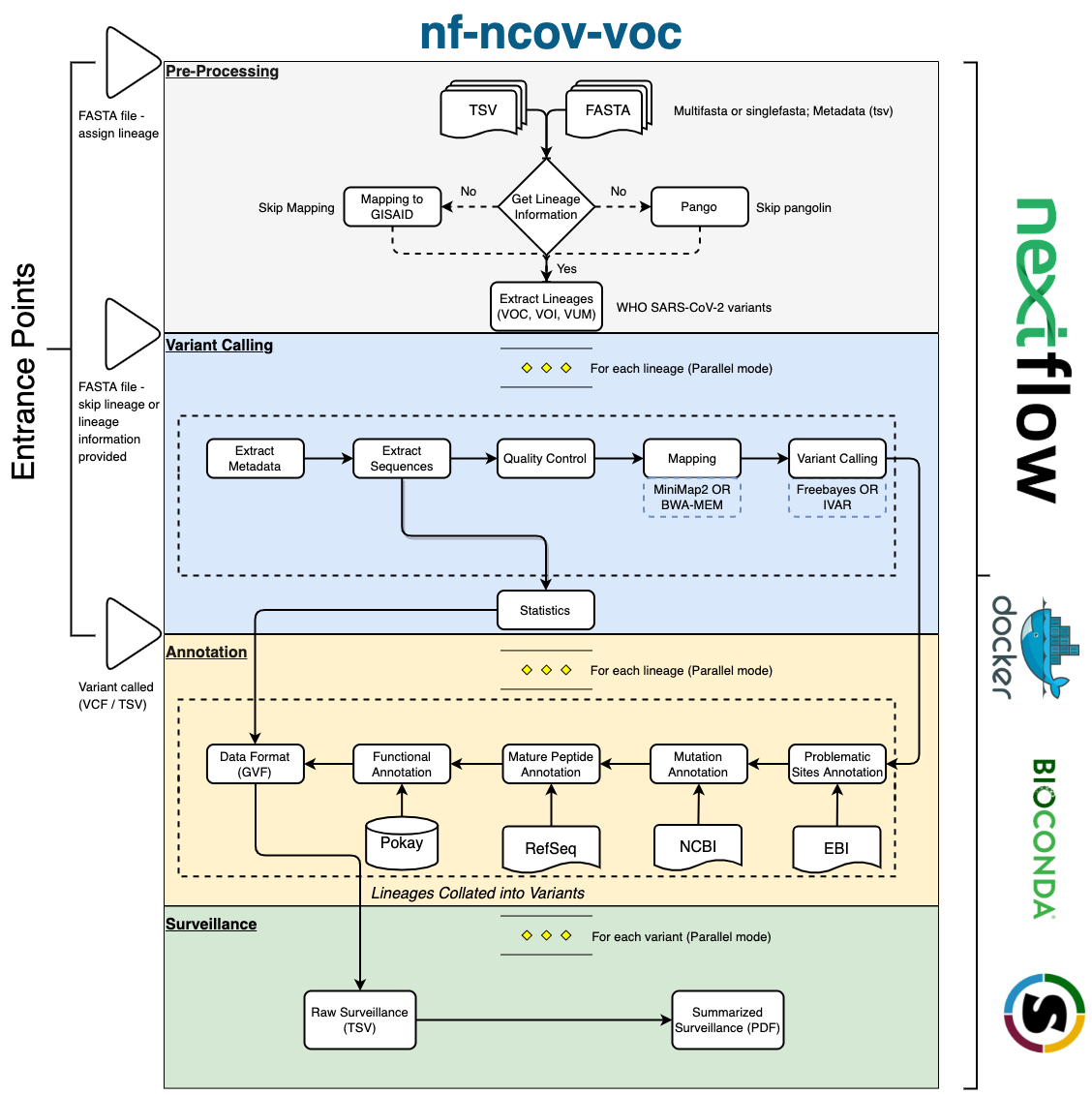
nf-ncov-voc is a bioinformatics analysis workflow used for performing variant calling on SARS-CoV-2 genomes to identify and profile mutations in Variants of Concern (VOCs), Variants of Interest (VOIs) and Variants under Monitoring (VUMs). This workflow has four main stages - Preprocessing, Genomic Analysis (Variant Calling) , Functional Annotation and Surveillance. nf-ncov-voc workflow can be used in combination with an interactive visualization tool COVID-MVP or as a stand-alone high-throughput analysis tool to produce mutation profiles and surveillance reports.
As an input, nf-ncov-voc workflow requires SARS-CoV-2 consensus
sequences in FASTA format and Metadata file in TSV format.
Sequences in pre-processing stage are filtered using Metadata
variables, quality filtered and assigned lineages. Sequences
assigned as VOCs, VOIs and VUMs are then mapped to SARS-CoV-2 genome,
variant called and normalized in Genomic Analysis (Variant Calling)
module. Mutations called are then annotated in several stages
including flagging the potential contaminated sites, mutation
annotation, genomic feature annotation, mature peptide annotation
and finally respective biological functional impact using the
manually curated effort Pokay.
(lead by Paul Gordon @nodrogluap).
Finally, in the surveillance module, these functional profiles are
summarized using functional indicators to highlight key functions
and mutations responsible for them for e.g. P618H role in
convalescent plasma escape.
The workflow is built using Nextflow-
DSL2, a workflow
tool to run tasks across multiple compute infrastructures in a very
portable manner. It can use conda/Docker/Singularity
containers making installation trivial and results highly reproducible.
A detailed structure and each module of the workflow is presented below in the dataflow diagram
This module offers two ways to get lineage information for each
genome in FASTA file and listed respectively in Metadata file
unless a column pango_lineage is already available in which case
both options can be skipped. First option is to use
PANGOLIN to assign
lineages and merge the metadata with pangolin report. This
step can be skipped by passing --skip_pangolin. The second option
is to map input metadata to GISAID metadata
file (which can be provided by --gisaid_metadata parameter) if the
genomes are available in GISAID. This option is faster and
computationally less expensive, though limits to only genomes
available in GISAID. This option can be skipped by
using --skip_mapping.
This module currently supports two different modes - "reference" &
"user" which can be passed with --mode reference or --mode user. By default, --mode reference is activated which allows you
to build a reference library for each lineage and subsequently each
variant for comparative analysis. This mode can take FASTA file
with multiple genomes (recommended & default) or single
genome with a metadata file that should have one column atleast
(pango_lineage) as minimal metadata
(see Workflow Summary for detailed options).
The workflow has numerous options for several steps. For
example, in mode --reference user can use BWAMEM using --bwa
instead of MINIMAP2 (default) for mapping consensus sequences to
reference genome. Similarly, ivar with parameter --ivar for
variant calling instead of freebayes (default) option.
The user mode (--mode user) is by default active when using
interactive visualization through
COVID-MVP where a user can
upload GVF file for comparative analysis against the reference data.
Uploaded dataset can be a FASTA file or variant called VCF file.
In this module, the variant called VCF file for each lineage is
converted into a GVF (Genomic Variant Format) file and annotated
with functional information using
Pokay. GVF is a variant of
GFF3 format that is standardized for describing genomic mutations;
it is used here because it can describe mutations across multiple
rows, and because the "#attributes" column can store information in
custom key-value pairs. The key-value pairs added at this stage
include for each mutation: VOC/VOI status, clade-defining status
(for reference lineages), and functional annotations parsed using
vcf2gvf.py
file written in python.
Different GVF files for the same variant are then collated and
summarized into a TSV file that contains mutation prevalence,
profile and functional impact. Further TSV file is also summarized
as a more human friendly and impactful surveillance report in a
PDF format. Relevant/important indicators can be specified in the
tsv file.
This feature of surveillance reports can be used to identify new
clusters, important mutations, and track their transmission and
prevalence trends. However, if not required, this step can be
skipped using --skip_surveillance. An example of surveillance file
for Omicron variant using
VirusSeq Data Portal is available in
Docs
See the parameters docs for all available options when running the workflow.
-
Install
Nextflow(>=21.04.0) -
Install any of
Docker,SingularityorCondafor full pipeline reproducibility see recipes -
Download the pipeline and run with help for detailed parameter options:
nextflow run nf-ncov-voc/main.nf --help
N E X T F L O W ~ version 21.04.3 Launching `main.nf` [berserk_austin] - revision: 93ccc86071 Usage: nextflow run main.nf -profile [singularity | docker | conda) --prefix [prefix] --mode [reference | user] [workflow-options] Description: Variant Calling workflow for SARS-CoV-2 Variant of Concern (VOC) and Variant of Interest (VOI) consensus sequences to generate data for Visualization. All options set via CLI can be set in conf directory Nextflow arguments (single DASH): -profile Allowed values: conda & singularity Mandatory workflow arguments (mutually exclusive): --prefix A (unique) string prefix for output directory for each run. --mode A flag for user uploaded data through visualization app or high-throughput analyses (reference | user) (Default: reference) Optional: Input options: --seq Input SARS-CoV-2 genomes or consensus sequences (.fasta file) --meta Input Metadata file of SARS-CoV-2 genomes or consensus sequences (.tsv file) --userfile Specify userfile (fasta | vcf) (Default: None) --gisaid_metadata If lineage assignment is preferred by mapping metadata to GISAID metadata file, provide the metadata file (.tsv file) --variants Provide a variants file (.tsv) (Default: /Users/au572806/GitHub/nf-ncov-voc/assets/ncov_variants/variants_who.tsv) --outdir Output directory (Default: /Users/au572806/GitHub/nf-ncov-voc/results) --gff Path to annotation gff for variant consequence calling and typing. (Default: /Users/au572806/GitHub/nf-ncov-voc/assets/ncov_genomeFeatures/MN908947.3.gff3) --ref Path to SARS-CoV-2 reference fasta file (Default: /Users/au572806/GitHub/nf-ncov-voc/assets/ncov_refdb/*) --bwa_index Path to BWA index files (Default: /Users/au572806/GitHub/nf-ncov-voc/assets/ncov_refdb/*) Selection options: --ivar Run the iVar workflow instead of Freebayes(default) --bwamem Run the BWA workflow instead of MiniMap2(default) --skip_pangolin Skip PANGOLIN. Can be used if metadata already have lineage information or mapping is preferred method --skip_mapping Skip Mapping. Can be used if metadata already have lineage information or PANGOLIN is preferred method Preprocessing options: --startdate Start date (Submission date) to extract dataset (yyyy-mm-dd) (Default: "2020-01-01") --enddate Start date (Submission date) to extract dataset (yyyy-mm-dd) (Default: "2022-12-31") Genomic Analysis parameters: BBMAP --maxns Max number of Ns allowed in the sequence in qc process --minlength Minimun length of sequence required for sequences to pass qc filtration. Sequence less than minlength are not taken further IVAR/FREEBAYES --ploidy Ploidy (Default: 1) --mpileupDepth Mpileup depth (Default: unlimited) --var_FreqThreshold Variant Calling frequency threshold for consensus variant (Default: 0.75) --var_MaxDepth Maximum reads per input file depth to call variant (mpileup -d, Default: 0) --var_MinDepth Minimum coverage depth to call variant (ivar variants -m, freebayes -u Default: 10) --var_MinFreqThreshold Minimum frequency threshold to call variant (ivar variants -t, Default: 0.25) --varMinVariantQuality Minimum mapQ to call variant (ivar variants -q, Default: 20) Surveillance parameters: --virusseq True/False (Default: False). If your data is from VirusSeq Data Portal (Canada's Nation COVID-19 genomics data portal). Passing this argument adds an acknowledgment statement to the surveillance report. see https://virusseq-dataportal.ca/acknowledgements
-
Start running your own analysis!
-
Typical command for reference mode when Metadata File don't have lineage information:
nextflow nf-ncov-voc/main.nf \ -profile <conda, singularity, docker> \ --prefix <testing> \ --mode reference \ --startdate <2020-01-01> \ --enddate <2020-01-01> \ --seq <Sequence File> \ --meta <Metadata File> \ --skip_mapping \ --outdir <Output Dir> -
Typical command for reference mode when Metadata File already have lineage information:
nextflow nf-ncov-voc/main.nf \ -profile <conda, singularity, docker> \ --prefix <testing> \ --mode reference \ --startdate <2020-01-01> \ --enddate <2020-01-01> \ --seq <Sequence File> \ --meta <Metadata File> \ --skip_mapping \ --skip_pangolin \ --outdir <Output Dir> -
An executable Python script called
functional_annotation.pyhas been provided if you would like to update the functional annotations fromPOKAY. This will create a new file which should replace the current file in assets/functional_annotation.
-
This workflow and scripts are written and conceptually designed by
Many thanks to others who have helped out and contributed along the way too, including (but not limited to)*: Canadian COVID Genomics Network - VirusSeq, Data Analytics Working Group
For further information or help, don't hesitate to get in touch at mzanwar@sfu.ca or wwshiao
An extensive list of references for the tools used by the workflow can be found in the CITATIONS.md file.